Deposition Date
2013-09-28
Release Date
2014-01-01
Last Version Date
2026-03-25
Entry Detail
PDB ID:
4MZ1
Keywords:
Title:
Crystal Structure of the Inosine 5'-monophosphate Dehydrogenase, with a Internal Deletion of CBS Domain from Campylobacter jejuni complexed with inhibitor compound P12
Biological Source:
Source Organism(s):
Campylobacter jejuni subsp. jejuni (Taxon ID: 192222)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.40 Å
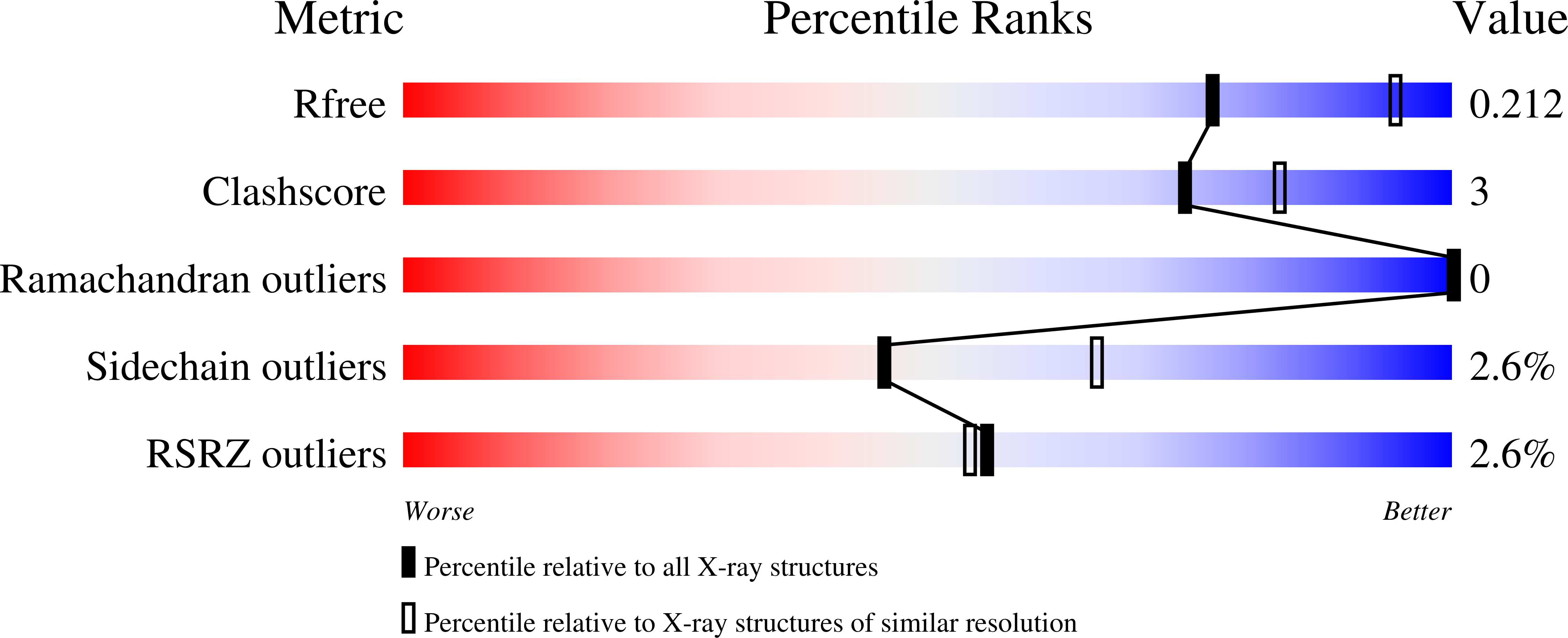
R-Value Free:
0.21
R-Value Work:
0.17
R-Value Observed:
0.17
Space Group:
I 4 2 2


