Deposition Date
2005-06-10
Release Date
2005-11-22
Last Version Date
2024-02-14
Entry Detail
PDB ID:
1ZYQ
Keywords:
Title:
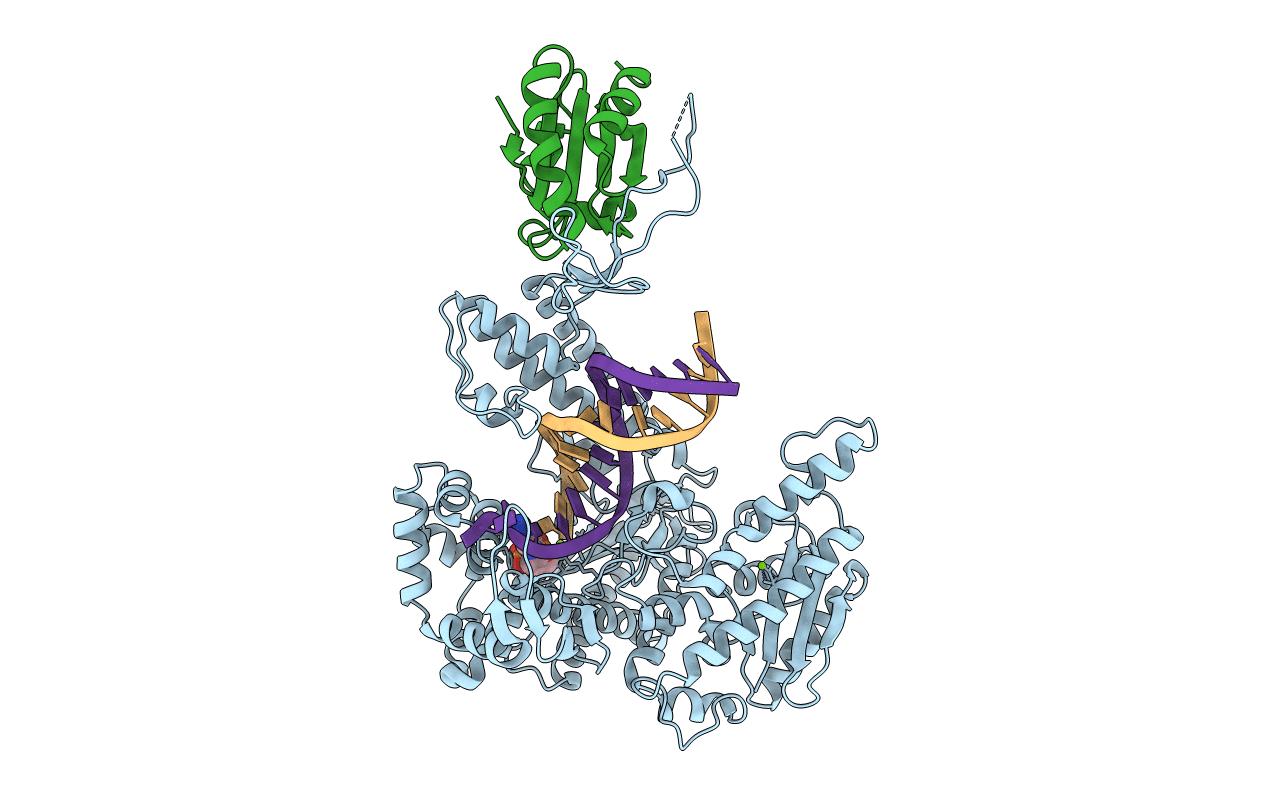
T7 DNA polymerase in complex with 8oG and incoming ddATP
Biological Source:
Source Organism(s):
Enterobacteria phage T7 (Taxon ID: 10760)
Escherichia coli (Taxon ID: 562)
Escherichia coli (Taxon ID: 562)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.70 Å
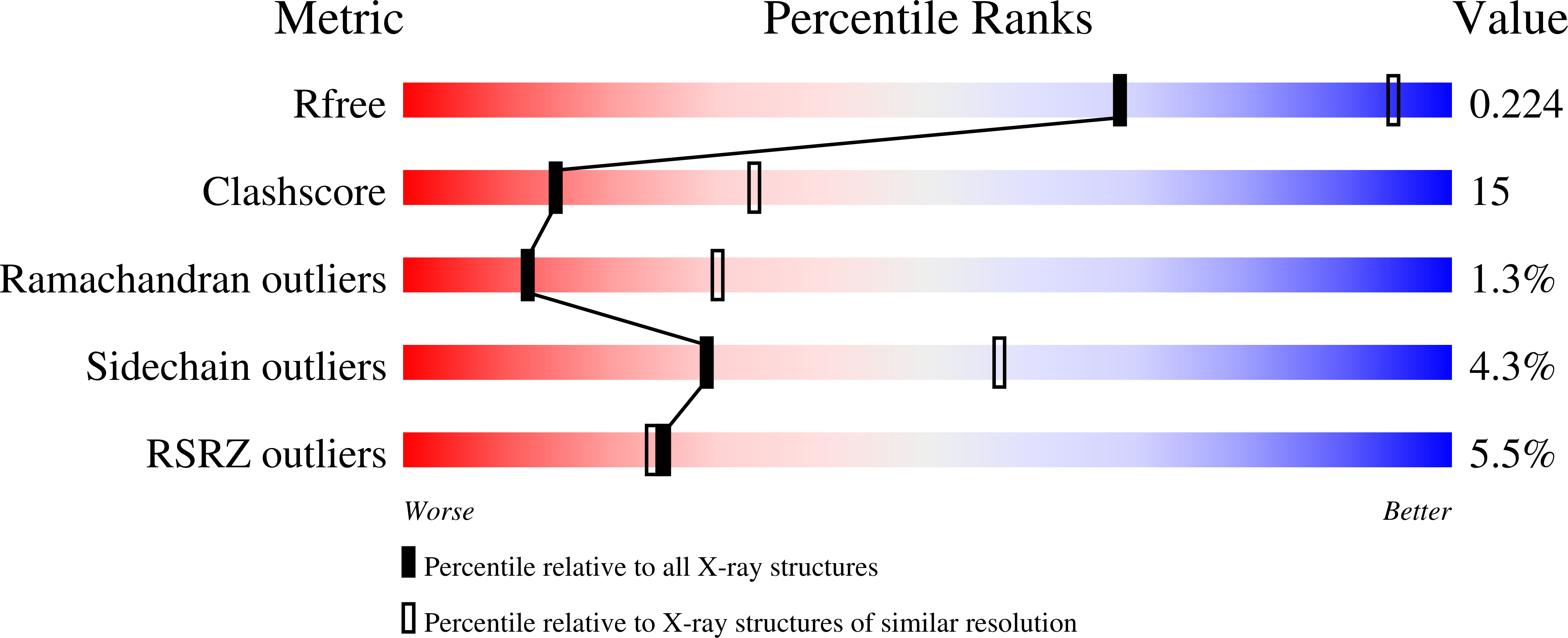
R-Value Free:
0.27
R-Value Work:
0.22
R-Value Observed:
0.22
Space Group:
P 21 21 2


