Deposition Date
2024-12-02
Release Date
2025-08-06
Last Version Date
2026-02-18
Entry Detail
PDB ID:
9KTJ
Keywords:
Title:
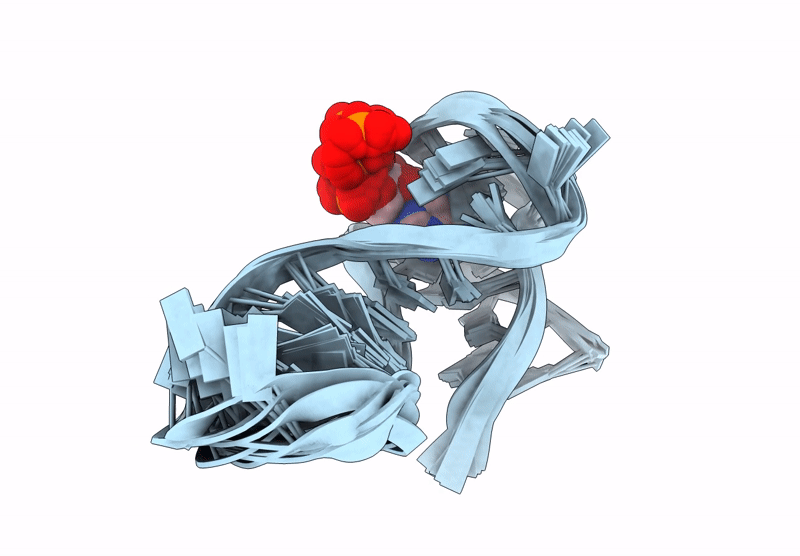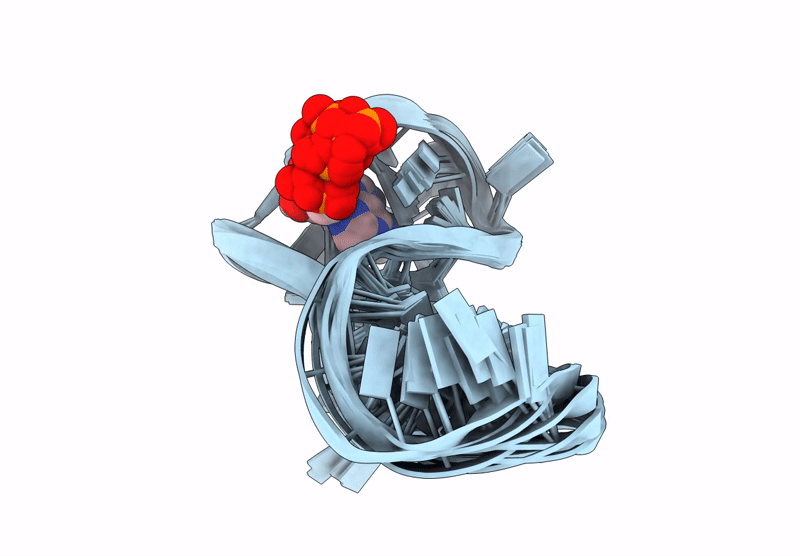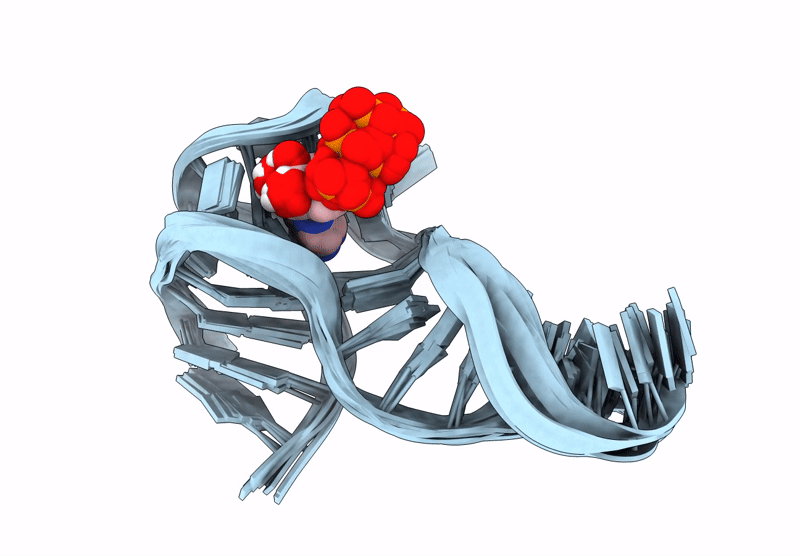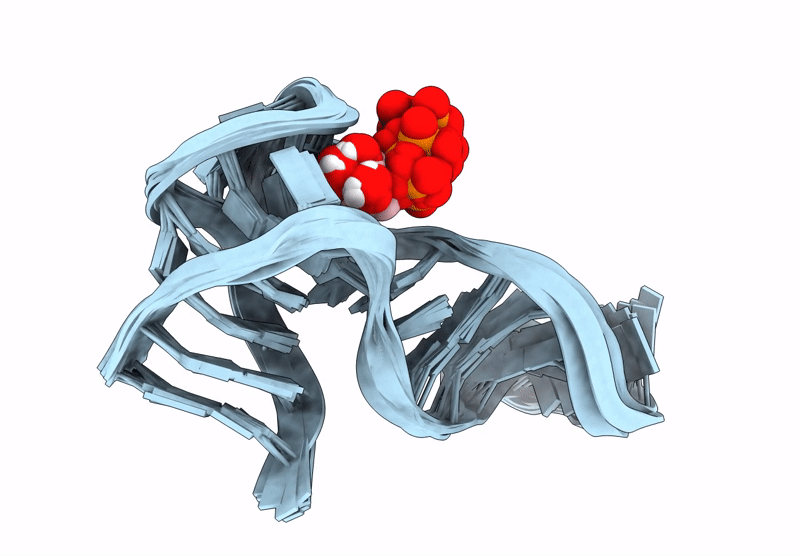
Solution NMR structures of ATP-binding DNA aptamer in complex with ATP
Biological Source:
Source Organism(s):
synthetic construct (Taxon ID: 32630)
Method Details:
Experimental Method:
Conformers Calculated:
100
Conformers Submitted:
10
Selection Criteria:
structures with the lowest energy