Deposition Date
2018-08-01
Release Date
2018-11-14
Last Version Date
2025-09-17
Entry Detail
PDB ID:
6ADQ
Keywords:
Title:
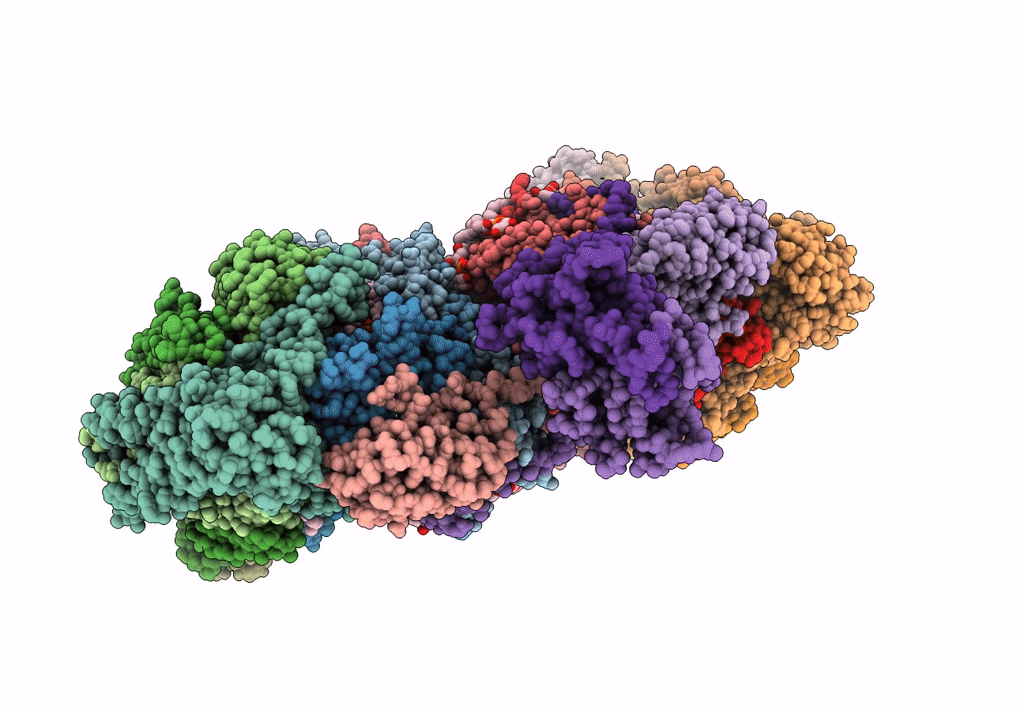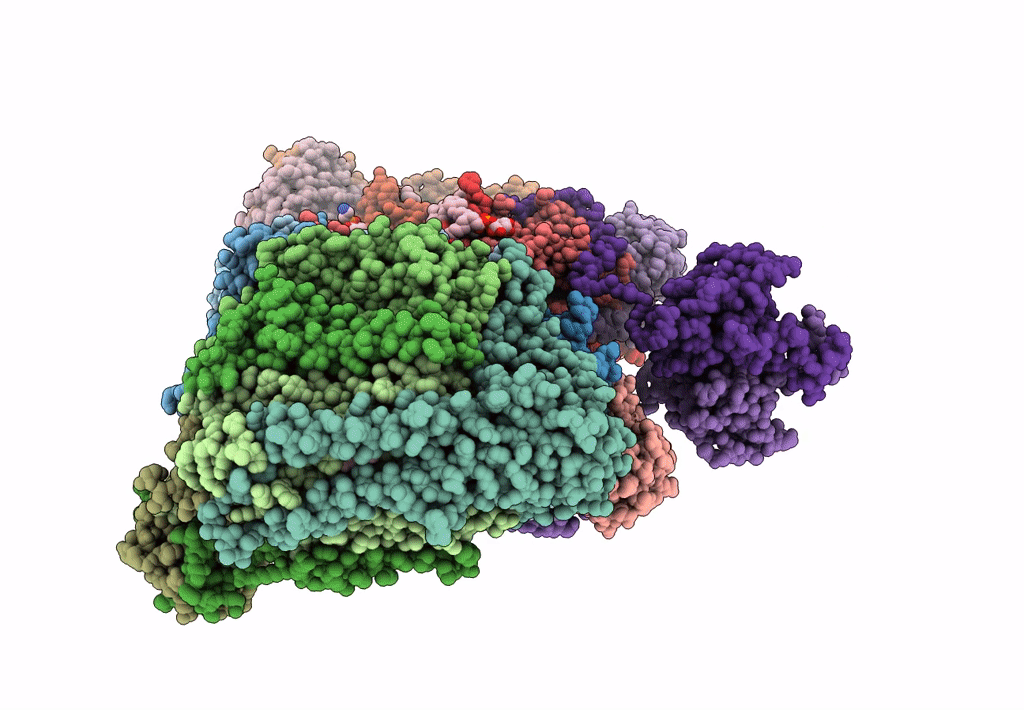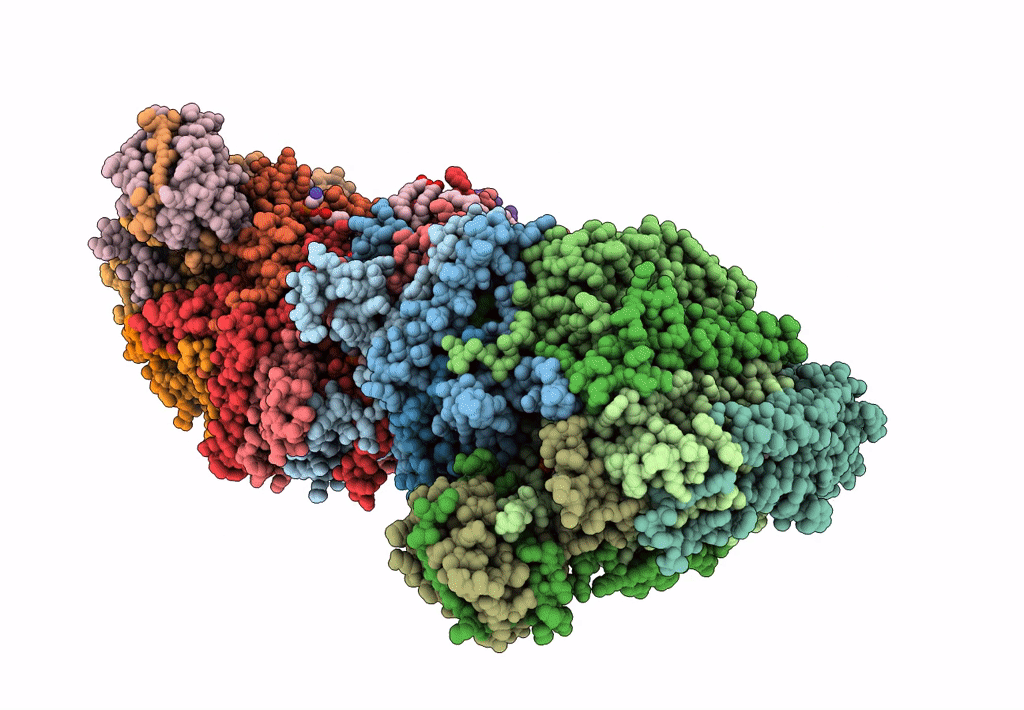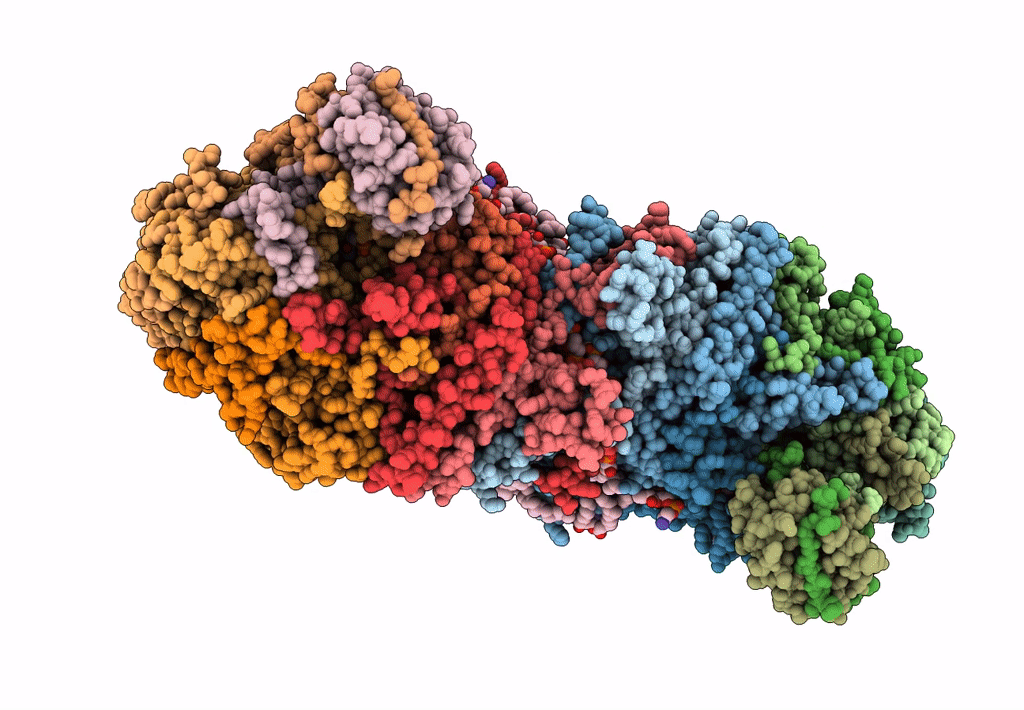
Respiratory Complex CIII2CIV2SOD2 from Mycobacterium smegmatis
Biological Source:
Source Organism(s):
Mycobacterium smegmatis MC2 51 (Taxon ID: 1445611)
Method Details:
Experimental Method:
Resolution:
3.50 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE