Deposition Date
2022-07-04
Release Date
2023-10-25
Last Version Date
2026-03-04
Entry Detail
PDB ID:
8AB5
Keywords:
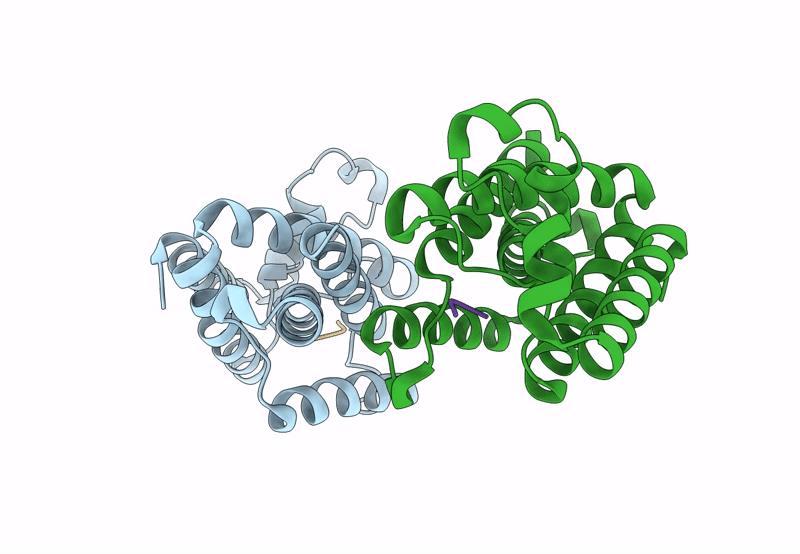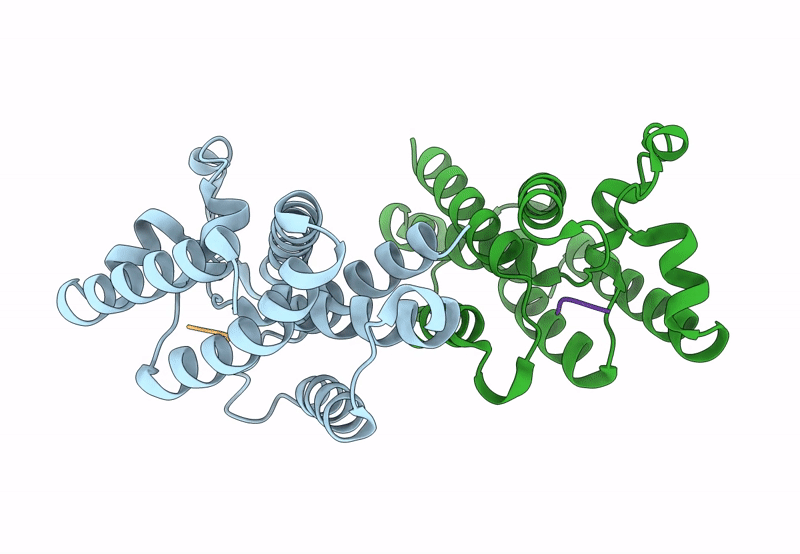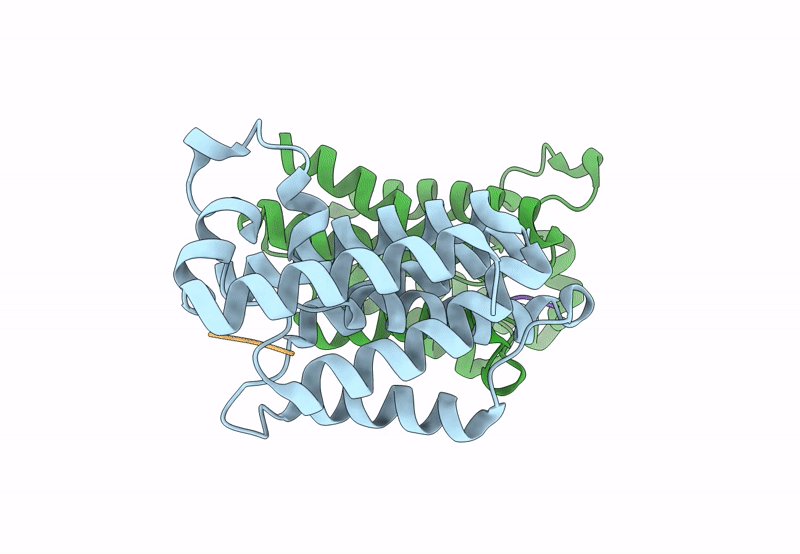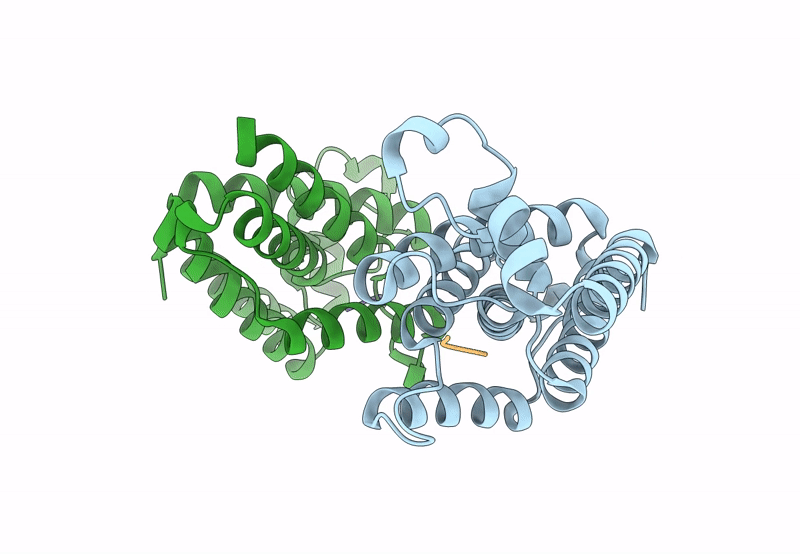
Title:
Structure of E. coli GlpG in complex with peptide derived inhibitor Ac-VRHA-conh-[4-(4-butyl)-phenoxy-1-phenyl-2-butyl]
Biological Source:
Source Organism(s):
Escherichia coli K-12 (Taxon ID: 83333)
Providencia stuartii (Taxon ID: 588)
Providencia stuartii (Taxon ID: 588)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.40 Å
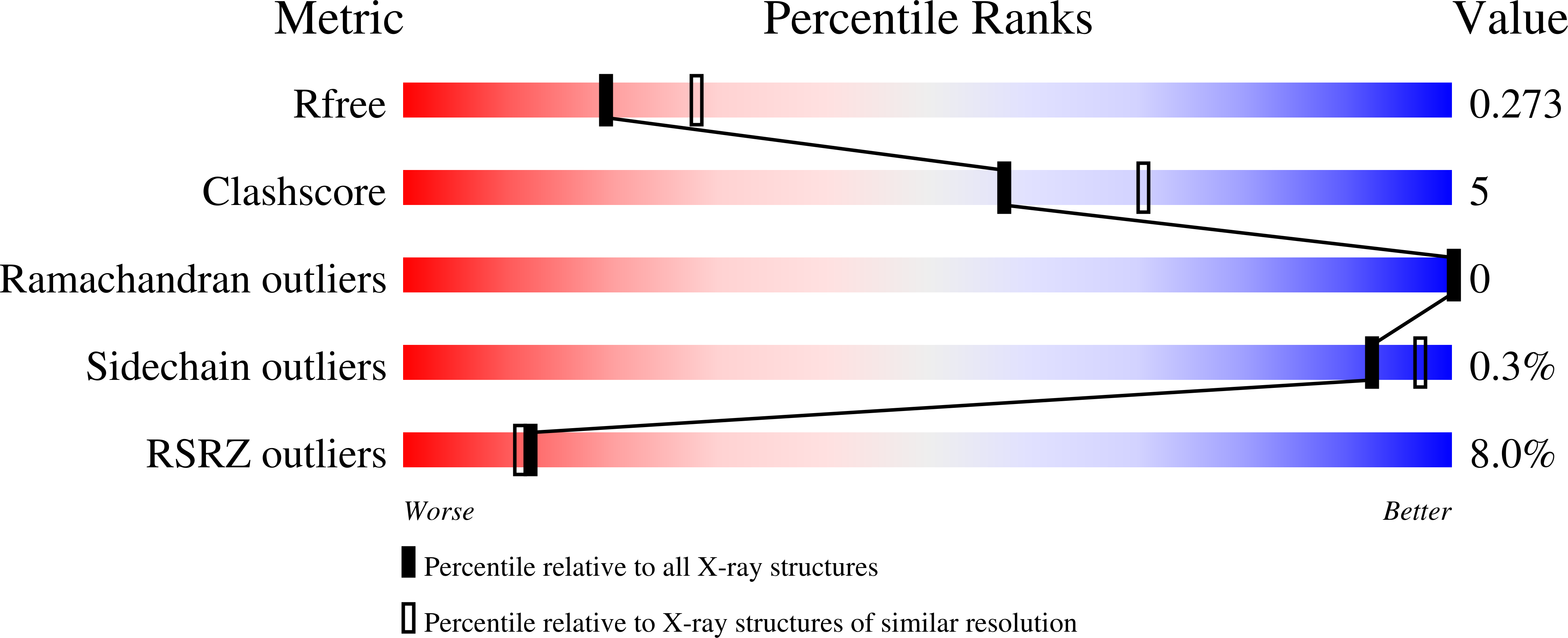
R-Value Free:
0.27
R-Value Work:
0.21
R-Value Observed:
0.22
Space Group:
P 1 21 1