Deposition Date
2023-12-07
Release Date
2024-02-21
Last Version Date
2024-03-13
Entry Detail
PDB ID:
8RD8
Keywords:
Title:
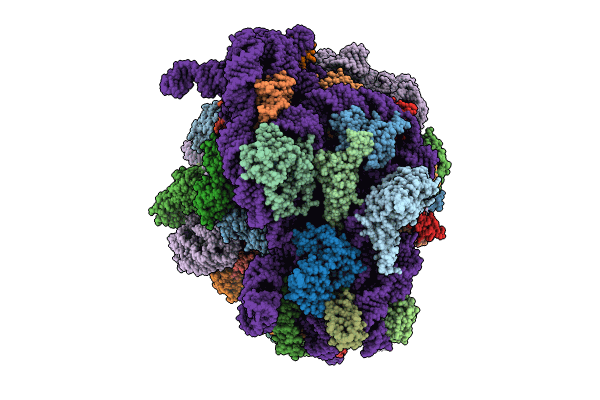
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).
Biological Source:
Source Organism(s):
Psychrobacter urativorans (Taxon ID: 45610)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.62 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE