Deposition Date
2011-05-13
Release Date
2011-08-31
Last Version Date
2024-02-28
Entry Detail
PDB ID:
3S0Q
Keywords:
Title:
Peptidase module of the peptidoglycan hydrolase RipA (Rv1477) from Mycobacterium tuberculosis, catalytic site mutant (Cys383Ala) at 1.45 resolution
Biological Source:
Source Organism(s):
Mycobacterium tuberculosis (Taxon ID: 1773)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
1.45 Å
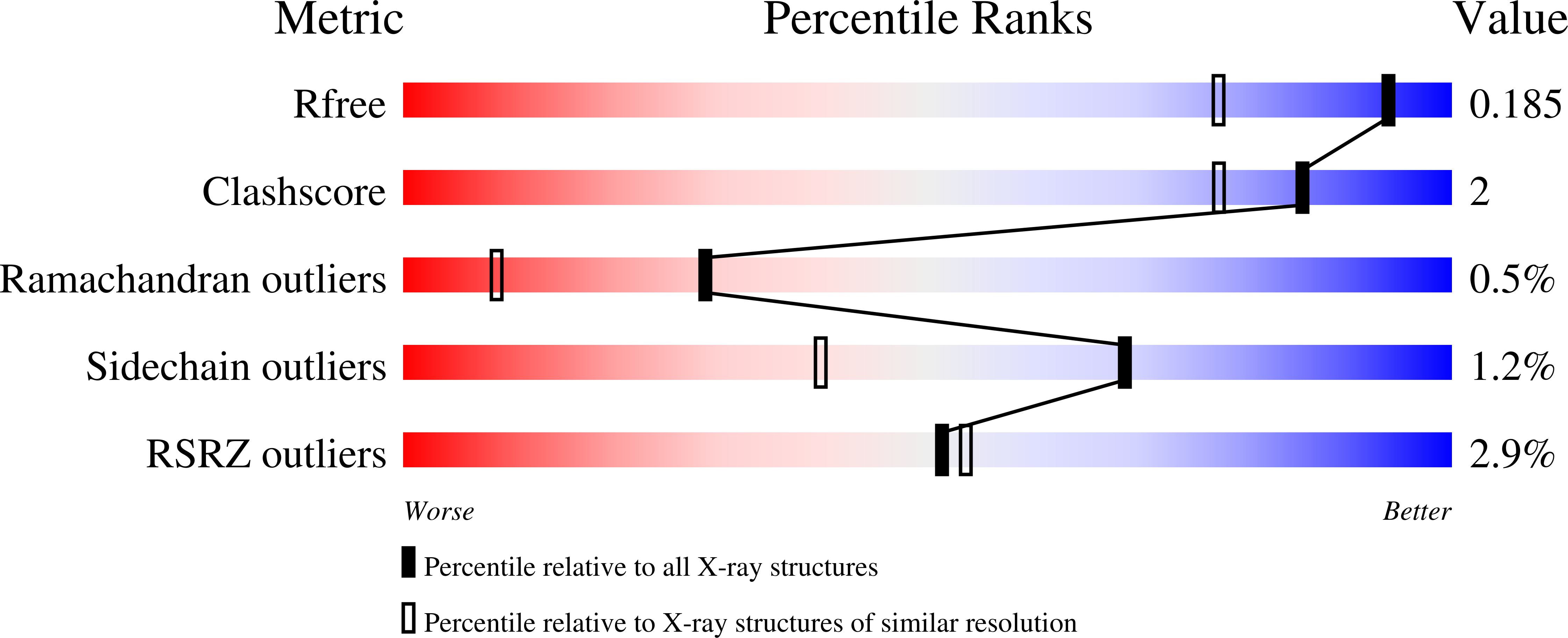
R-Value Free:
0.17
R-Value Work:
0.15
R-Value Observed:
0.15
Space Group:
P 21 21 21


