Deposition Date
2012-12-20
Release Date
2013-11-06
Last Version Date
2023-09-20
Entry Detail
PDB ID:
4IIT
Keywords:
Title:
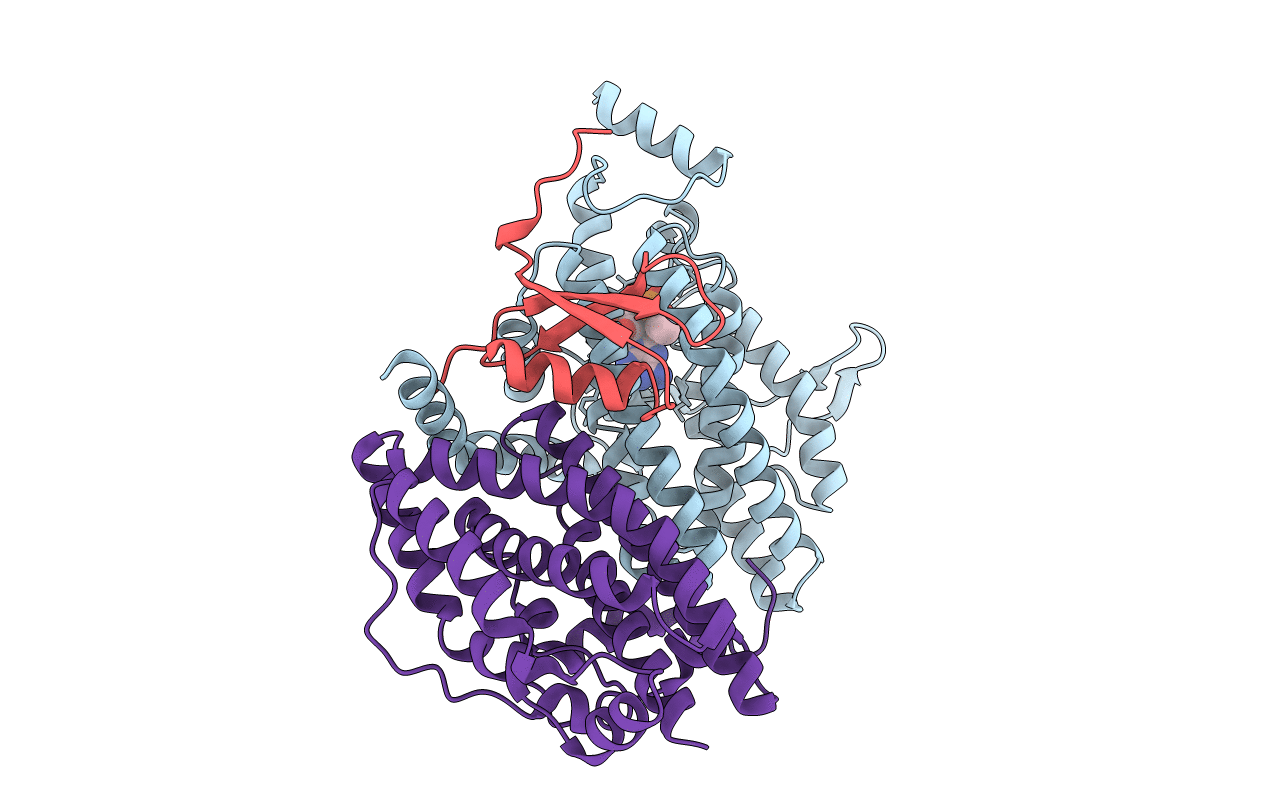
The Phenylacetyl-CoA monooxygenase PaaABC subcomplex with phenylacetyl-CoA
Biological Source:
Source Organism(s):
Klebsiella pneumoniae subsp. pneumoniae (Taxon ID: 272620)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
4.30 Å
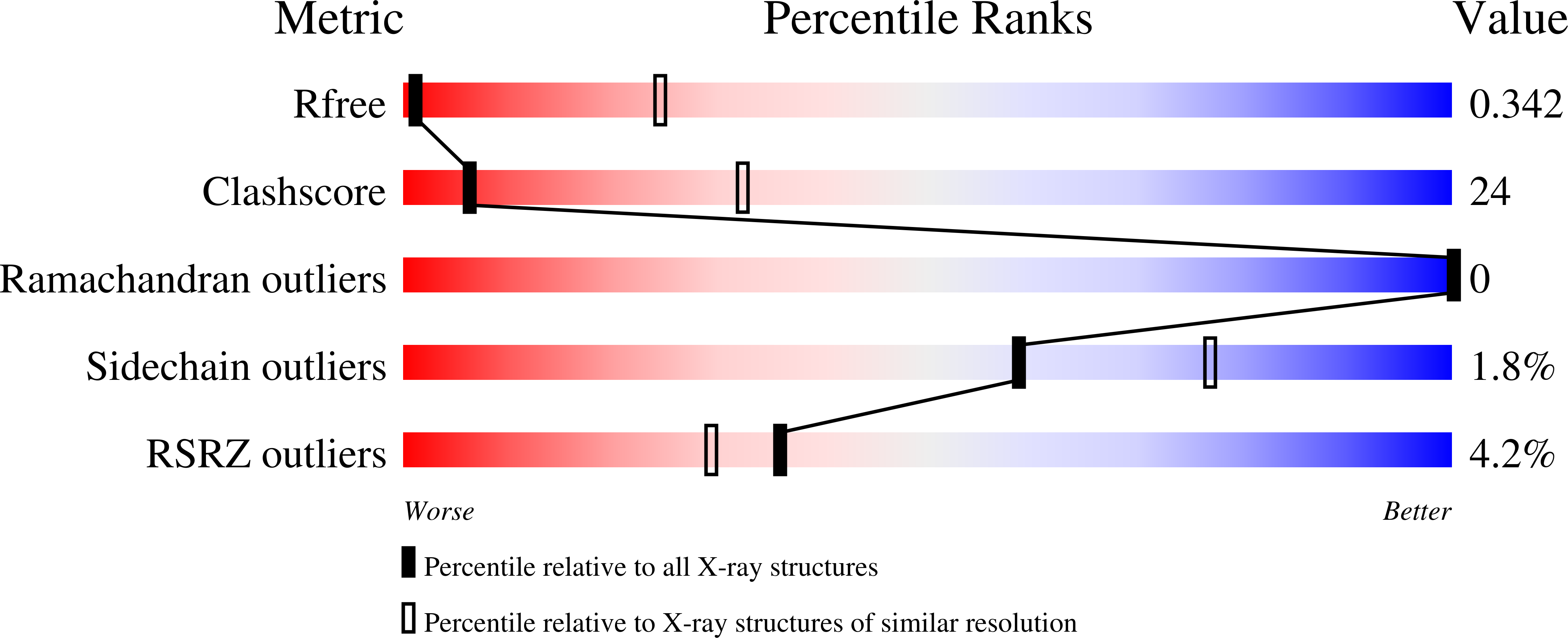
R-Value Free:
0.36
R-Value Work:
0.28
R-Value Observed:
0.29
Space Group:
P 63 2 2


