Deposition Date
2022-09-27
Release Date
2022-12-07
Last Version Date
2024-11-20
Entry Detail
PDB ID:
8B6L
Keywords:
Title:
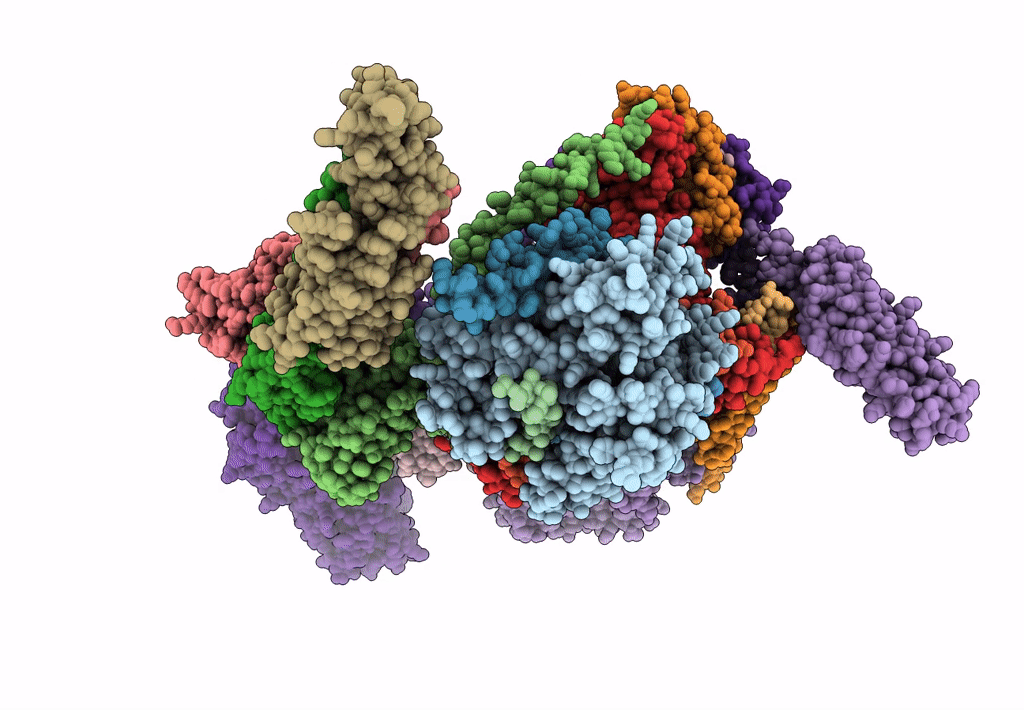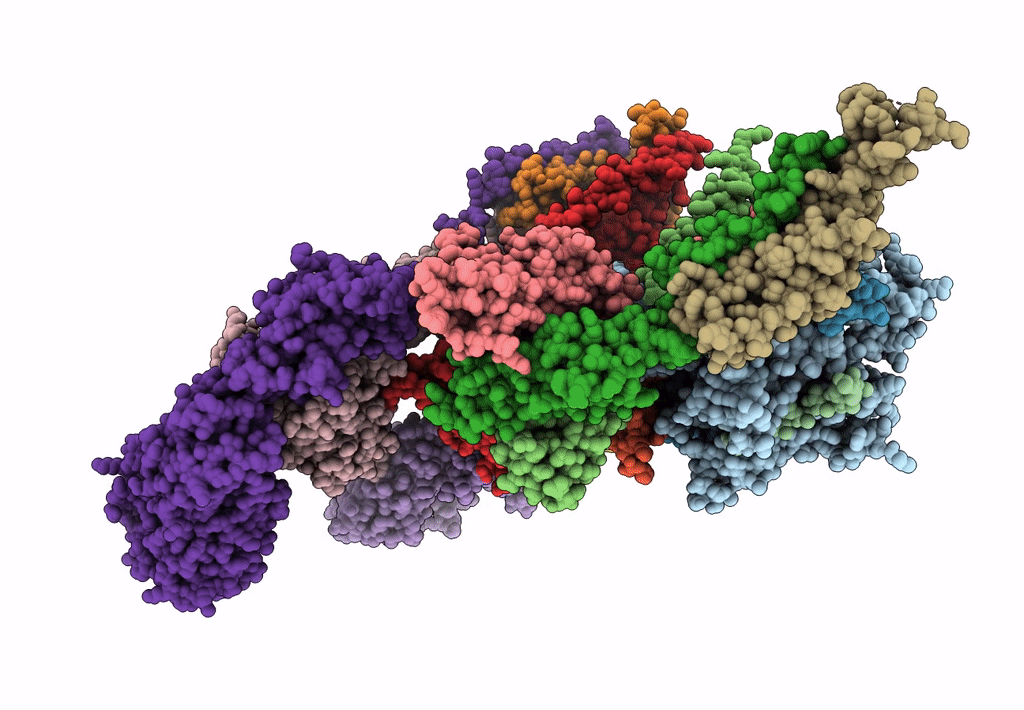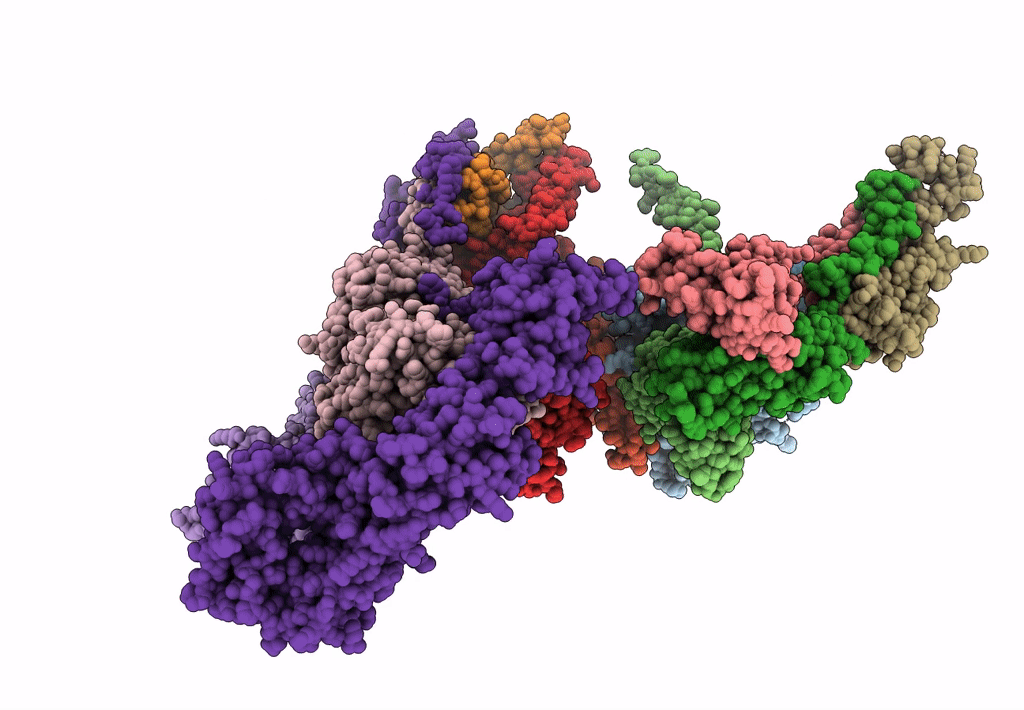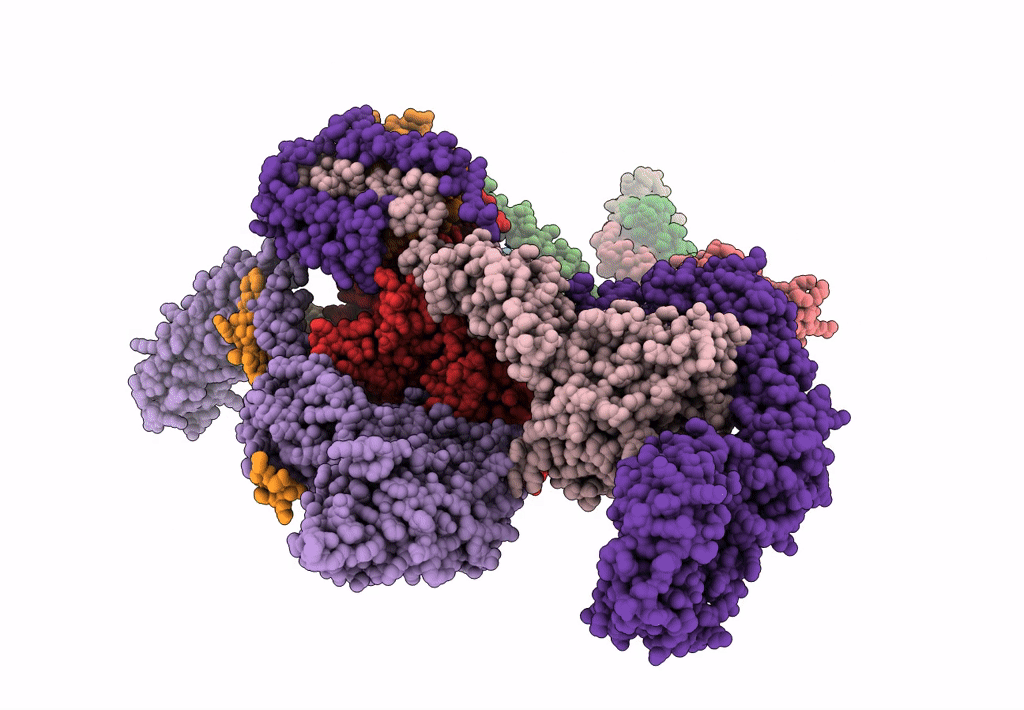
Subtomogram average of the human Sec61-TRAP-OSTA-translocon
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
Resolution:
7.60 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SUBTOMOGRAM AVERAGING