Deposition Date
2024-05-14
Release Date
2024-06-26
Last Version Date
2026-04-01
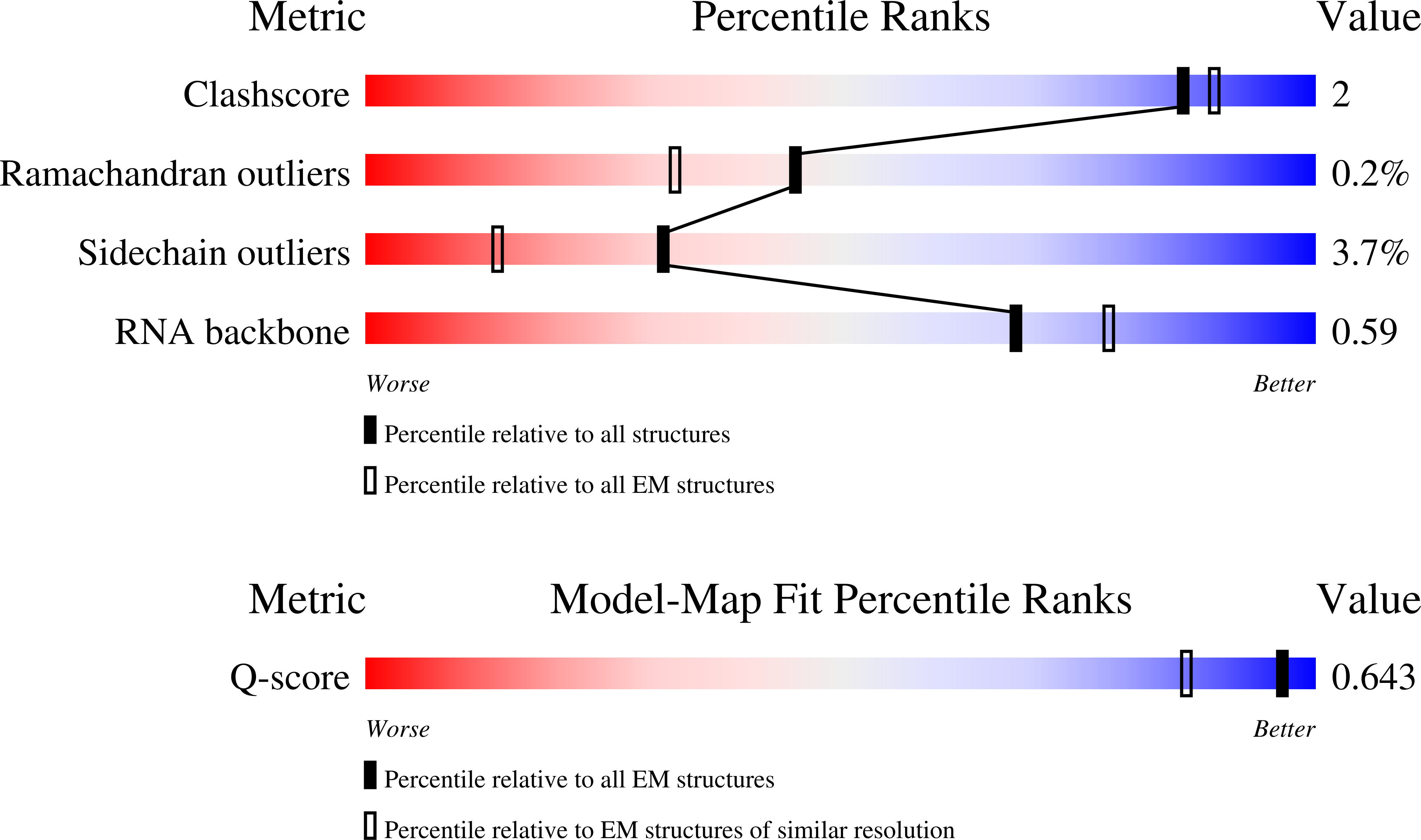
Method Details:
Experimental Method:
Resolution:
2.40 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE