Deposition Date
2024-08-12
Release Date
2025-01-22
Last Version Date
2026-02-18
Entry Detail
PDB ID:
9D4N
Keywords:
Title:
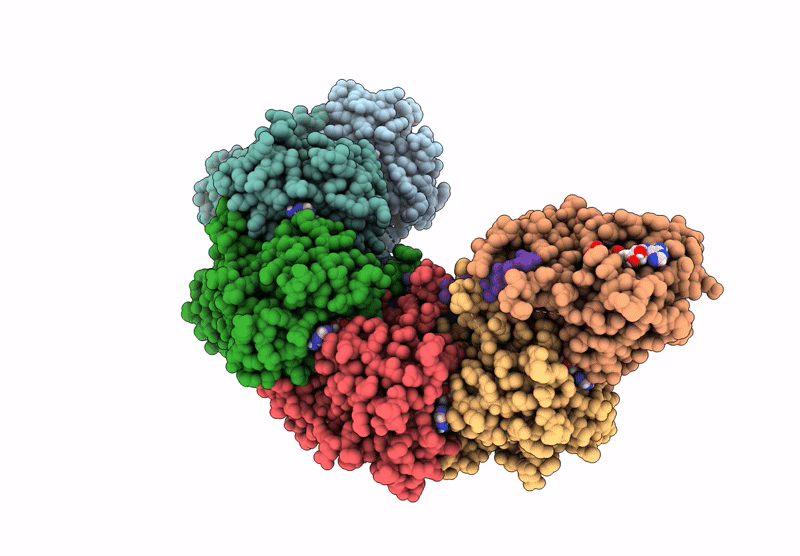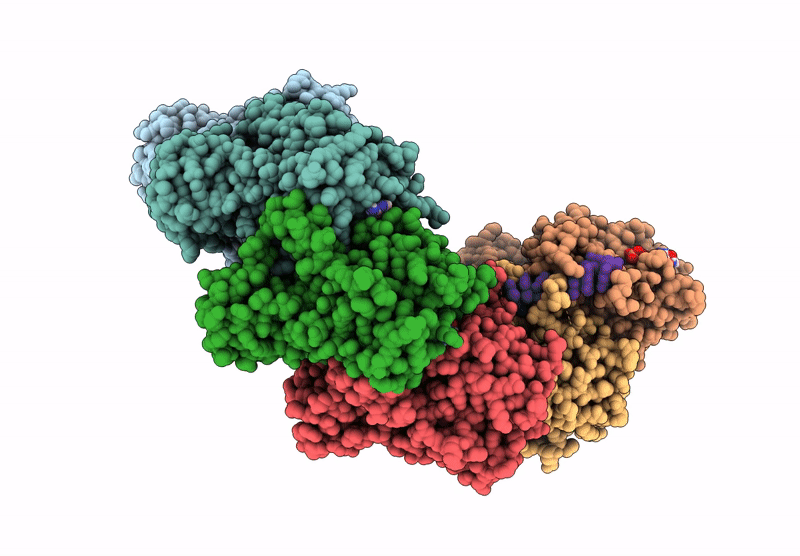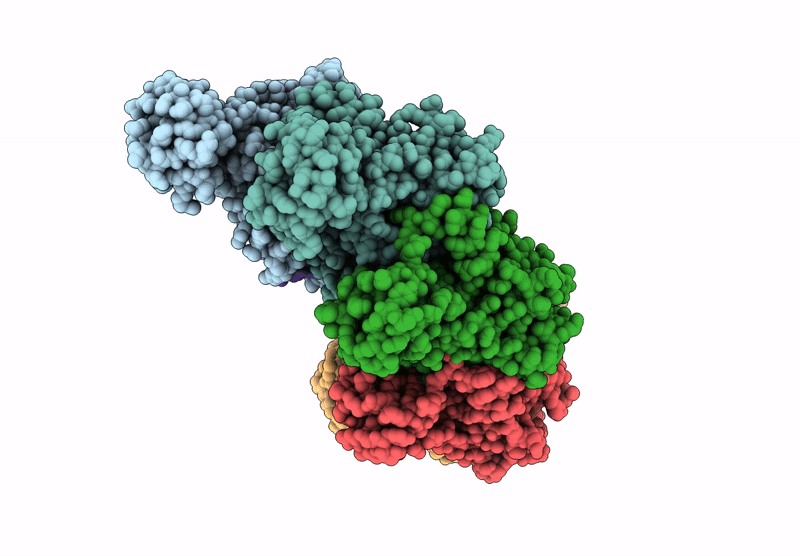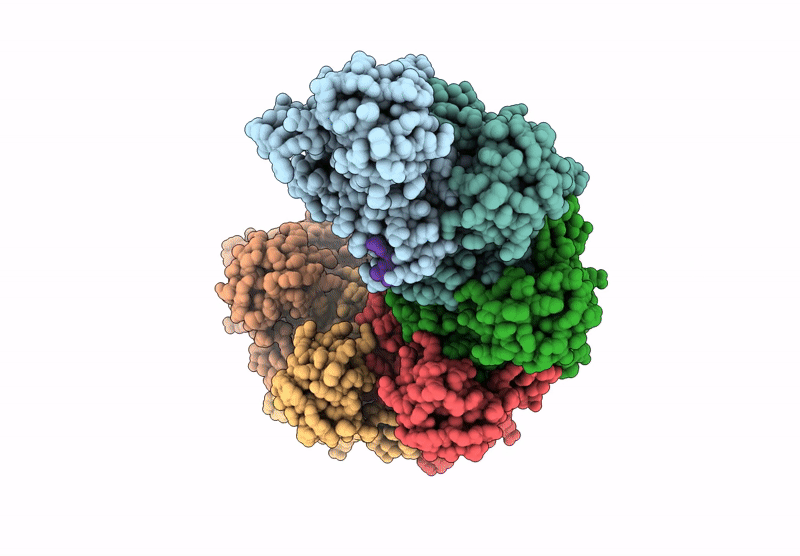
The cryo-EM structure of the yeast Dmc1 filament bound to ssDNA in the presence of ATP
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae (Taxon ID: 4932)
Expression System(s):
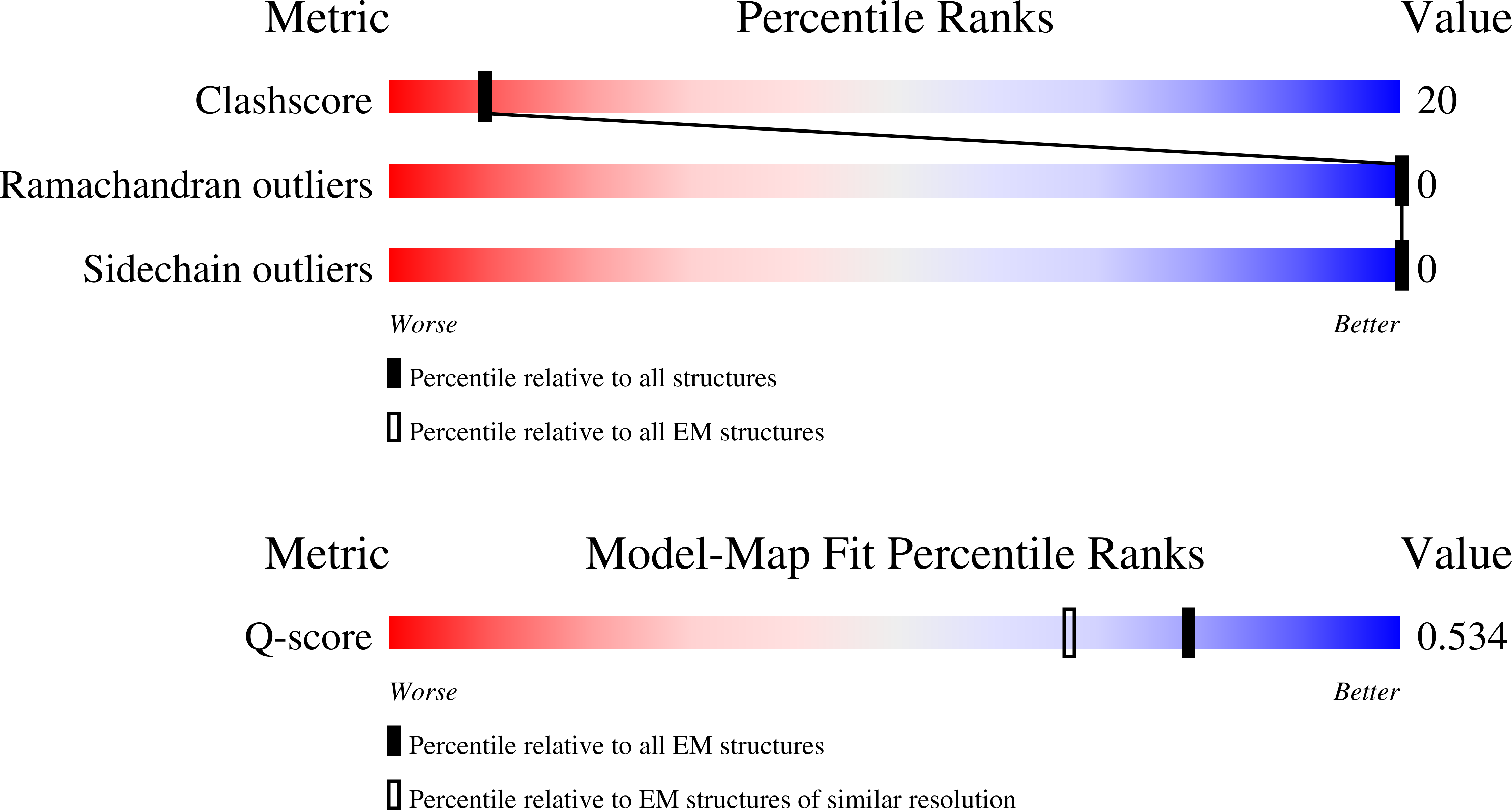
Method Details:
Experimental Method:
Resolution:
2.90 Å
Aggregation State:
FILAMENT
Reconstruction Method:
HELICAL