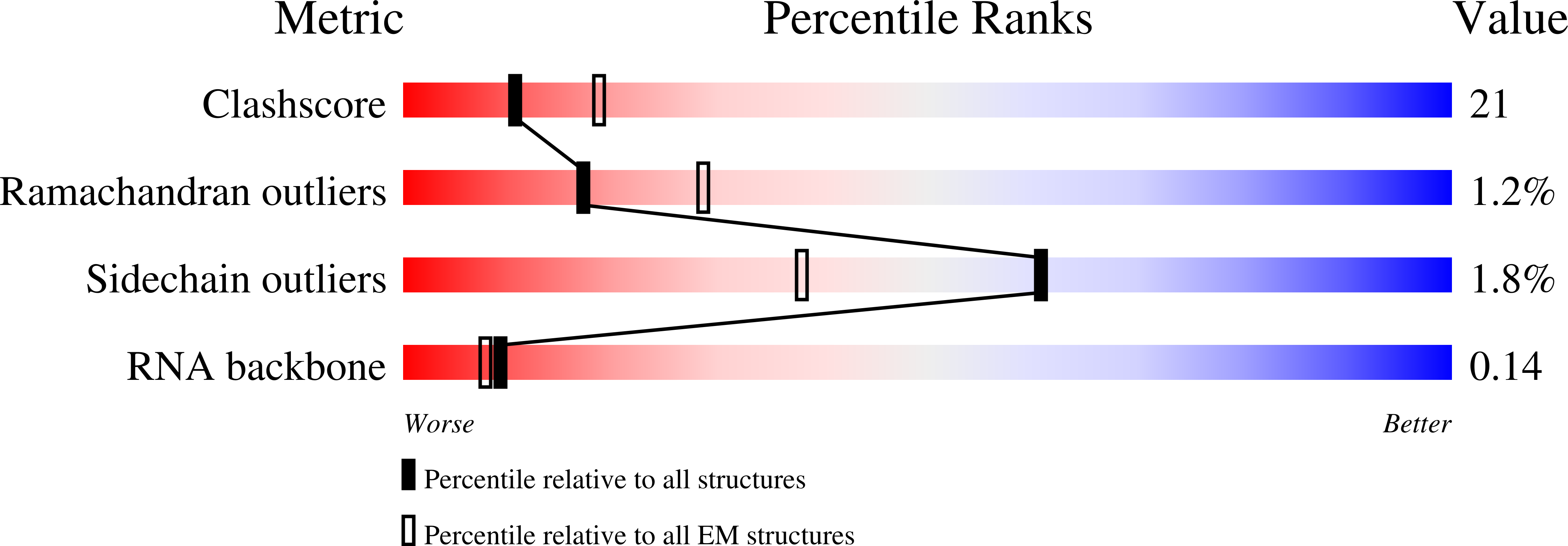
Deposition Date
2018-01-17
Release Date
2018-06-27
Last Version Date
2023-11-29
Entry Detail
PDB ID:
6C66
Keywords:
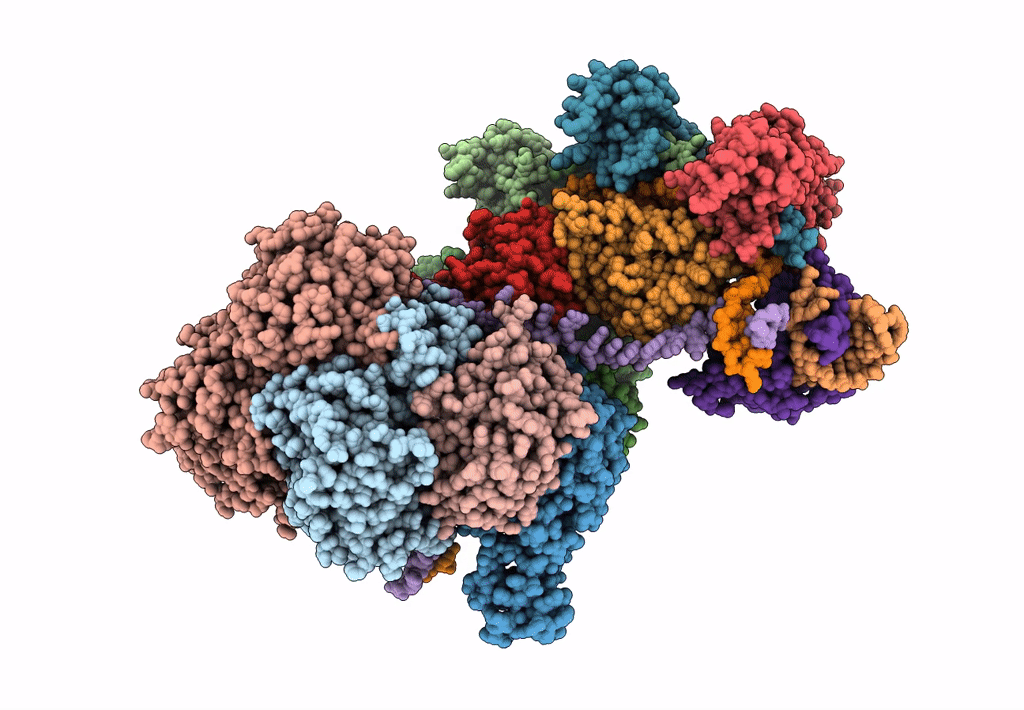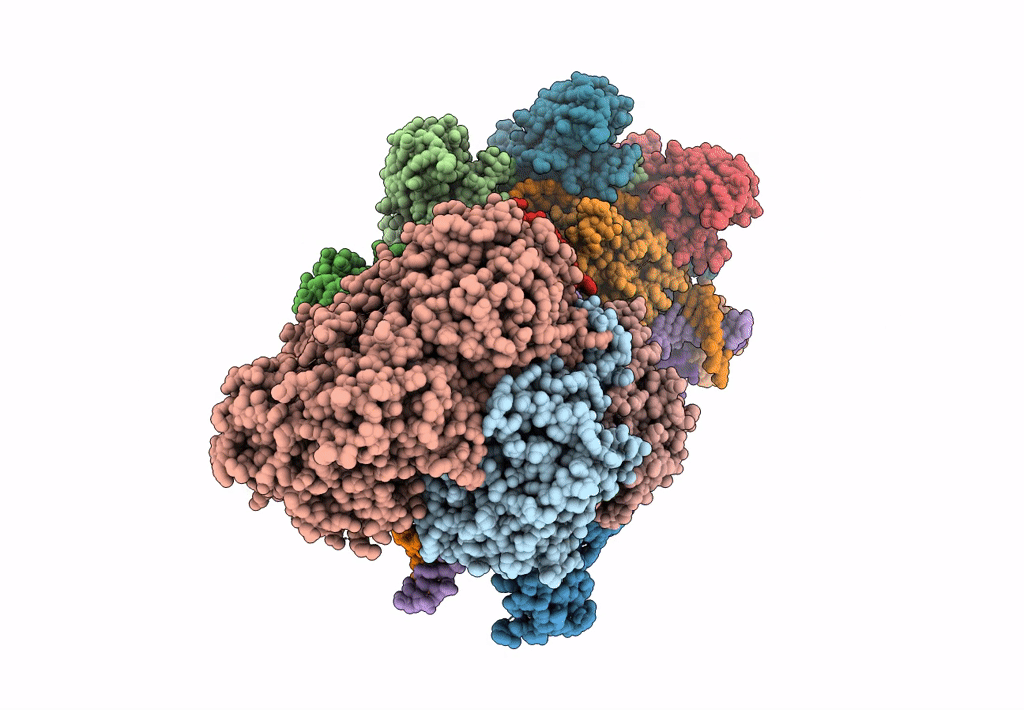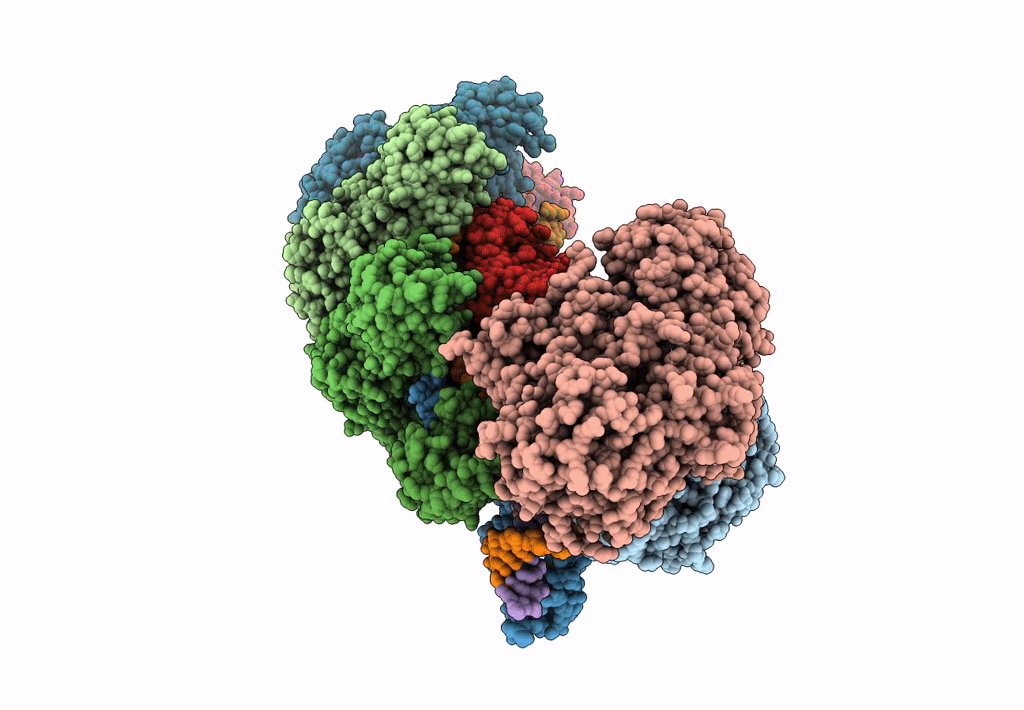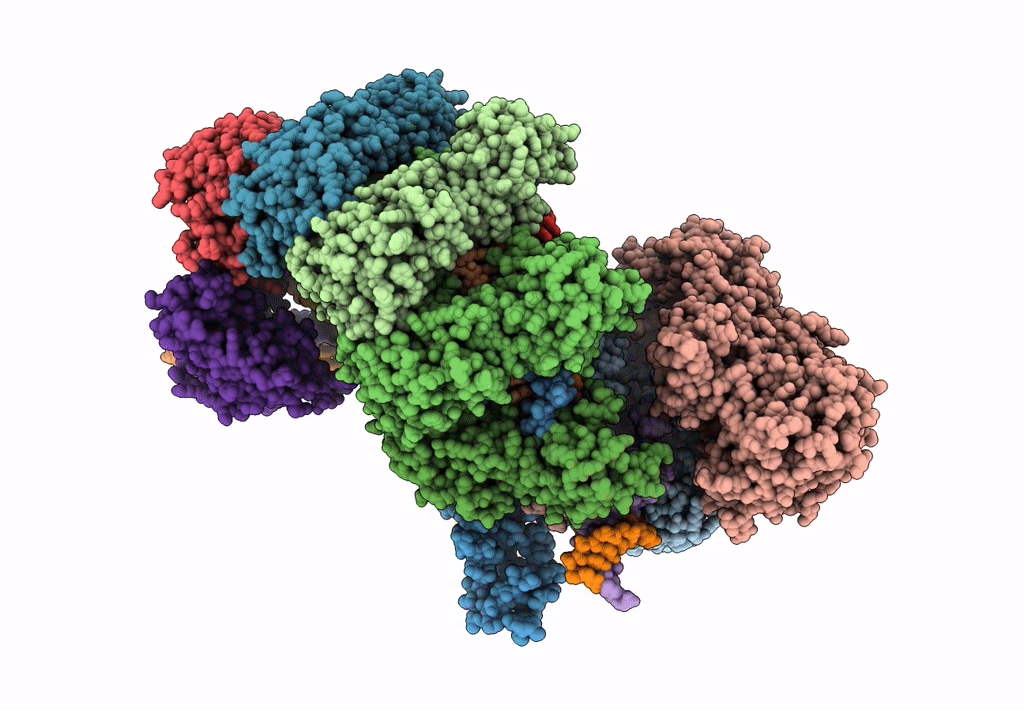
Title:
CRISPR RNA-guided surveillance complex, pre-nicking
Biological Source:
Source Organism(s):
Thermobifida fusca (strain YX) (Taxon ID: 269800)
Thermobifida fusca (Taxon ID: 2021)
Thermobifida fusca (Taxon ID: 2021)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
3.66 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE