Deposition Date
2001-06-04
Release Date
2001-11-28
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1JBG
Keywords:
Title:
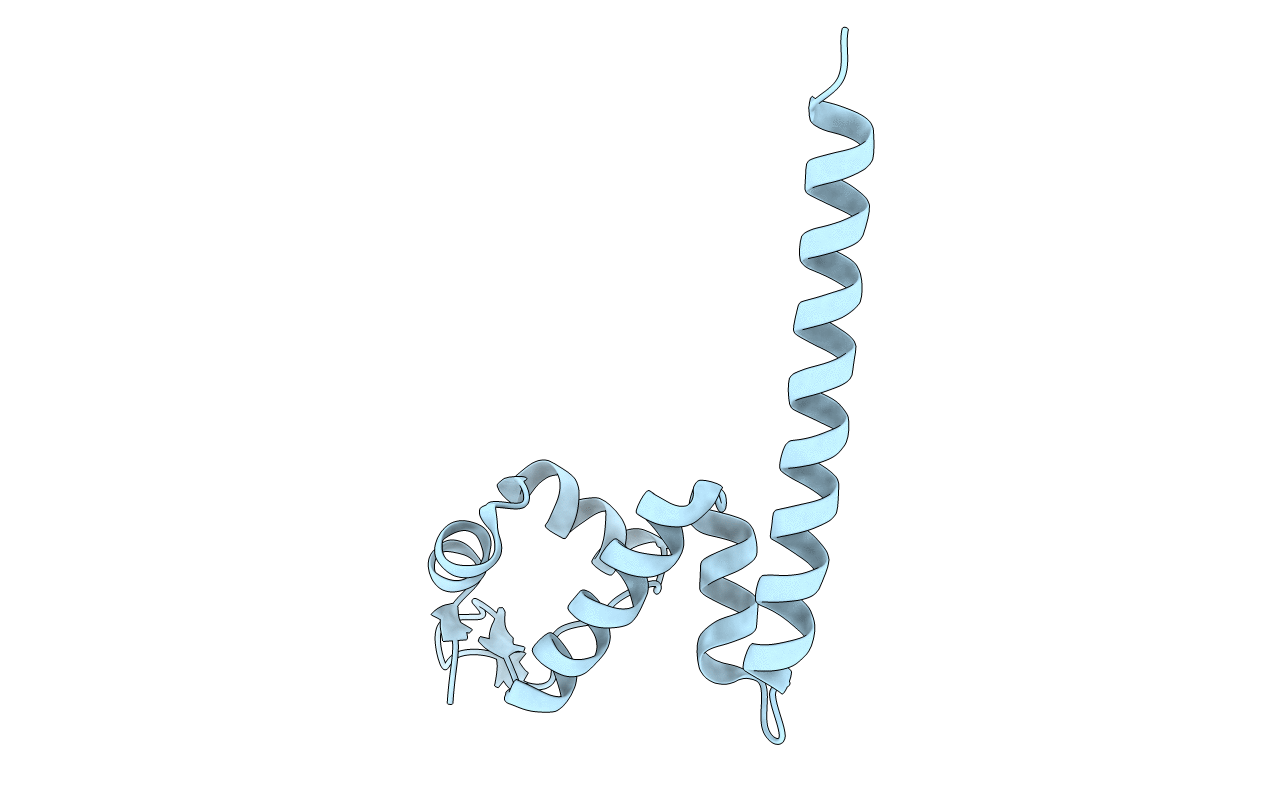
Crystal Structure of MtaN, the Bacillus subtilis Multidrug Transporter Activator, N-terminus
Biological Source:
Source Organism(s):
Bacillus subtilis (Taxon ID: 1423)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.75 Å
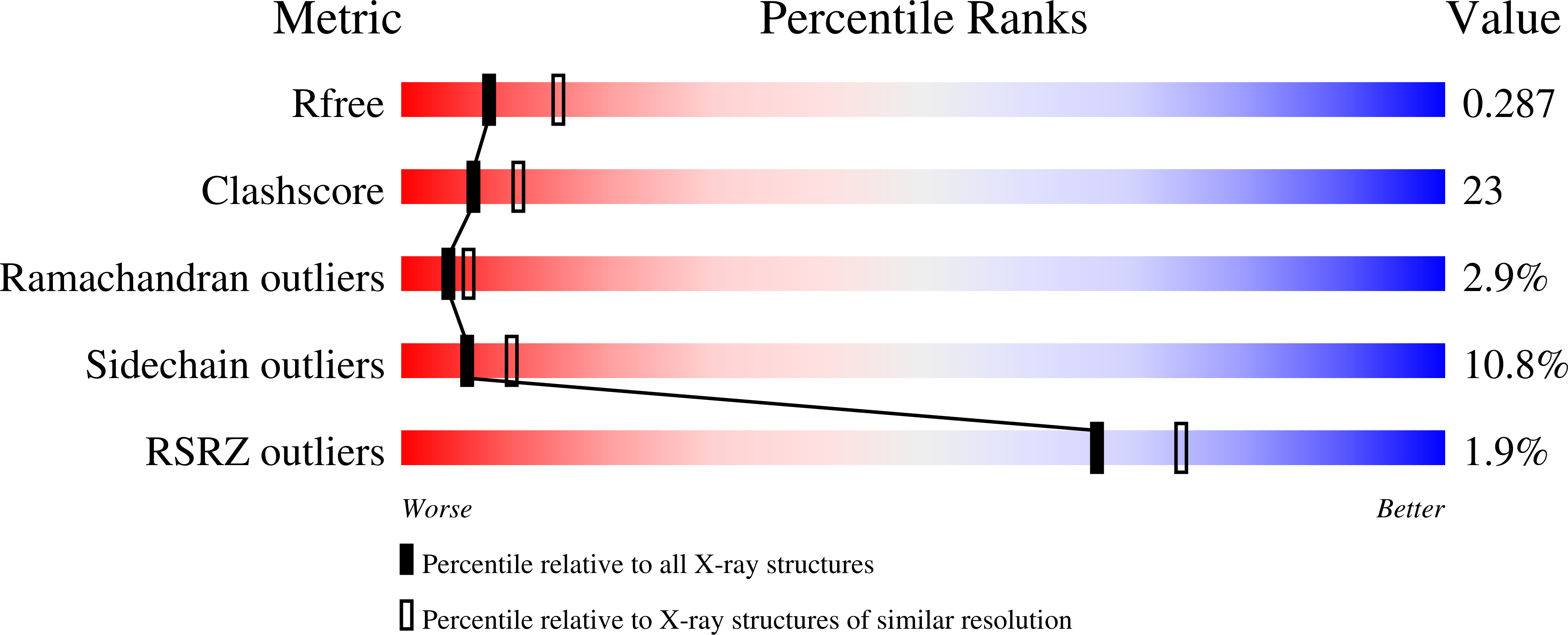
R-Value Free:
0.28
R-Value Work:
0.22
R-Value Observed:
0.23
Space Group:
I 21 21 21