Deposition Date
2025-02-24
Release Date
2026-01-21
Last Version Date
2026-01-21
Entry Detail
PDB ID:
9NH6
Keywords:
Title:
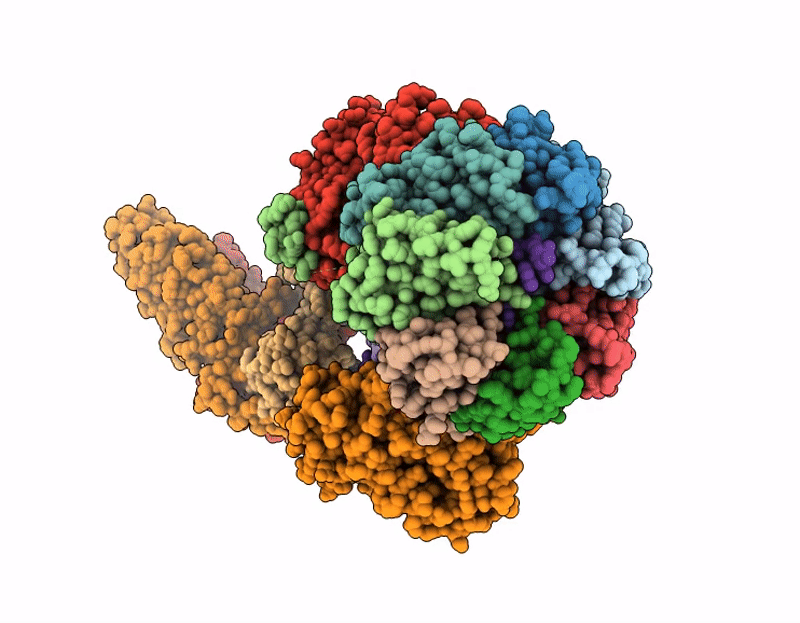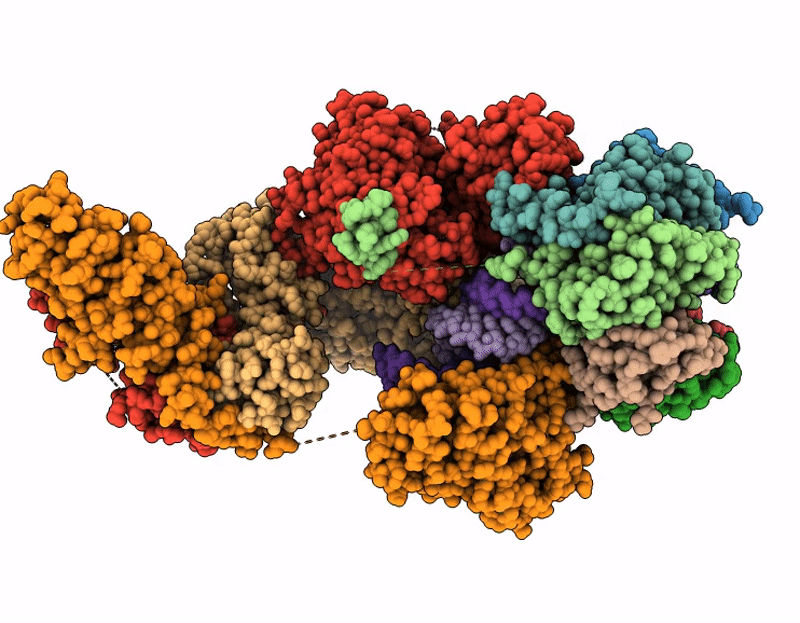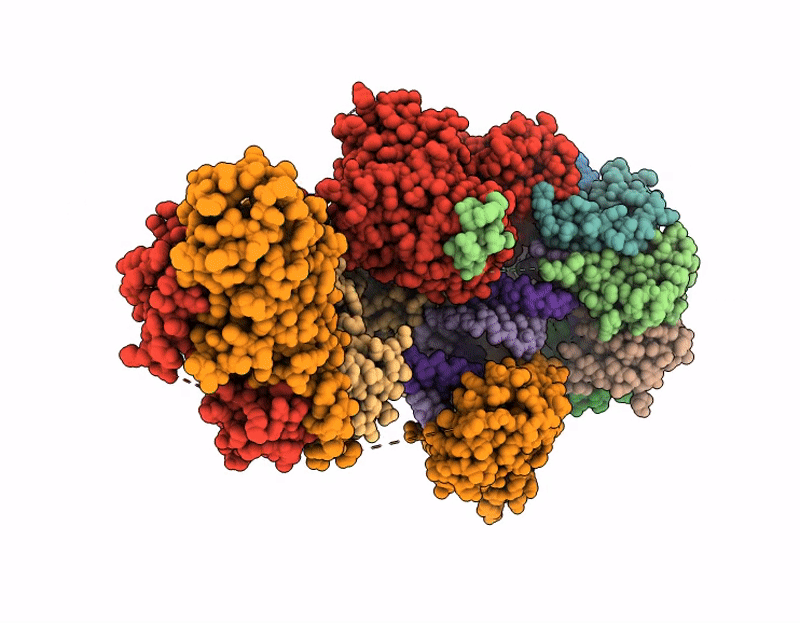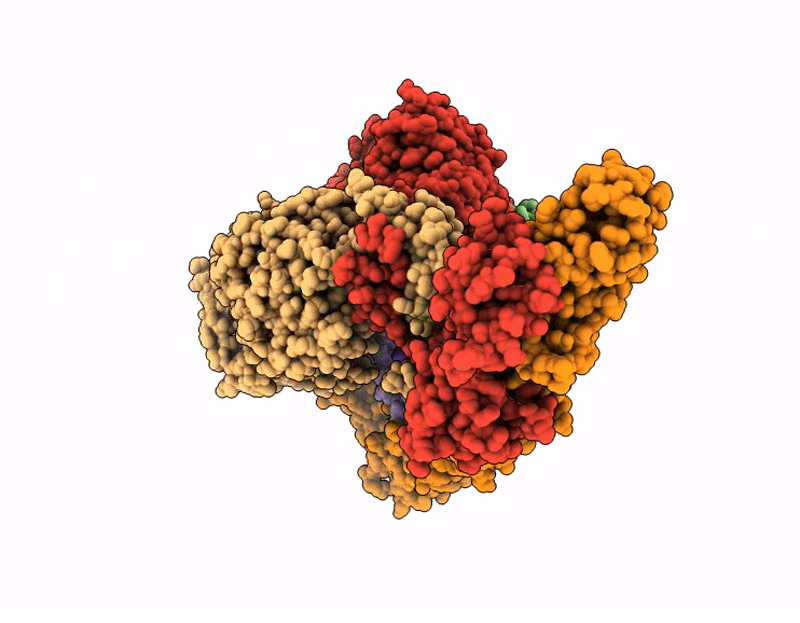
CRYO-EM STRUCTURE OF HUMAN U7 SNRNP WITH METHYLATED noncleavable H2A* SUBSTRATE PRE-MRNA (COMPOSITE MAP)
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Trichoplusia ni (Taxon ID: 7111)
Trichoplusia ni (Taxon ID: 7111)
Expression System(s):
Method Details:
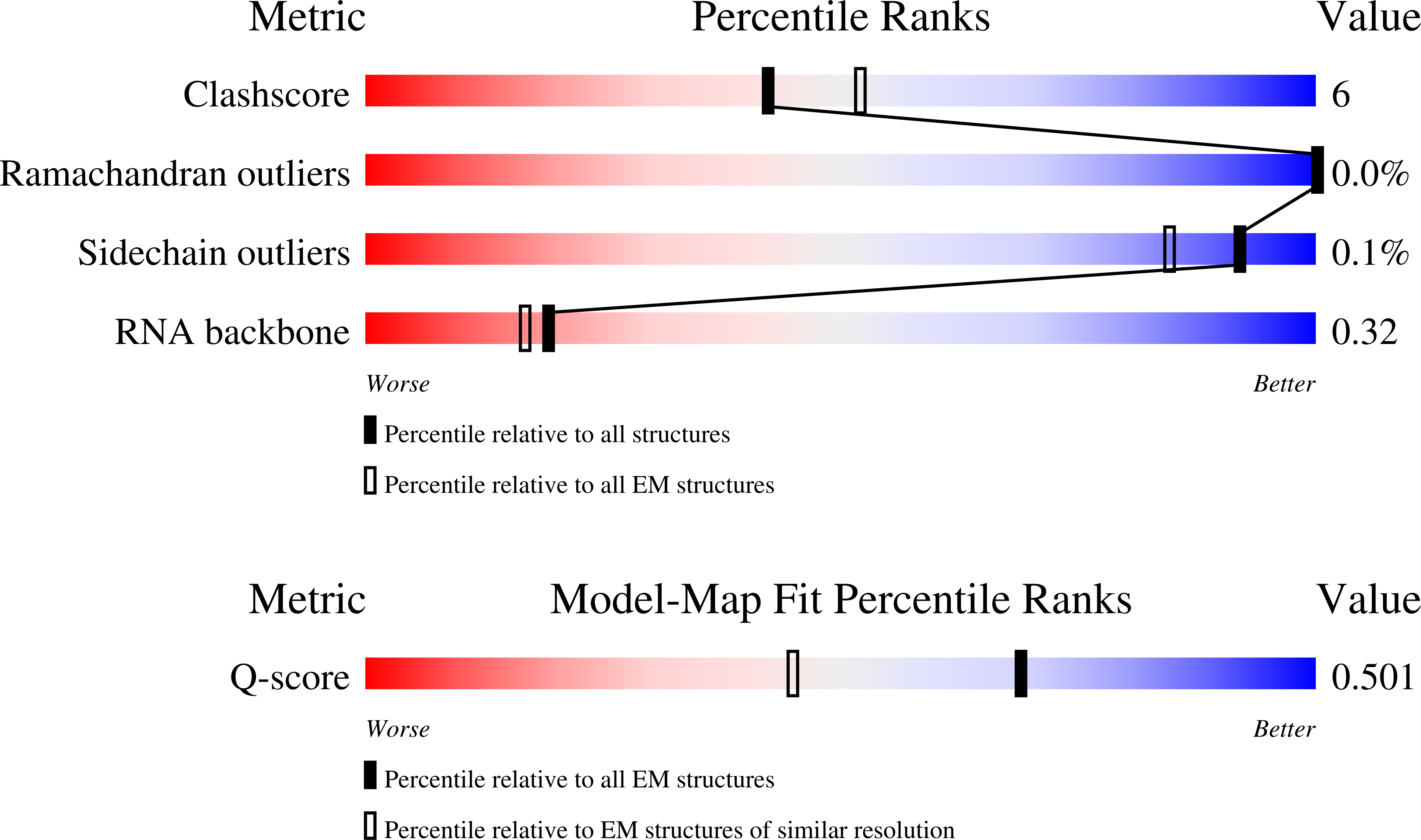
Experimental Method:
Resolution:
2.82 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE