Deposition Date
2019-01-17
Release Date
2020-08-05
Last Version Date
2024-05-15
Entry Detail
PDB ID:
6QI5
Keywords:
Title:
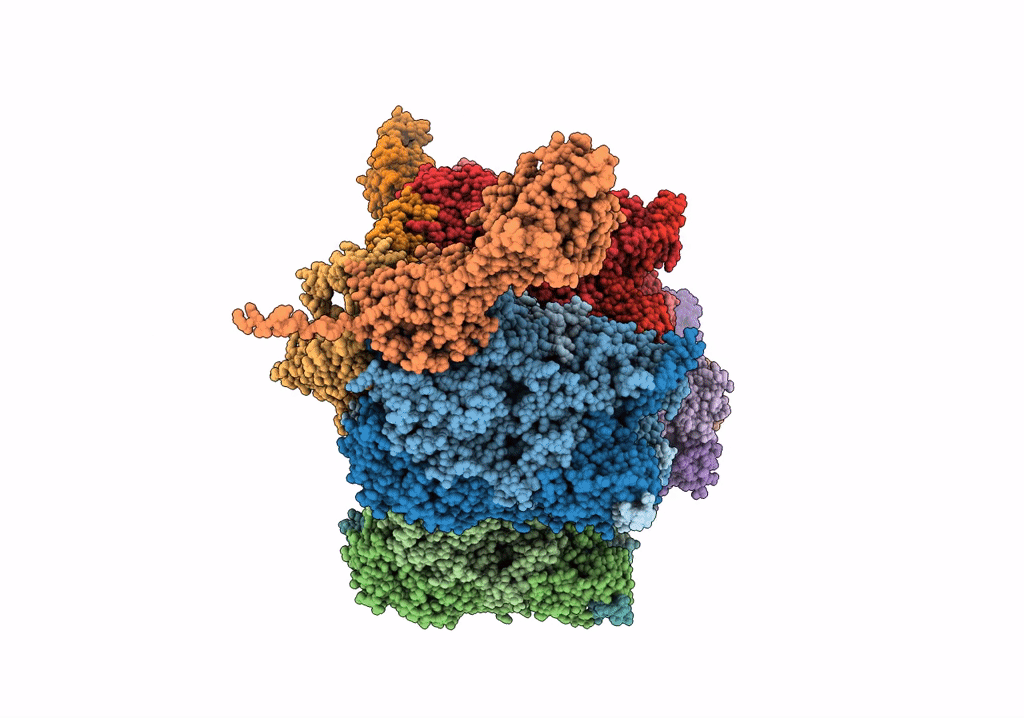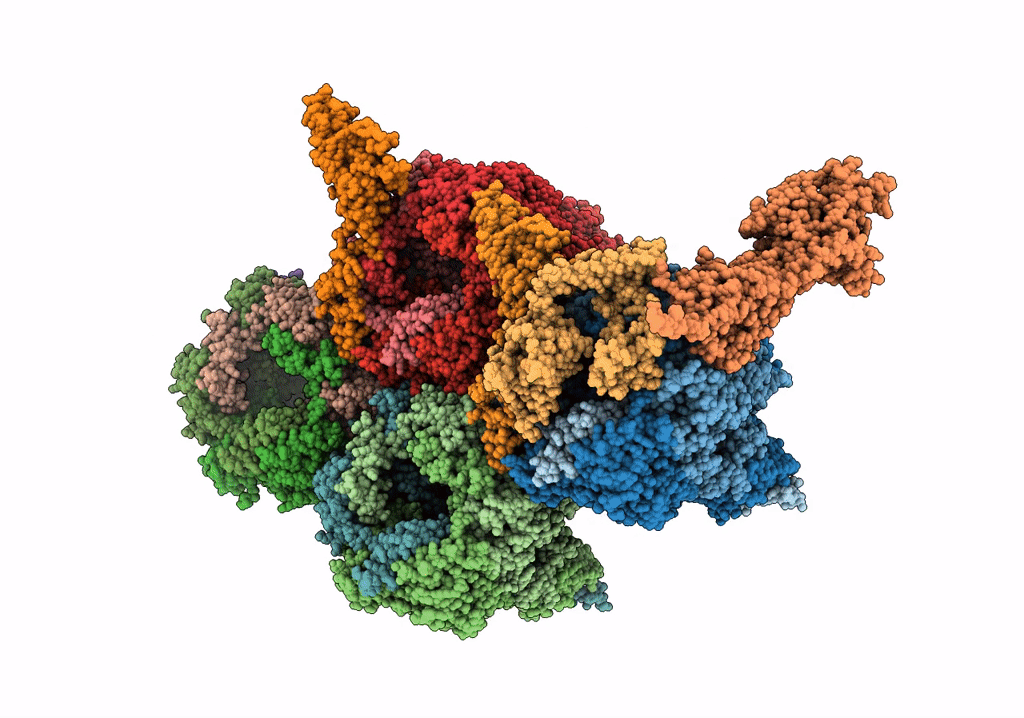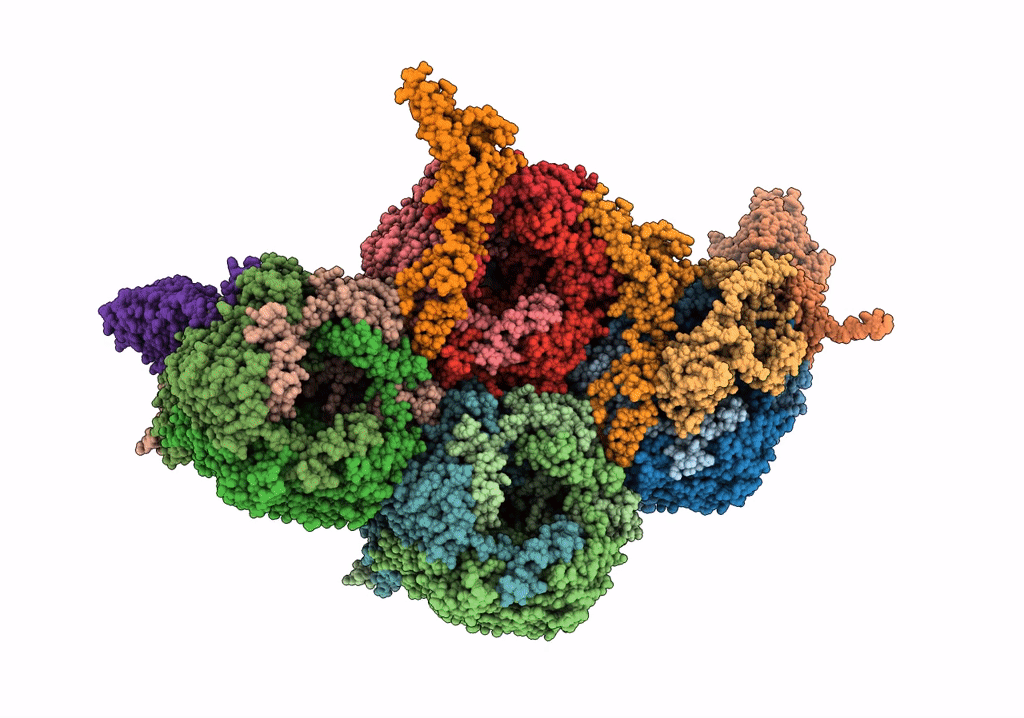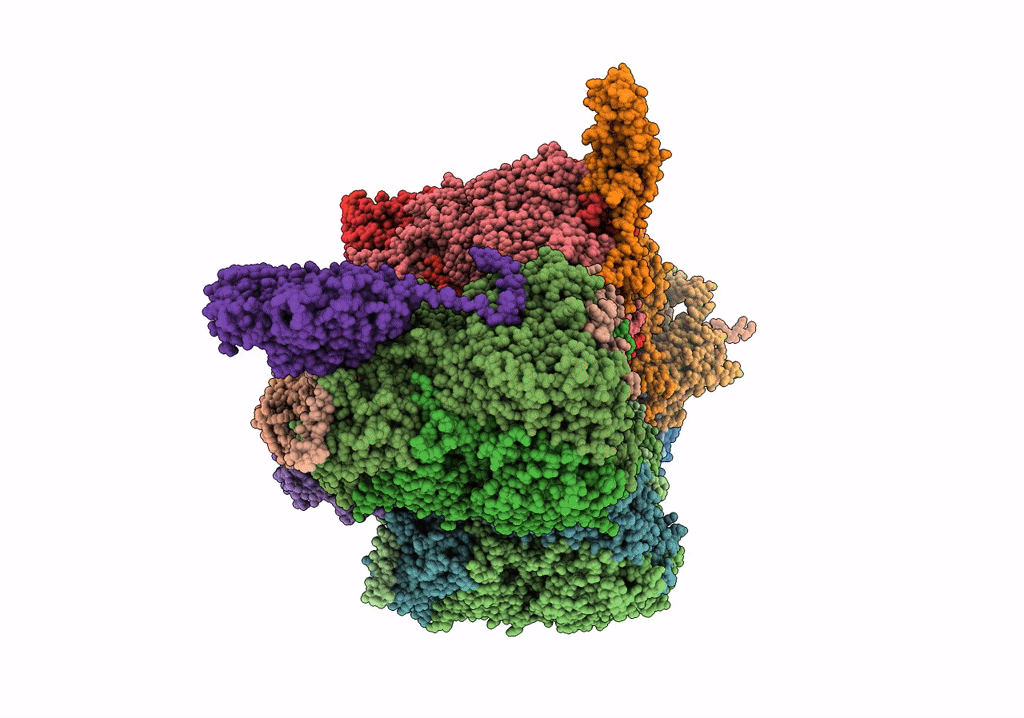
Near Atomic Structure of an Atadenovirus Shows a possible gene duplication event and Intergenera Variations in Cementing Proteins
Biological Source:
Source Organism(s):
Lizard adenovirus 2 (Taxon ID: 874272)
Method Details:
Experimental Method:
Resolution:
3.40 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE