Deposition Date
2007-11-22
Release Date
2014-07-09
Last Version Date
2024-11-13
Method Details:
Experimental Method:
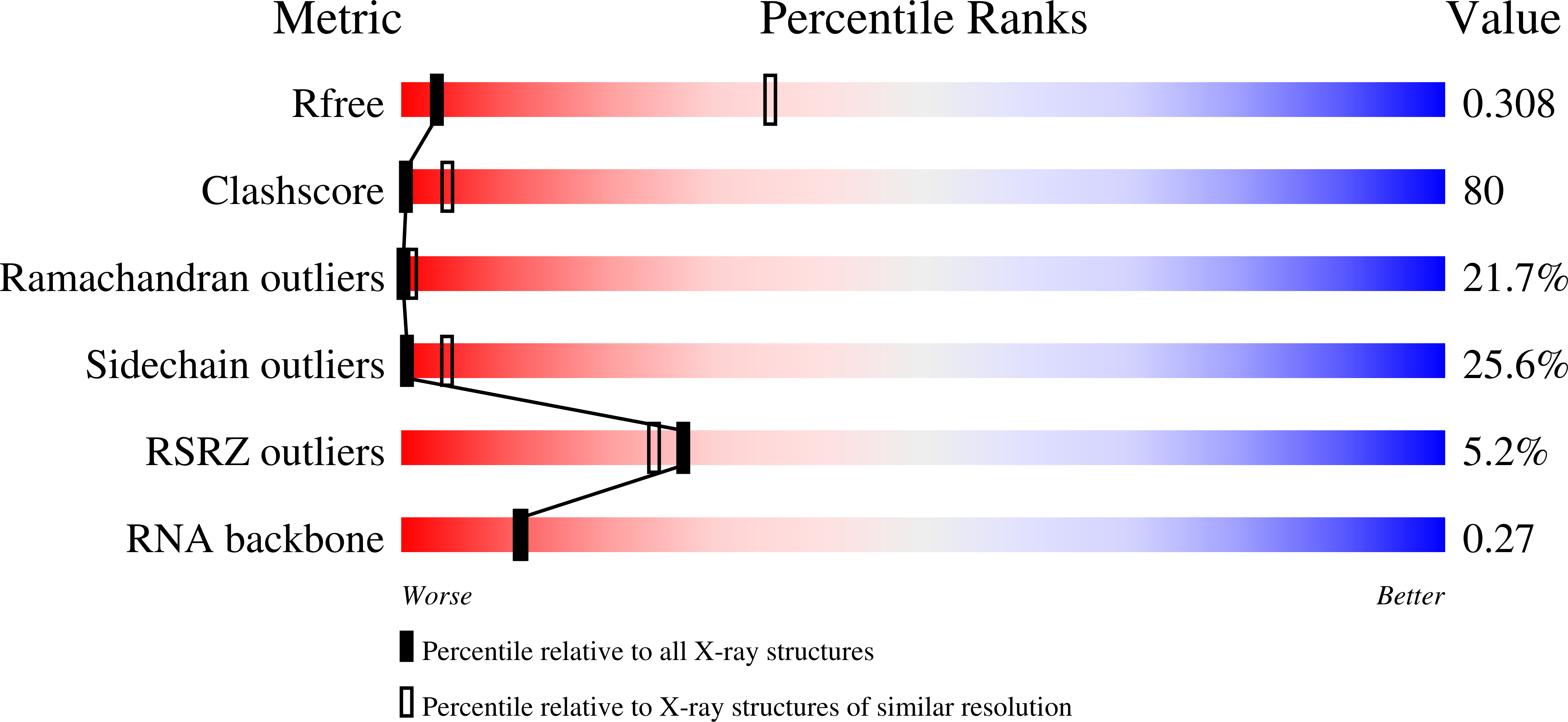
Resolution:
3.74 Å
R-Value Free:
0.32
R-Value Work:
0.25
Space Group:
P 21 21 21