Deposition Date
2023-12-04
Release Date
2024-05-01
Last Version Date
2024-05-01
Entry Detail
PDB ID:
8V84
Keywords:
Title:
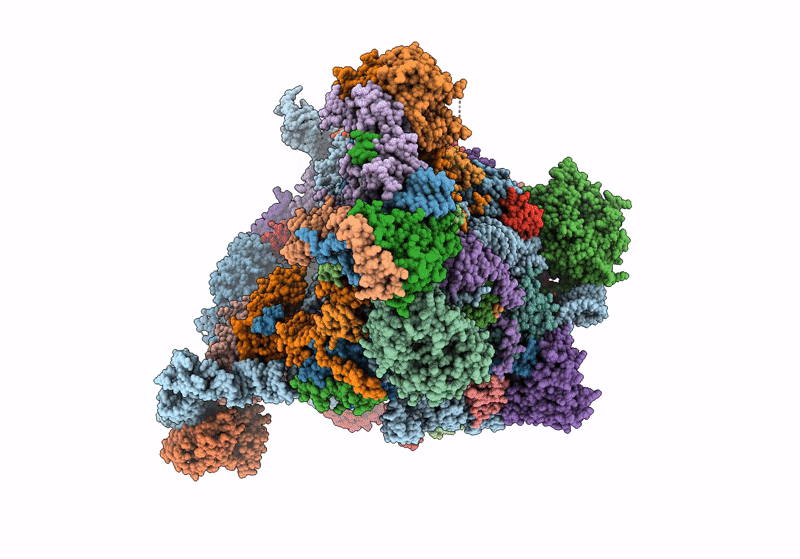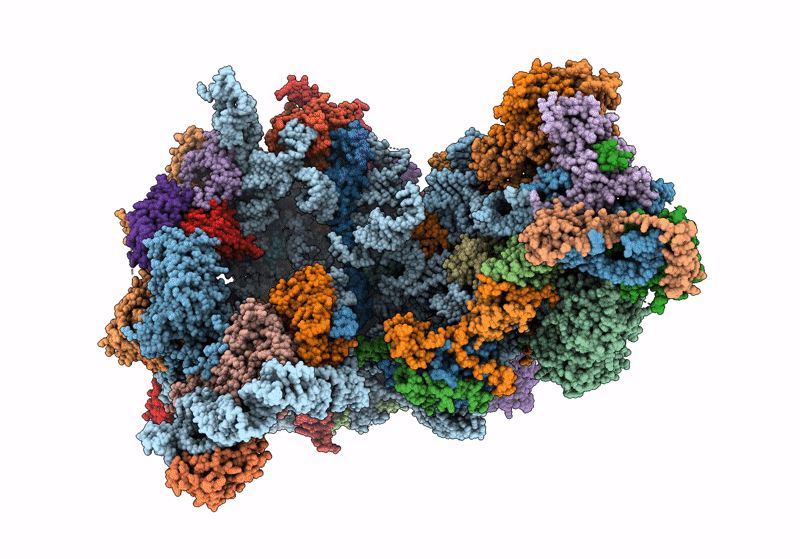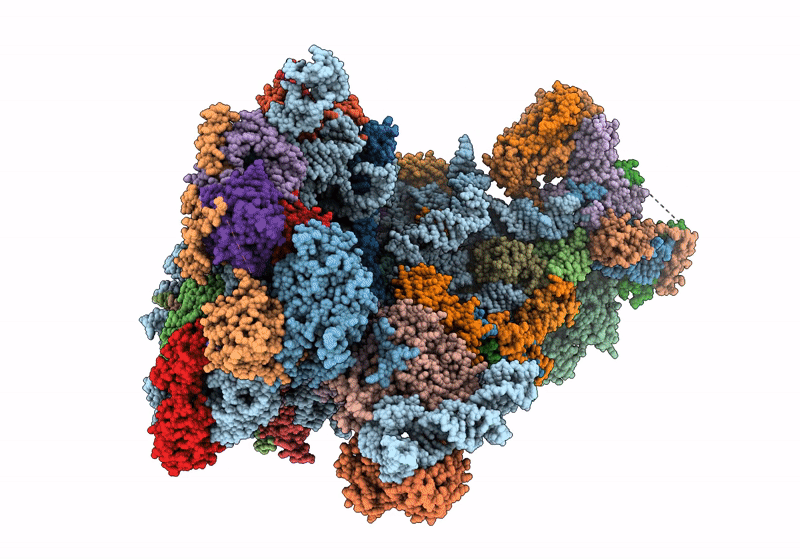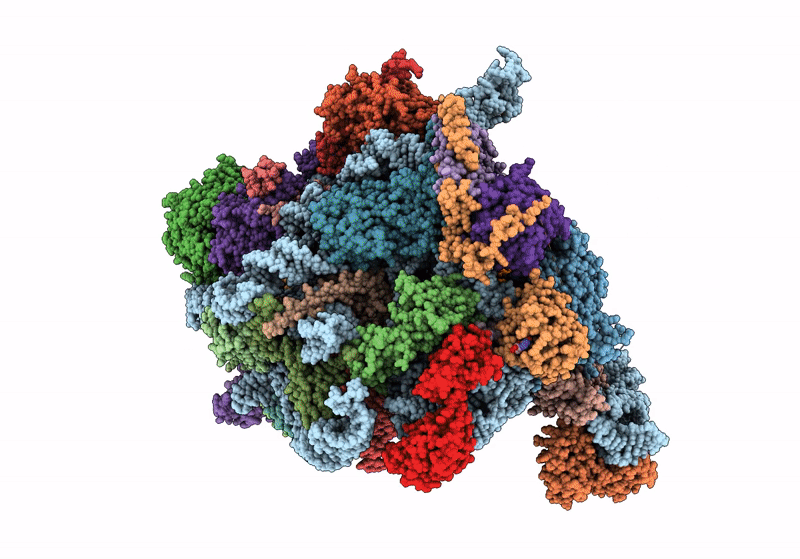
60S ribosome biogenesis intermediate (Dbp10 catalytic structure - Overall map)
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae BY4741 (Taxon ID: 1247190)
Saccharomyces cerevisiae BY4741-AV16 (Taxon ID: 1390929)
Saccharomyces cerevisiae BY4741-AV16 (Taxon ID: 1390929)
Method Details:
Experimental Method:
Resolution:
2.70 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE