Deposition Date
2025-05-27
Release Date
2025-08-06
Last Version Date
2025-08-06
Entry Detail
PDB ID:
9OTU
Keywords:
Title:
Crystal Structure of Salmonella FraB Deglycase, E214A Mutant, Crystal Form 5
Biological Source:
Source Organism(s):
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.87 Å
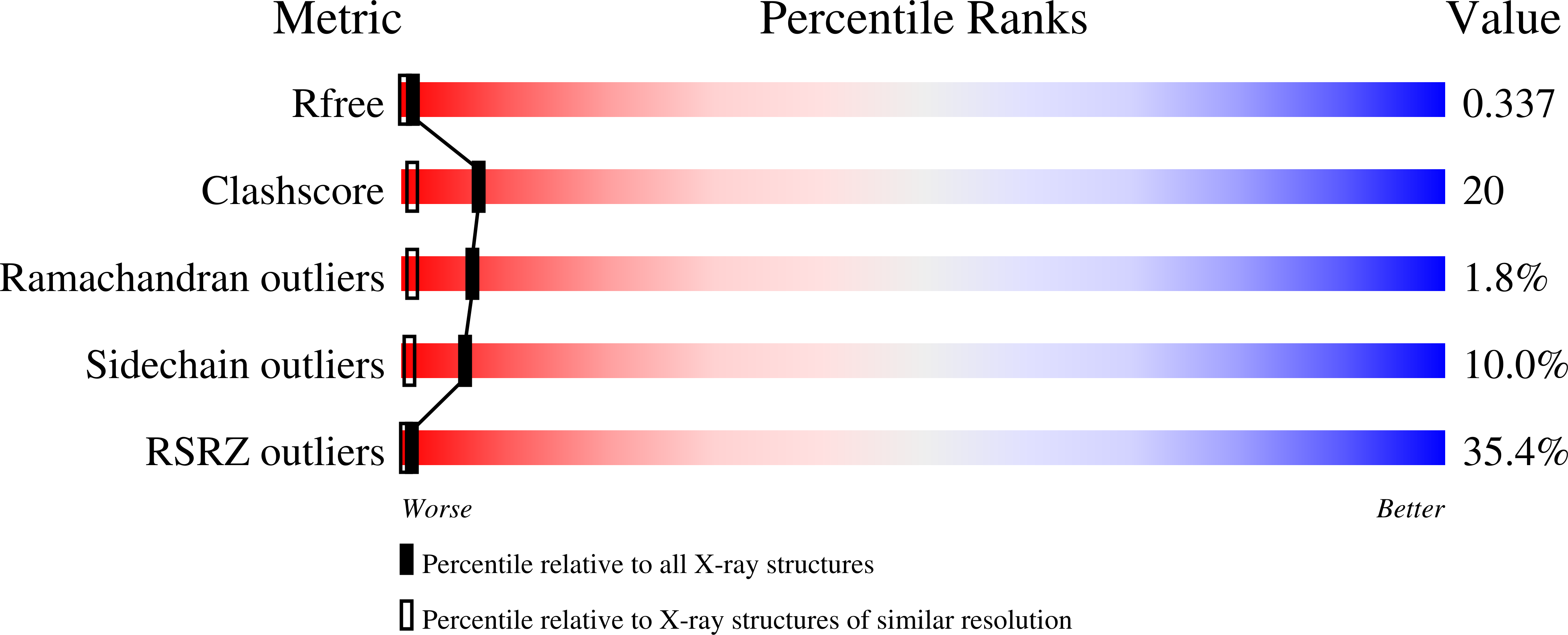
R-Value Free:
0.33
R-Value Work:
0.26
Space Group:
P 32 2 1


