Deposition Date
2013-04-18
Release Date
2013-05-15
Last Version Date
2024-03-20
Entry Detail
PDB ID:
3J3T
Keywords:
Title:
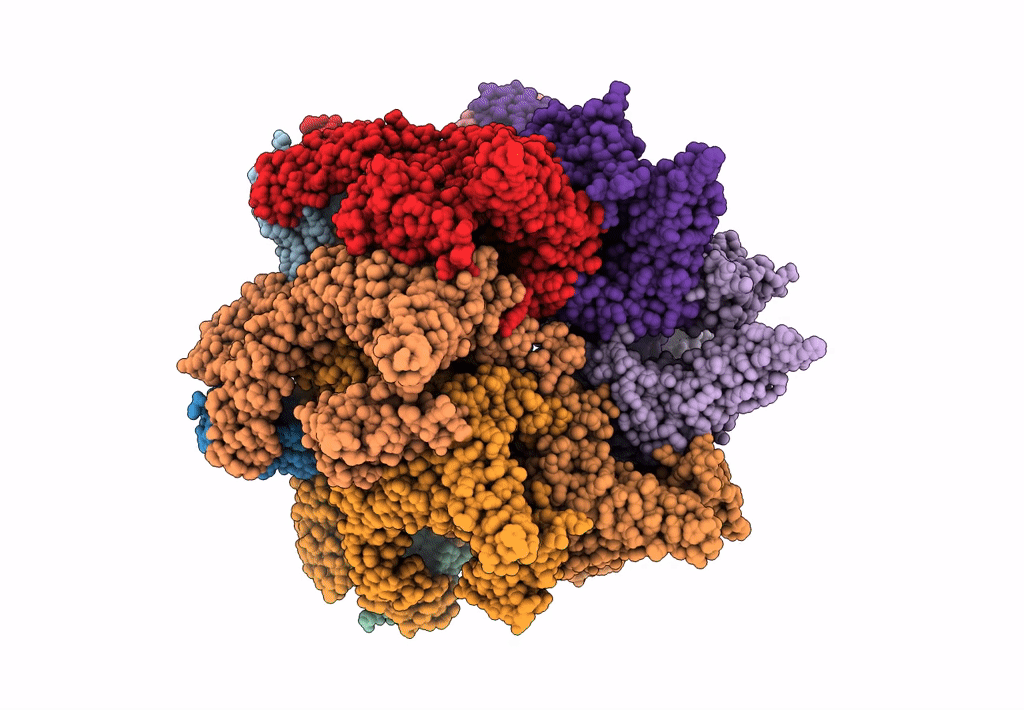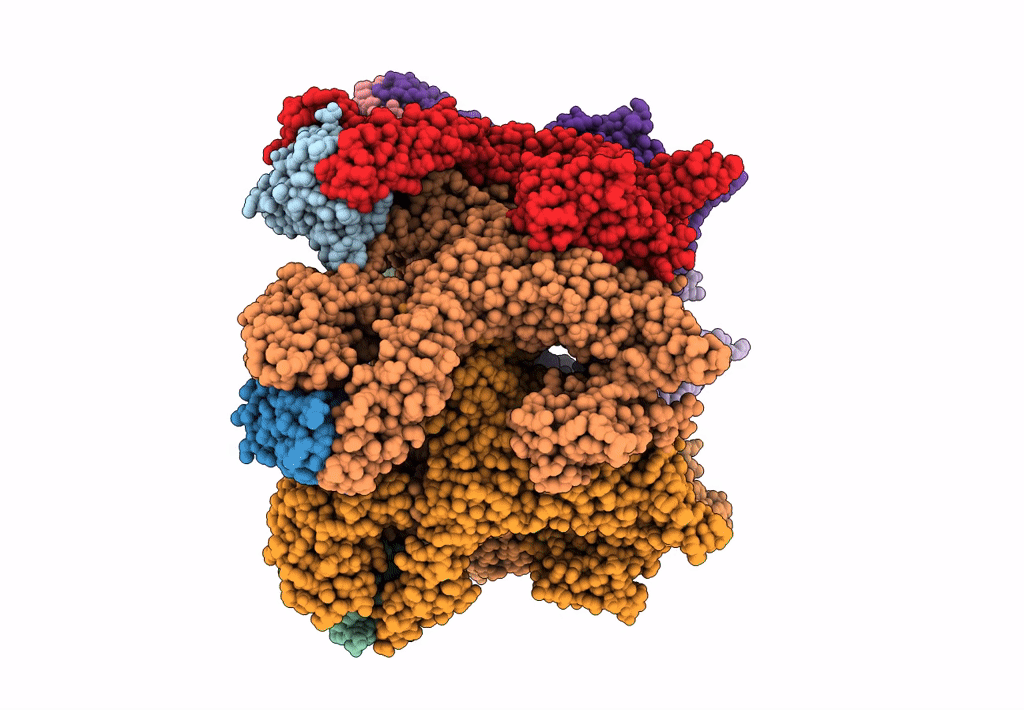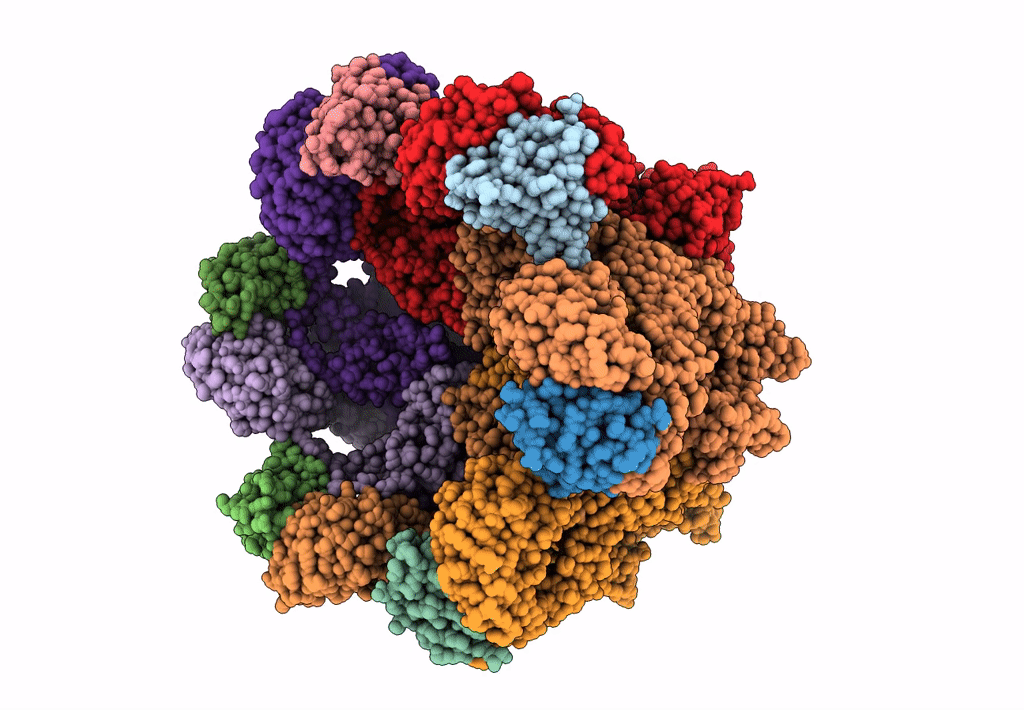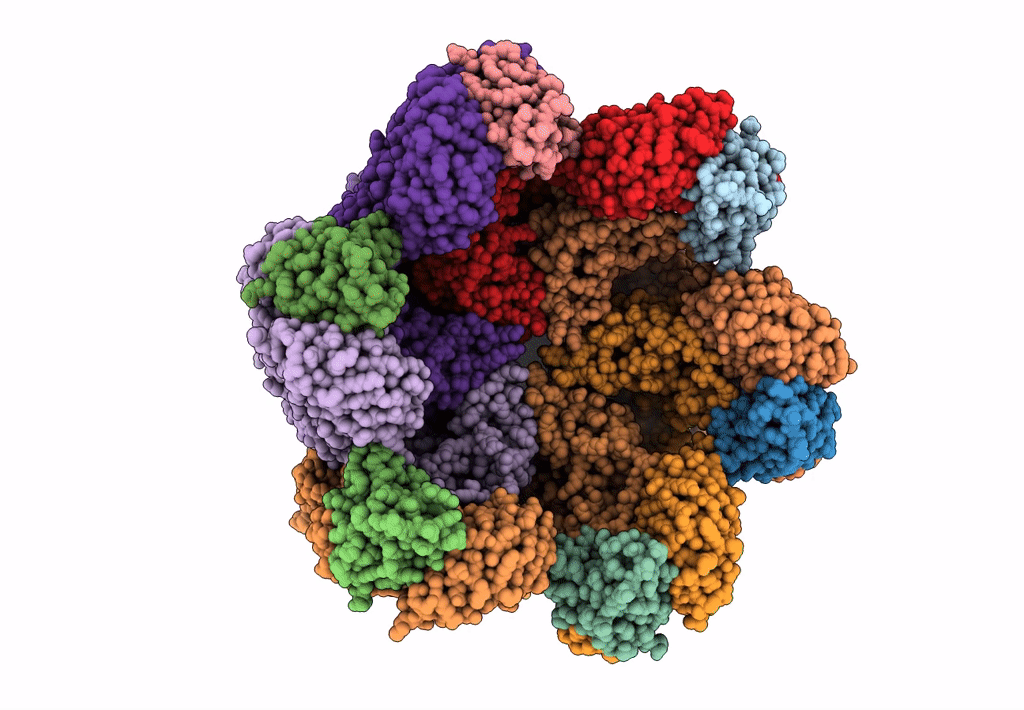
Structural dynamics of the MecA-ClpC complex revealed by cryo-EM
Biological Source:
Source Organism(s):
Bacillus subtilis (Taxon ID: 224308)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
9.00 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE