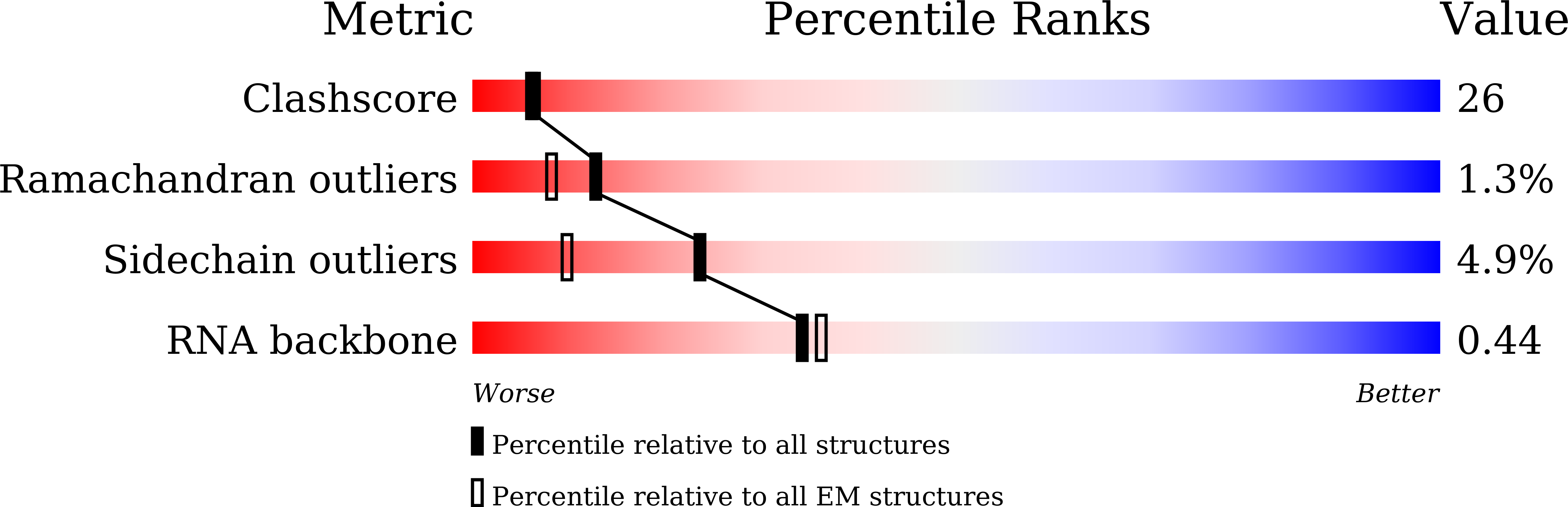
Deposition Date
2018-01-17
Release Date
2018-09-19
Last Version Date
2024-11-13
Entry Detail
PDB ID:
5Z56
Keywords:
Title:
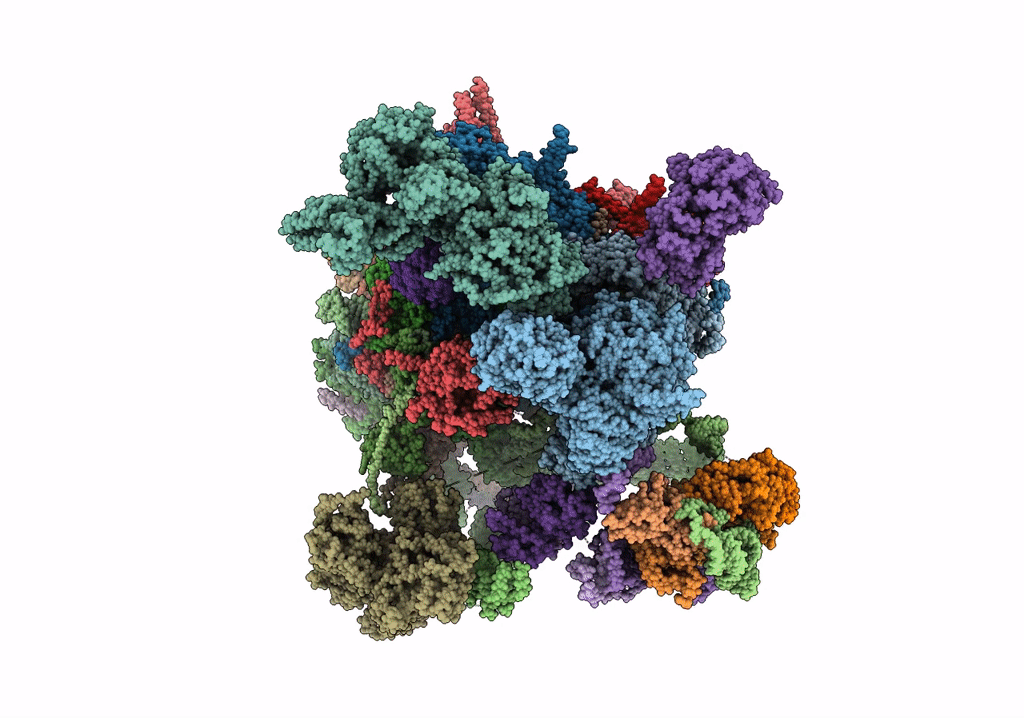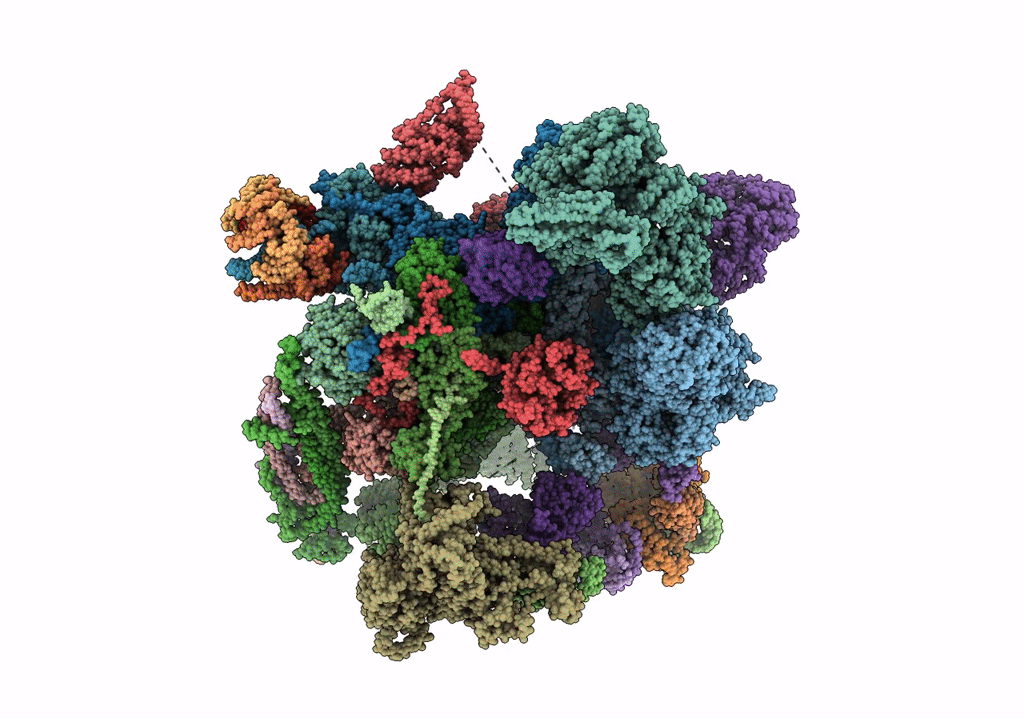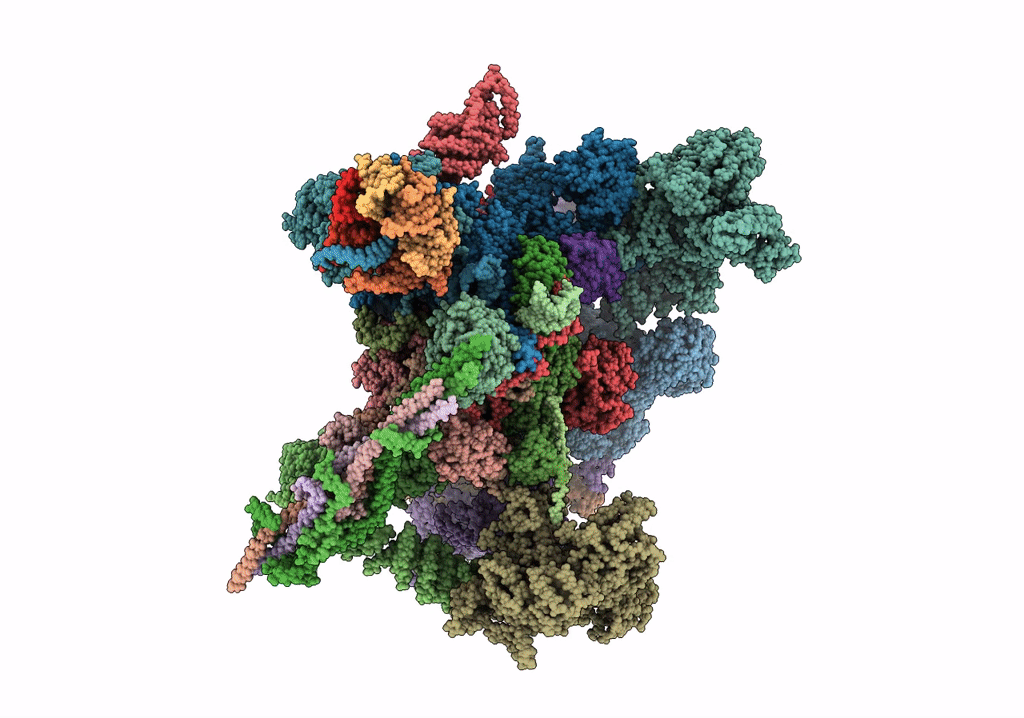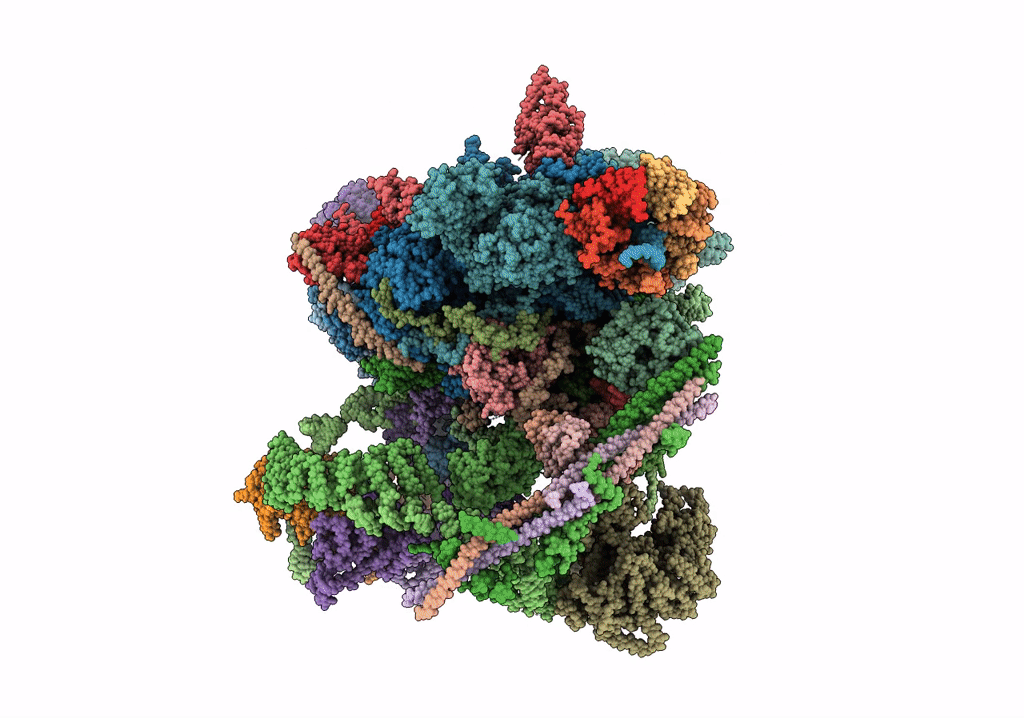
cryo-EM structure of a human activated spliceosome (mature Bact) at 5.1 angstrom.
Biological Source:
Source Organism(s):
unidentified adenovirus (Taxon ID: 10535)
Homo sapiens (Taxon ID: 9606)
Homo sapiens (Taxon ID: 9606)
Method Details:
Experimental Method:
Resolution:
5.10 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE