Deposition Date
2023-10-05
Release Date
2024-06-19
Last Version Date
2024-12-25
Entry Detail
PDB ID:
8UGJ
Keywords:
Title:
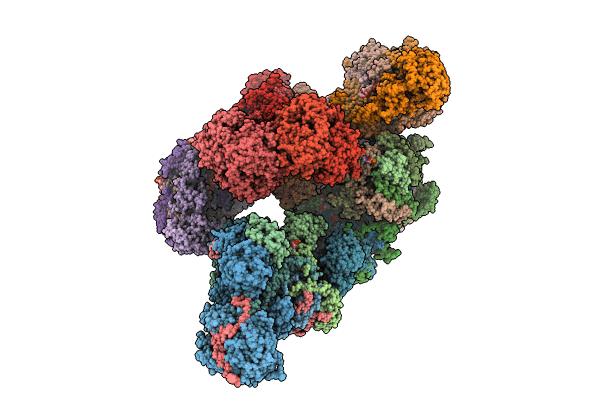
In-situ structure of typeB supercomplex in respiratory chain (composite)
Biological Source:
Source Organism(s):
Sus scrofa (Taxon ID: 9823)
Method Details:
Experimental Method:
Resolution:
2.30 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE