Deposition Date
2013-03-23
Release Date
2013-06-26
Last Version Date
2023-09-20
Entry Detail
PDB ID:
4JTI
Keywords:
Title:
Crystal structure of F114R/R117Q/F139G mutant of 3-deoxy-D-manno-octulosonate 8-phosphate synthase (KDO8PS) from Neisseria meningitidis
Biological Source:
Source Organism(s):
Neisseria meningitidis (Taxon ID: 122586)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.75 Å
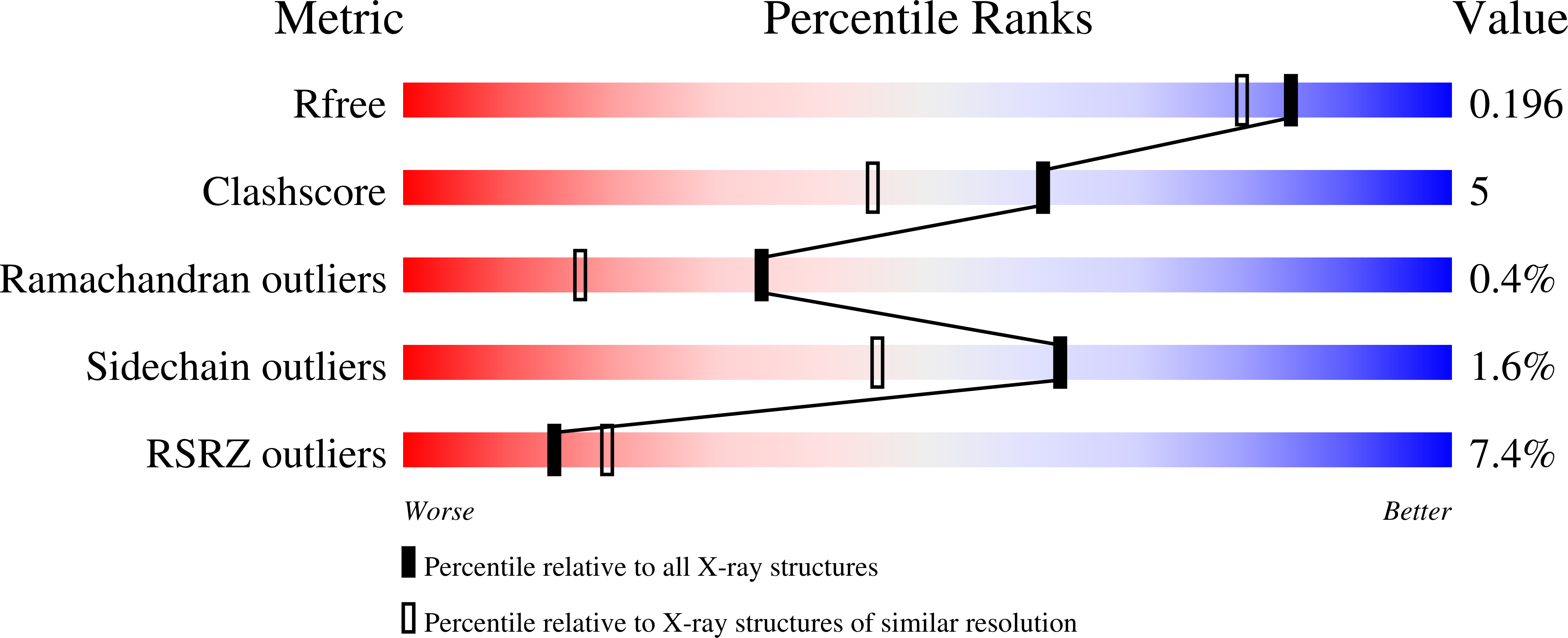
R-Value Free:
0.18
R-Value Work:
0.15
R-Value Observed:
0.16
Space Group:
P 21 21 21


