Deposition Date
2017-06-12
Release Date
2017-11-15
Last Version Date
2023-11-15
Entry Detail
PDB ID:
5W4L
Keywords:
Title:
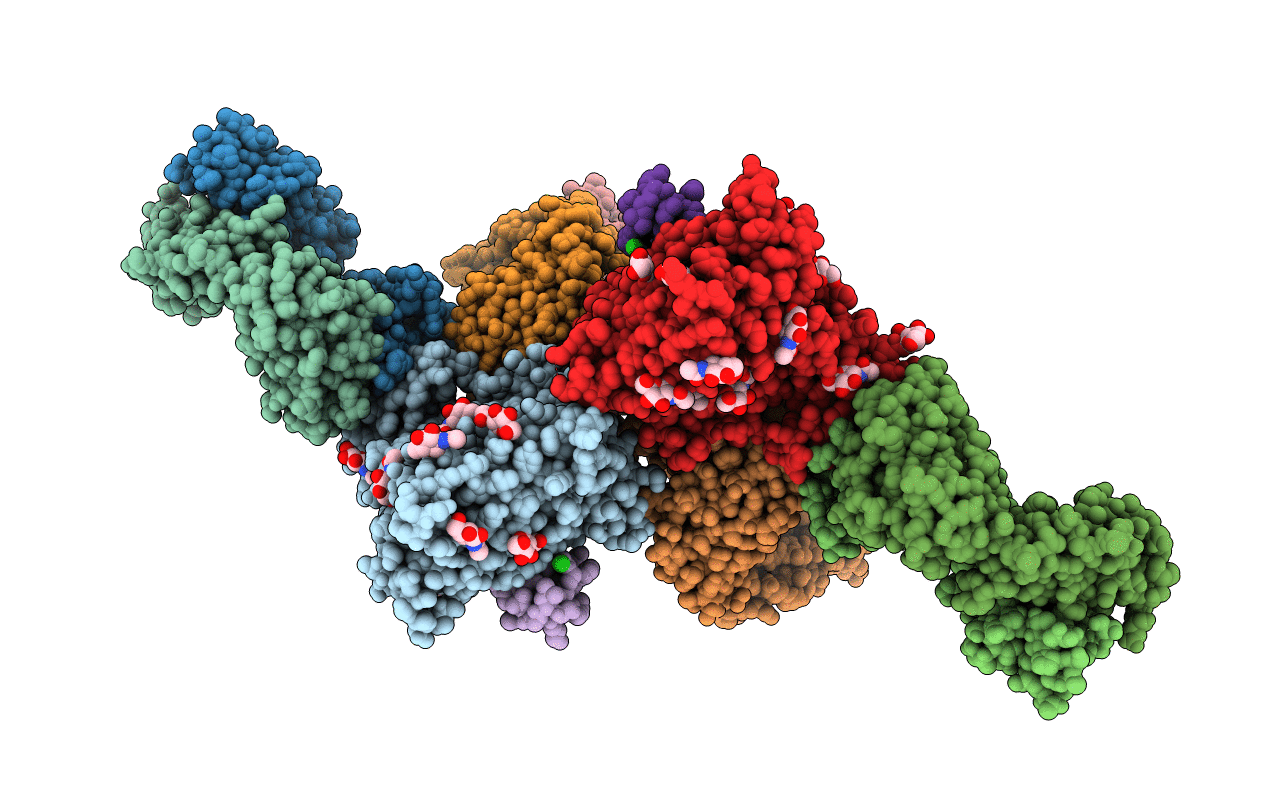
Crystal structure of the non-neutralizing and ADCC-potent C11-like antibody N12-i3 in complex with HIV-1 clade A/E gp120, the CD4 mimetic M48U1, and the antibody N5-i5.
Biological Source:
Source Organism(s):
Human immunodeficiency virus 1 (Taxon ID: 11676)
Homo sapiens (Taxon ID: 9606)
synthetic construct (Taxon ID: 32630)
Homo sapiens (Taxon ID: 9606)
synthetic construct (Taxon ID: 32630)
Expression System(s):
Method Details:
Experimental Method:
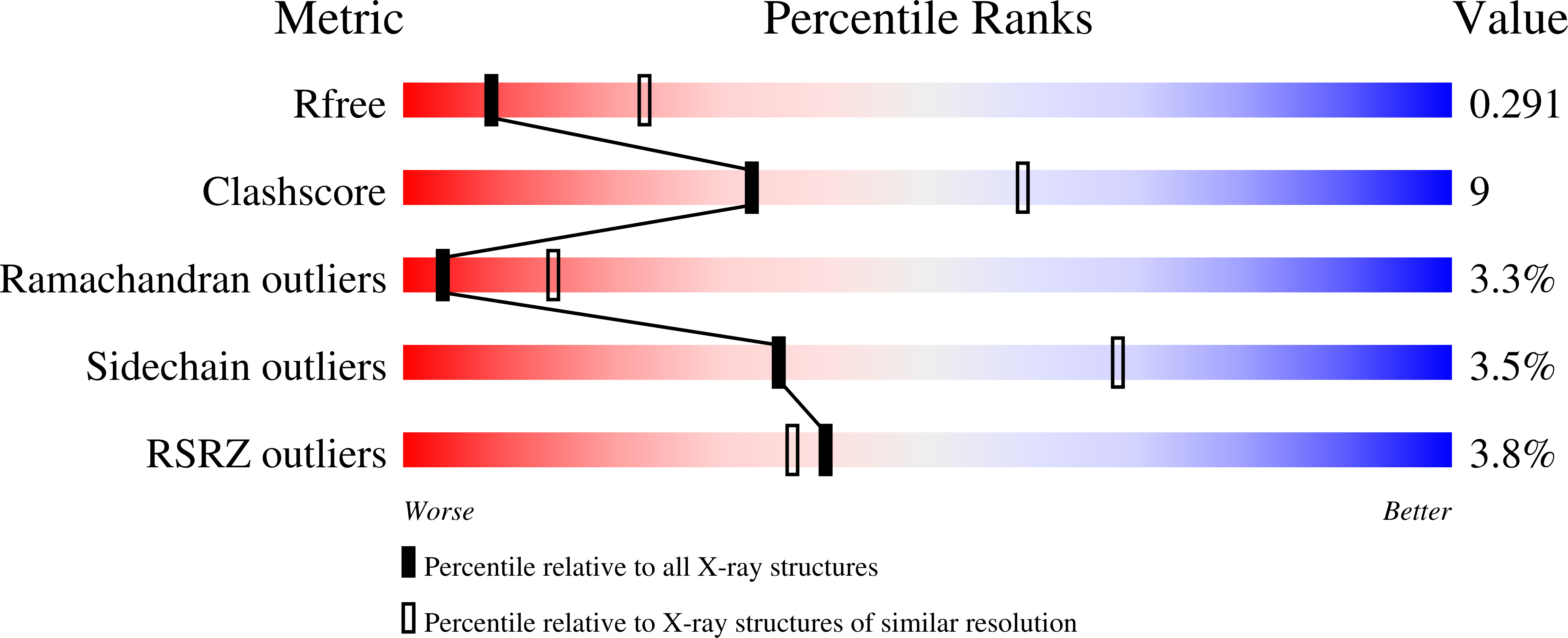
Resolution:
2.92 Å
R-Value Free:
0.29
R-Value Work:
0.23
R-Value Observed:
0.23
Space Group:
C 1 2 1