Deposition Date
1998-05-26
Release Date
1998-08-12
Last Version Date
2024-05-22
Entry Detail
PDB ID:
1BF3
Keywords:
Title:
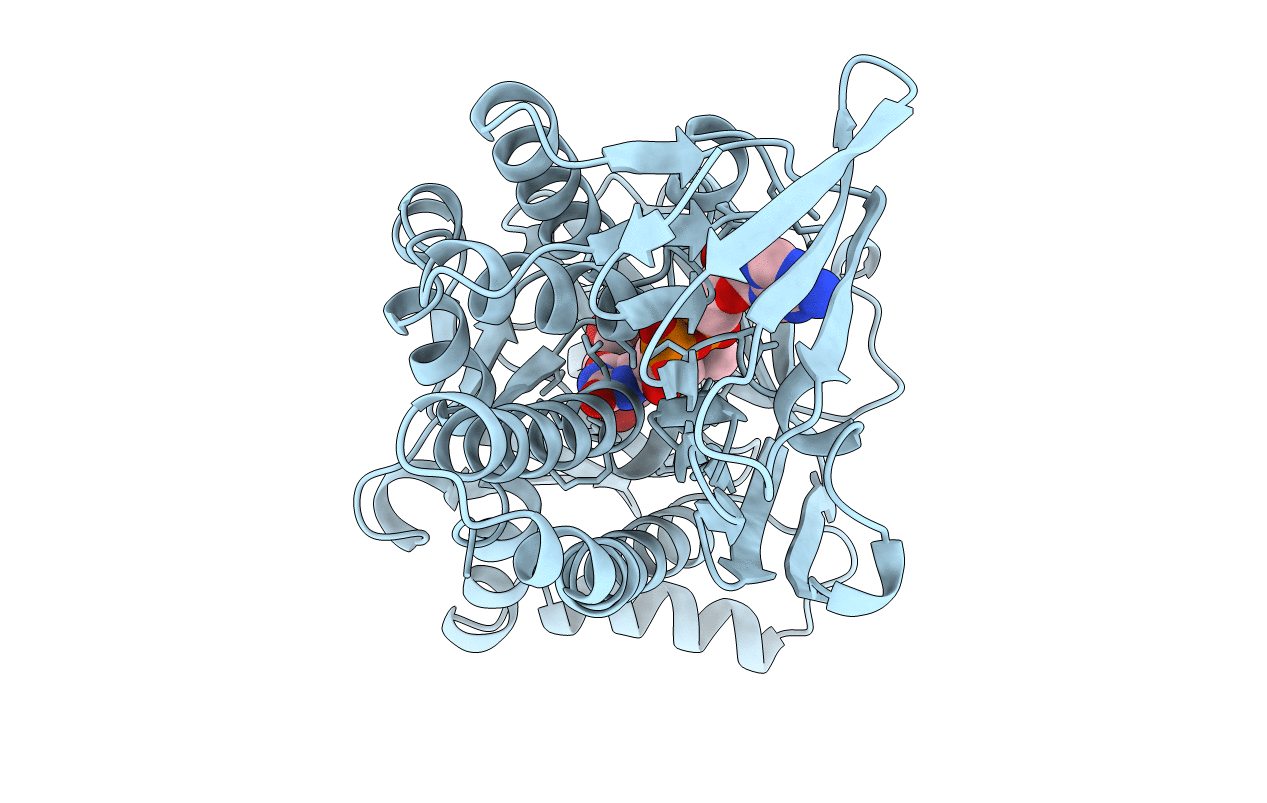
P-HYDROXYBENZOATE HYDROXYLASE (PHBH) MUTANT WITH CYS 116 REPLACED BY SER (C116S) AND ARG 42 REPLACED BY LYS (R42K), IN COMPLEX WITH FAD AND 4-HYDROXYBENZOIC ACID
Biological Source:
Source Organism(s):
Pseudomonas fluorescens (Taxon ID: 294)
Expression System(s):
Method Details: