Deposition Date
2022-12-05
Release Date
2023-05-10
Last Version Date
2024-11-06
Entry Detail
PDB ID:
8FDW
Keywords:
Title:
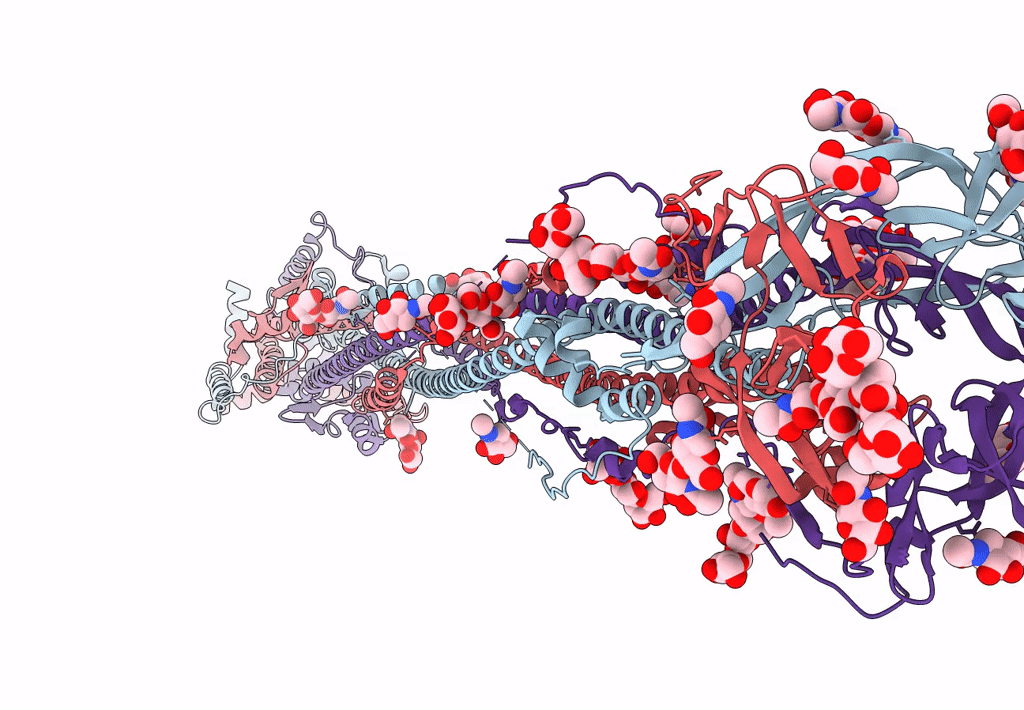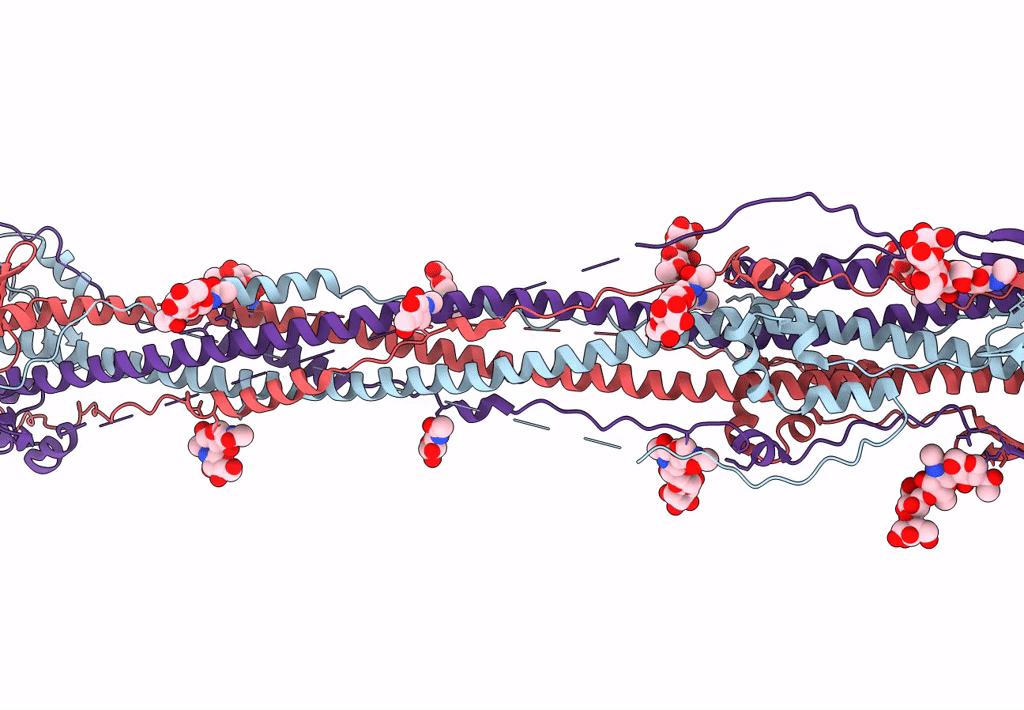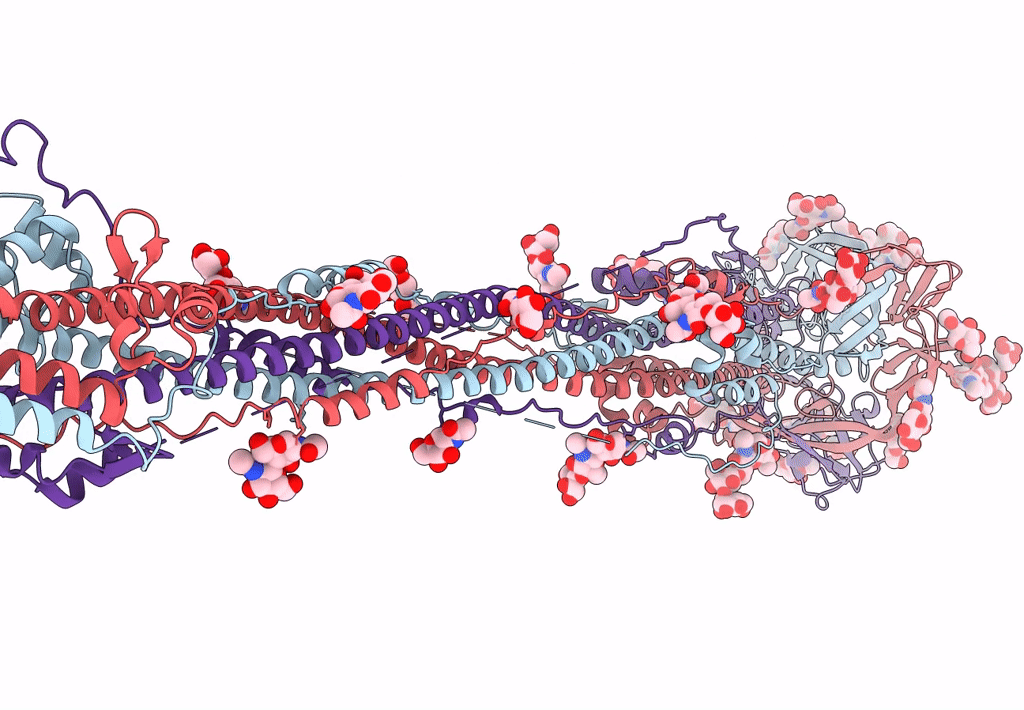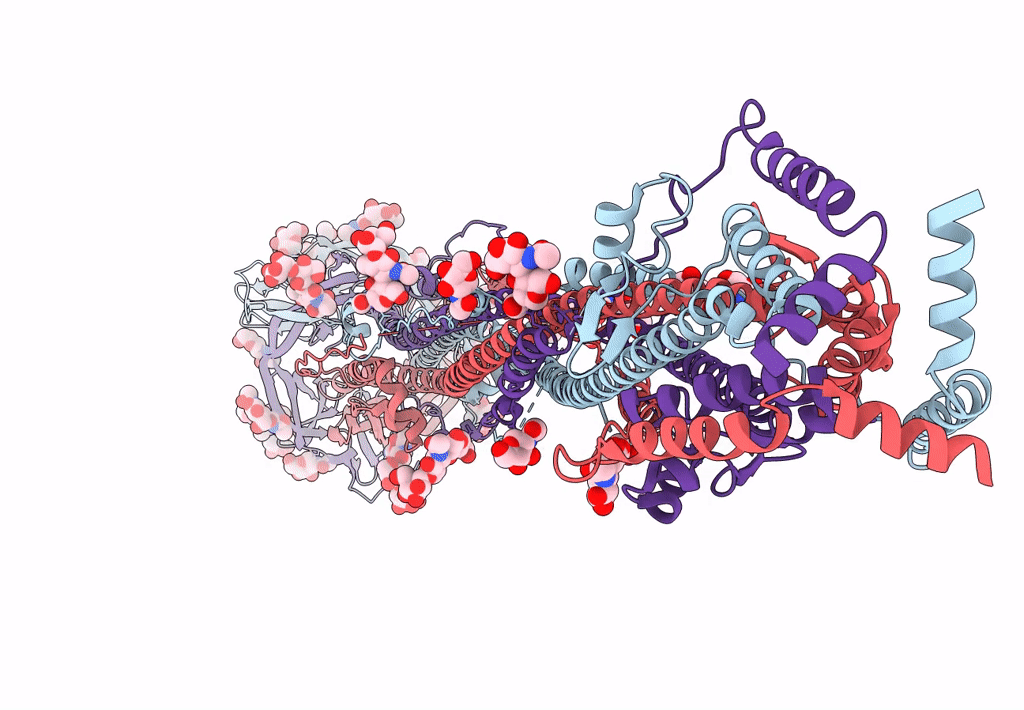
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Biological Source:
Source Organism(s):
Severe acute respiratory syndrome coronavirus (Taxon ID: 2901879)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.90 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE