Deposition Date
2023-01-09
Release Date
2023-06-14
Last Version Date
2026-04-29
Entry Detail
PDB ID:
8FS3
Keywords:
Title:
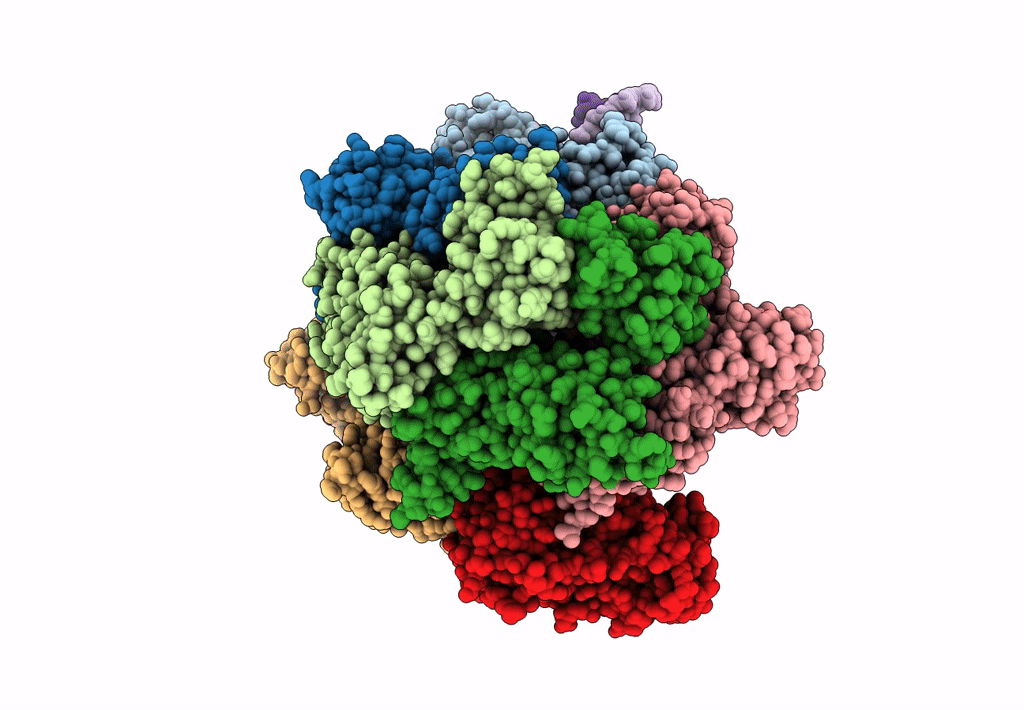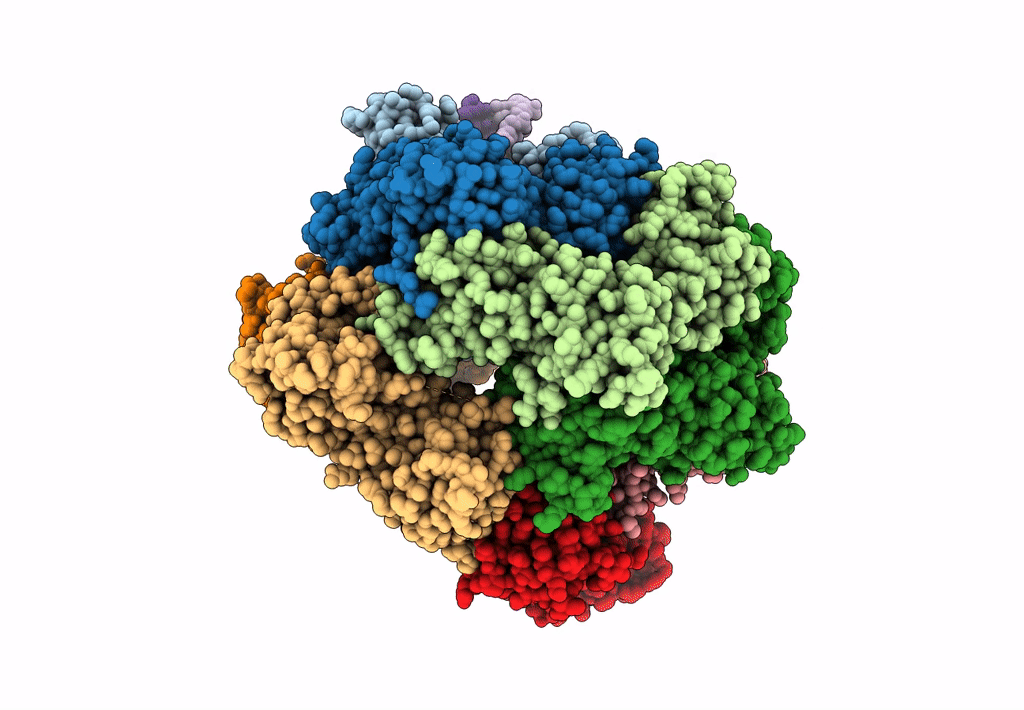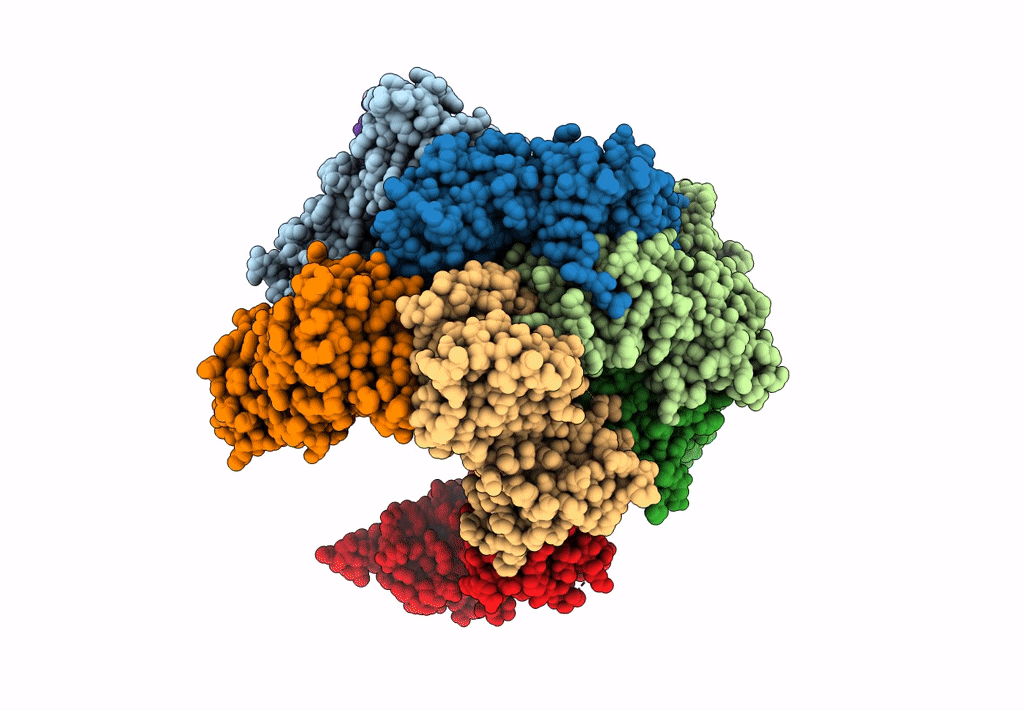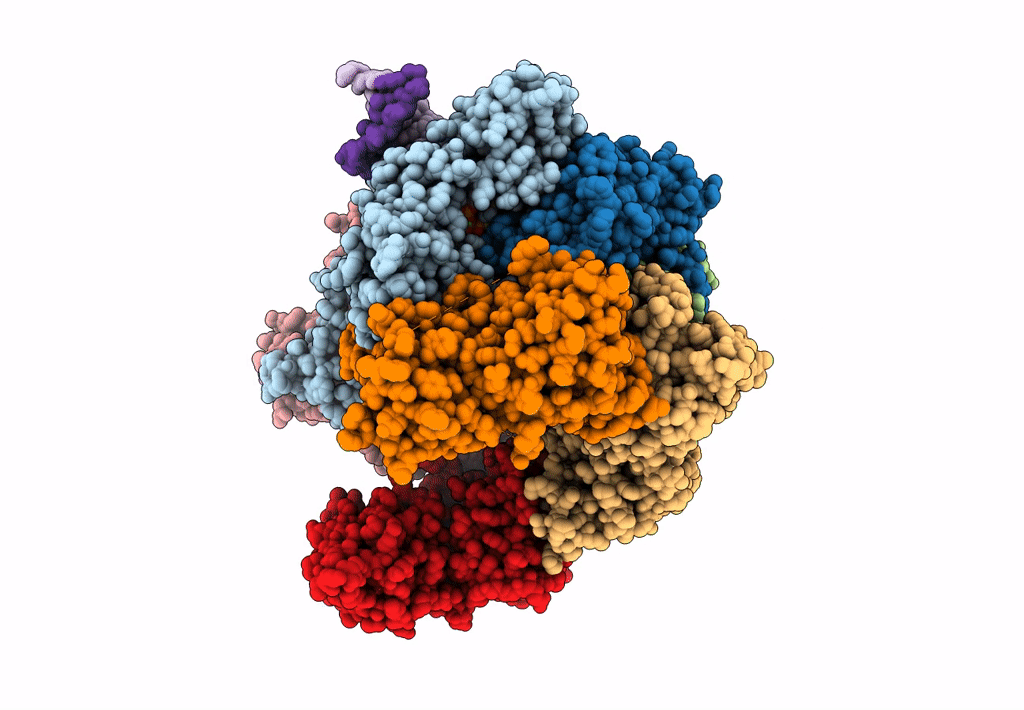
Structure of S. cerevisiae Rad24-RFC loading the 9-1-1 clamp onto a 10-nt gapped DNA in step 1 (open 9-1-1 and shoulder bound DNA only)
Biological Source:
Source Organism(s):
Saccharomyces cerevisiae (Taxon ID: 4932)
synthetic construct (Taxon ID: 32630)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.93 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE