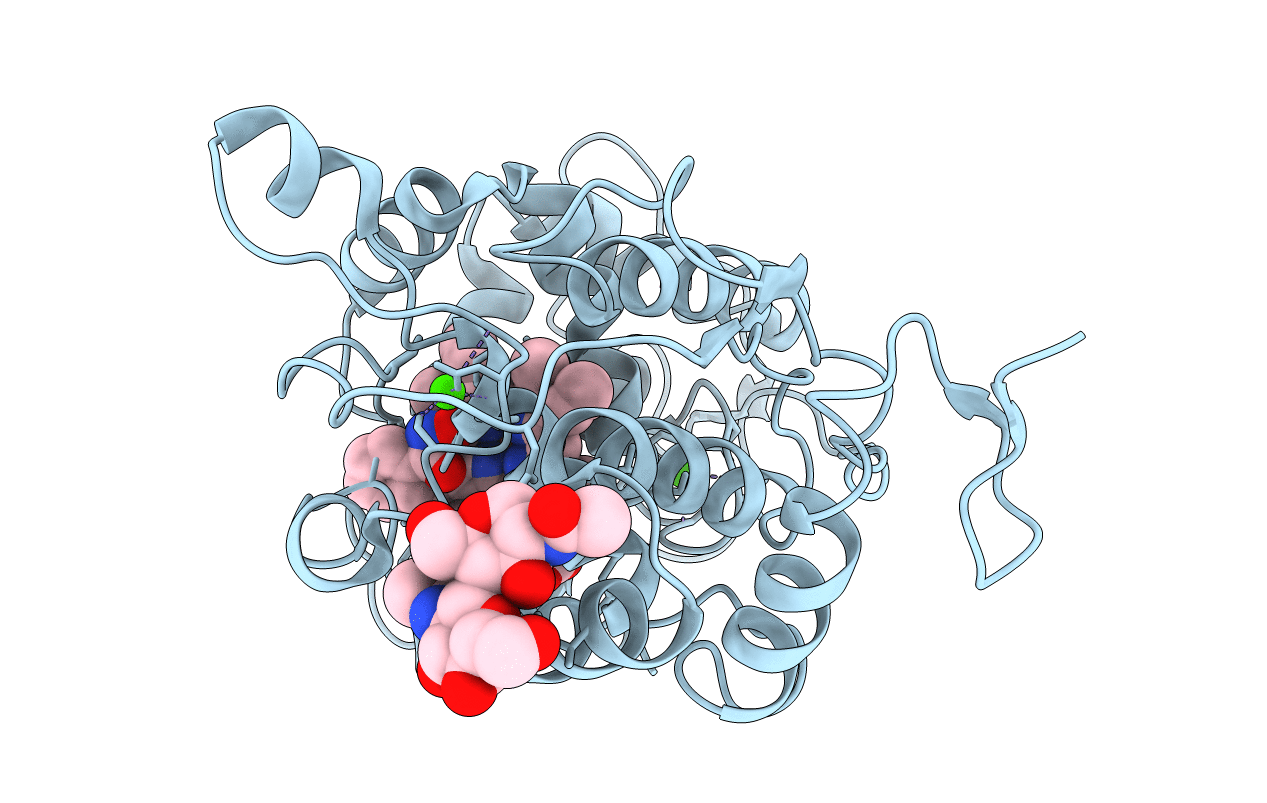
Deposition Date
1997-07-01
Release Date
1998-07-01
Last Version Date
2024-11-06
Entry Detail
PDB ID:
1HSR
Keywords:
Title:
BINDING MODE OF BENZHYDROXAMIC ACID TO ARTHROMYCES RAMOSUS PEROXIDASE
Biological Source:
Source Organism(s):
'Arthromyces ramosus' (Taxon ID: 5451)
Method Details:
Experimental Method:
Resolution:
1.60 Å
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
P 42 21 2