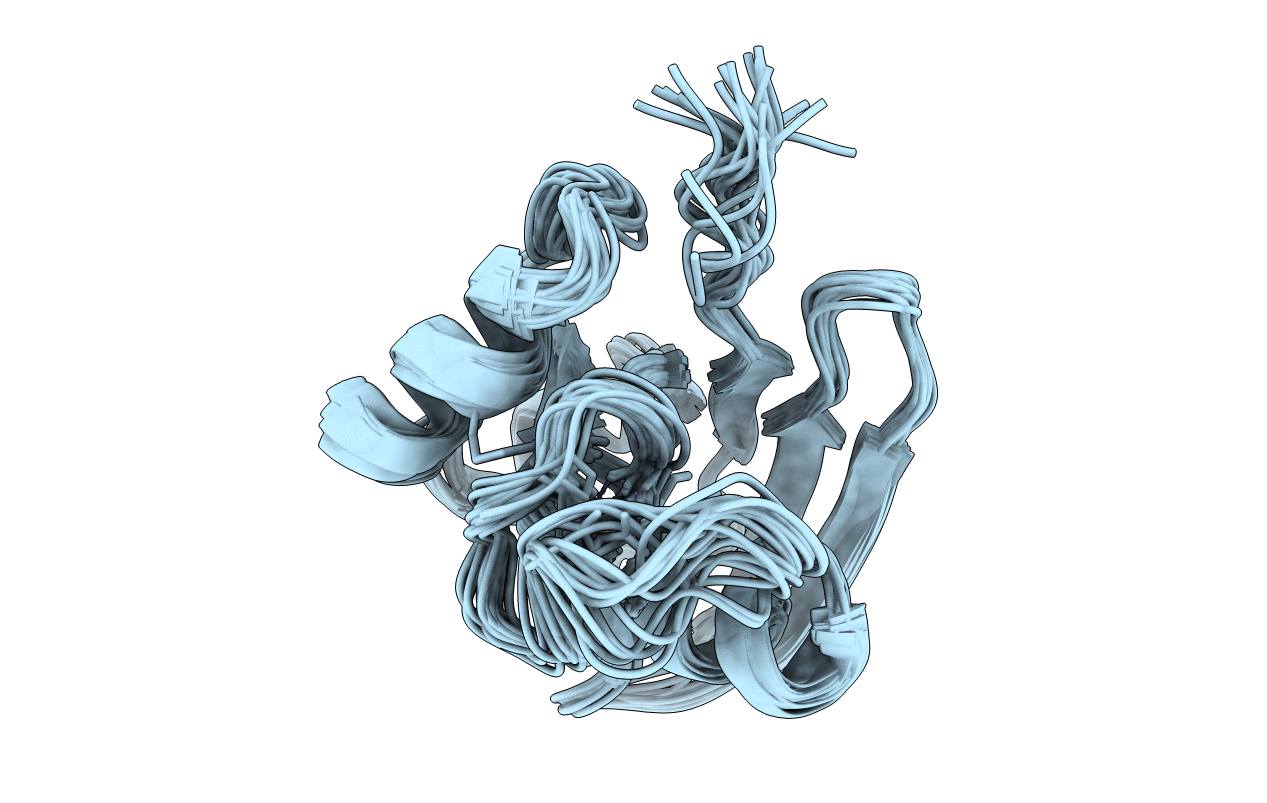
Deposition Date
1998-06-10
Release Date
1999-01-27
Last Version Date
2024-05-01
Entry Detail
Biological Source:
Source Organism(s):
Pseudomonas putida (Taxon ID: 303)
Expression System(s):
Method Details:
Experimental Method:
Conformers Calculated:
100
Conformers Submitted:
20
Selection Criteria:
LEAST RESTRAINT VIOLATION