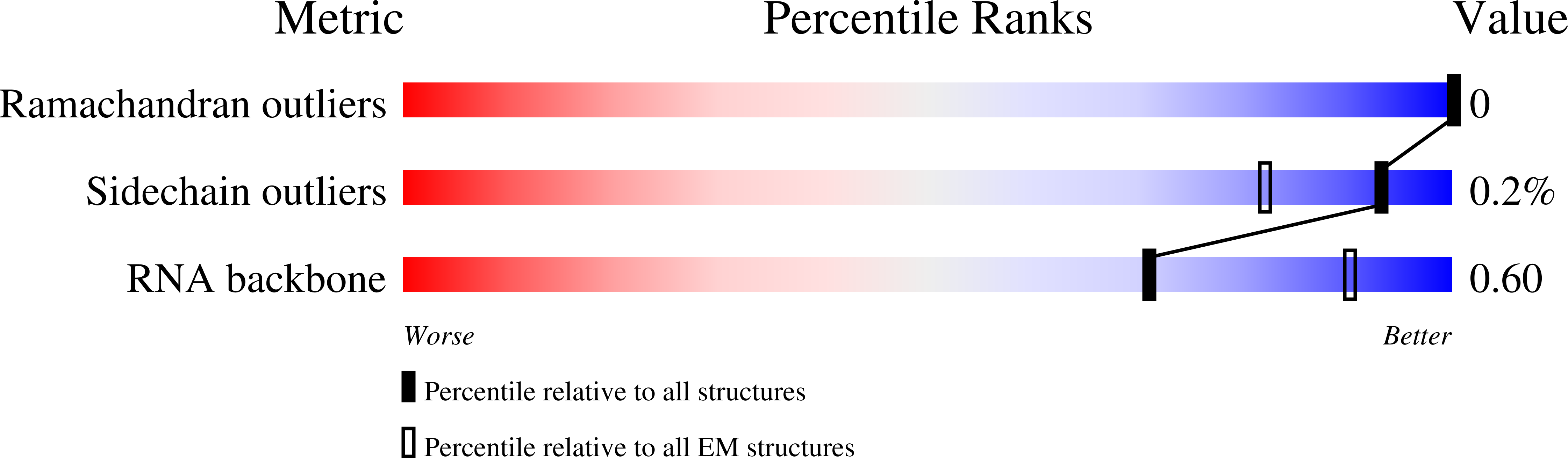
Deposition Date
2023-03-23
Release Date
2023-06-14
Last Version Date
2024-06-26
Entry Detail
PDB ID:
8OIT
Keywords:
Title:
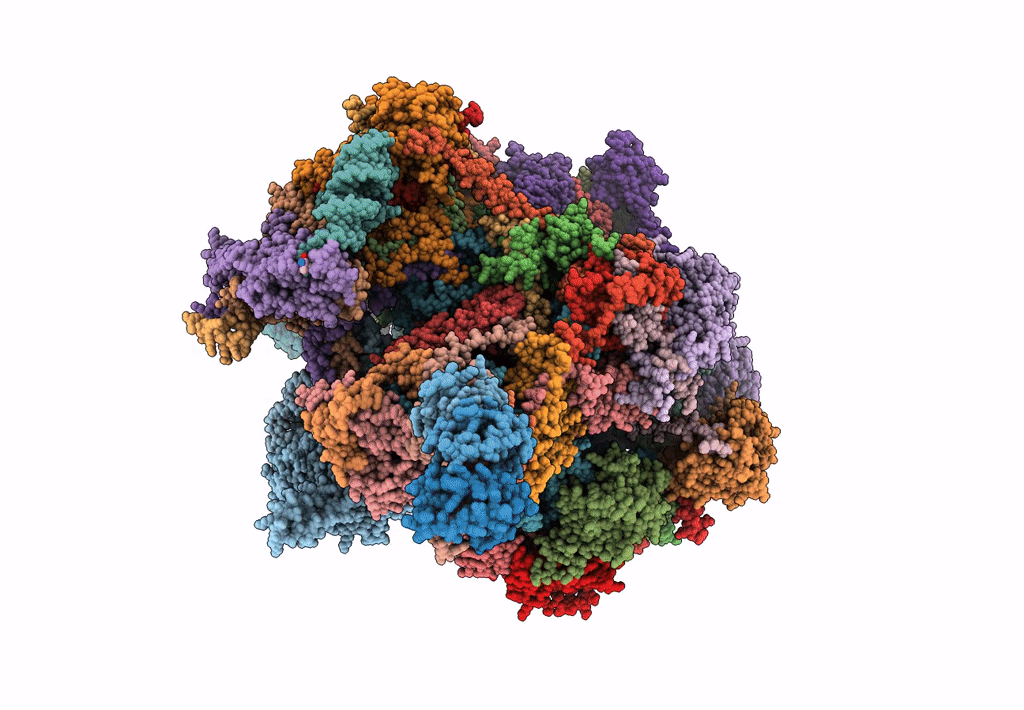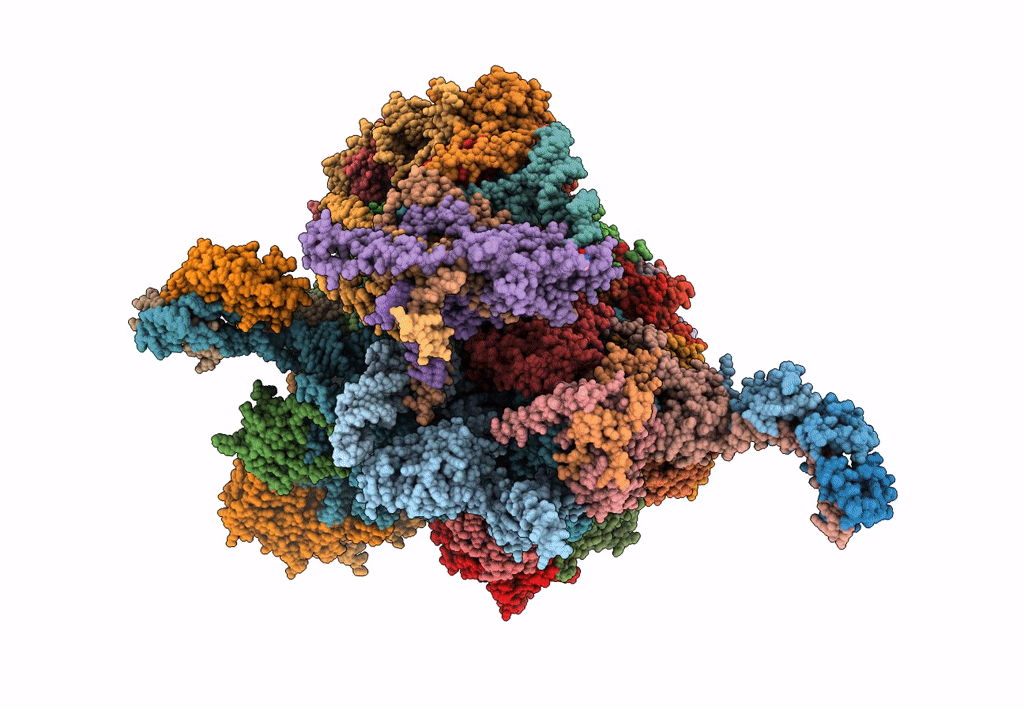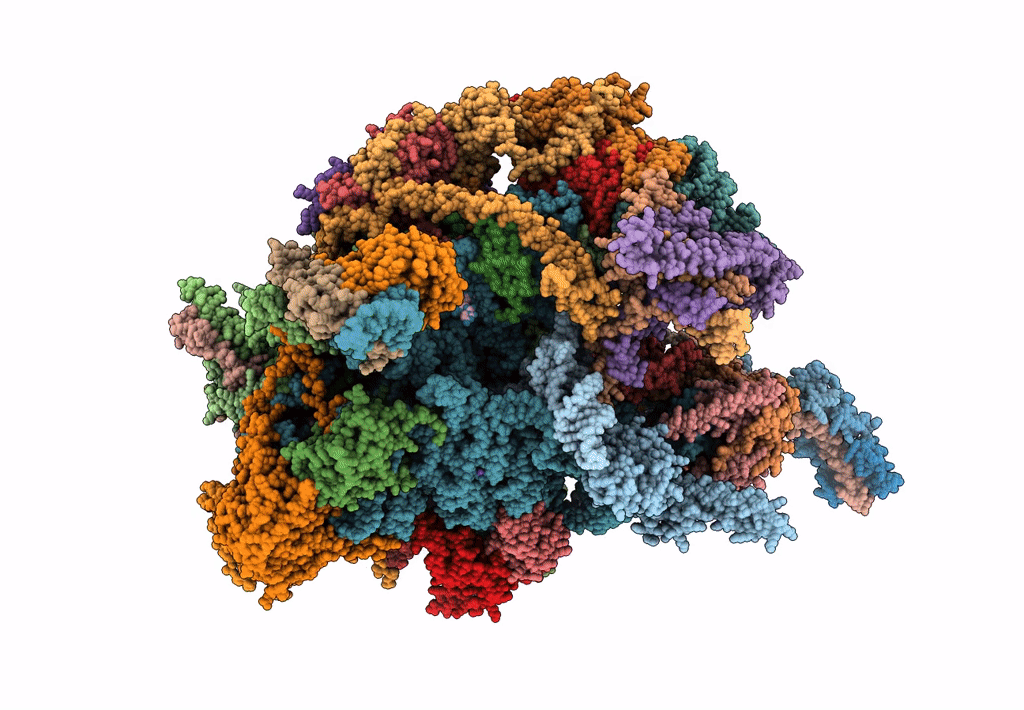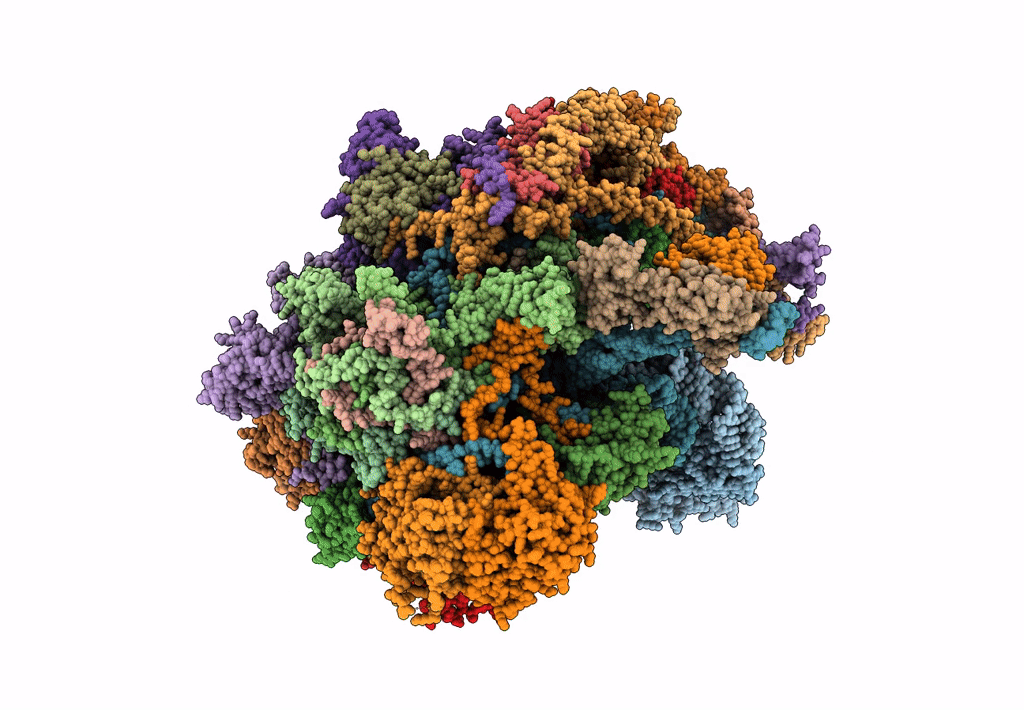
39S human mitochondrial large ribosomal subunit with mtRF1 and P-site tRNA
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.90 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE