Search Count: 90
All
Selected
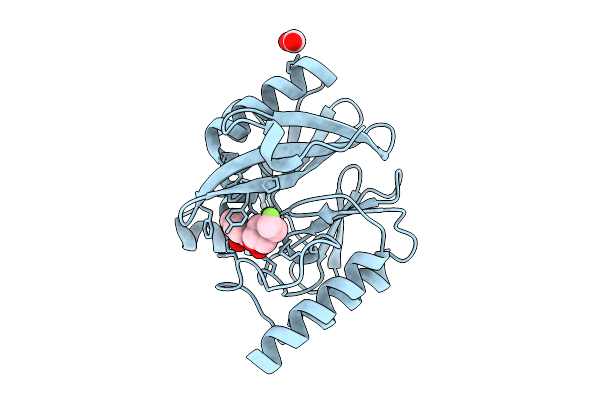 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ZN, A1EOX, FMT |
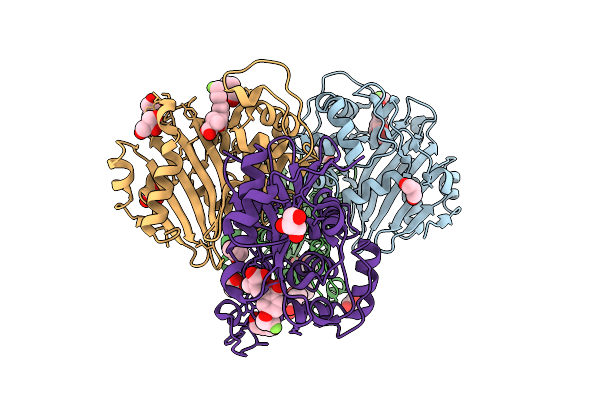 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: A1EO2, GOL, ZN, 1BO |
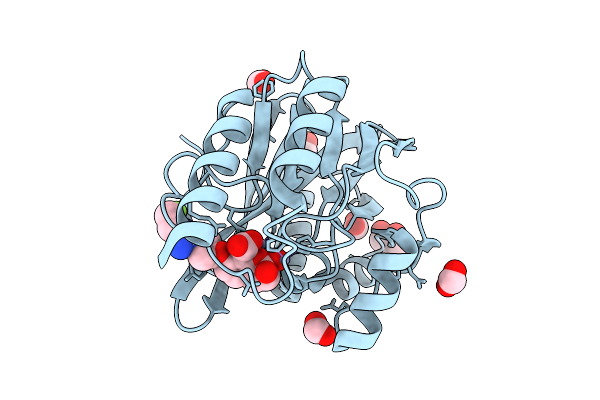 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ZN, A1EO0, GOL, FMT |
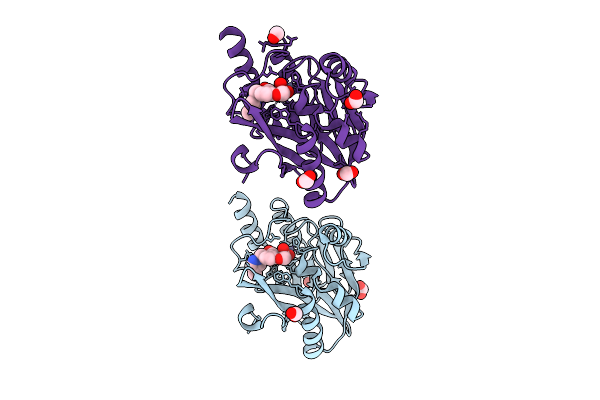 |
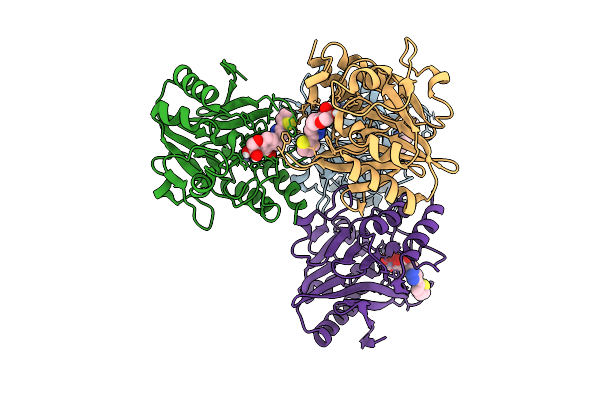
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: ZN, A1ENM, FMT |
 |
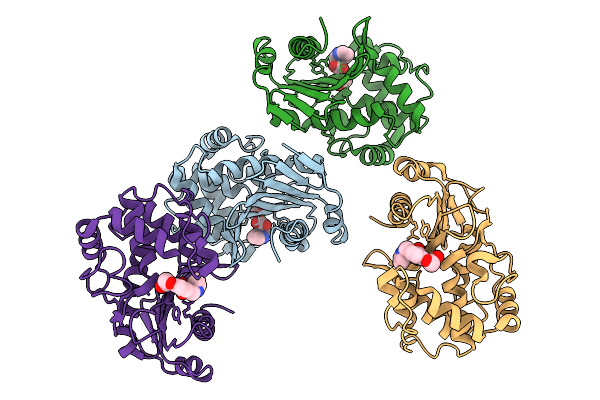
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2026-03-11 Classification: HYDROLASE Ligands: ZN, A1ENL |
 |
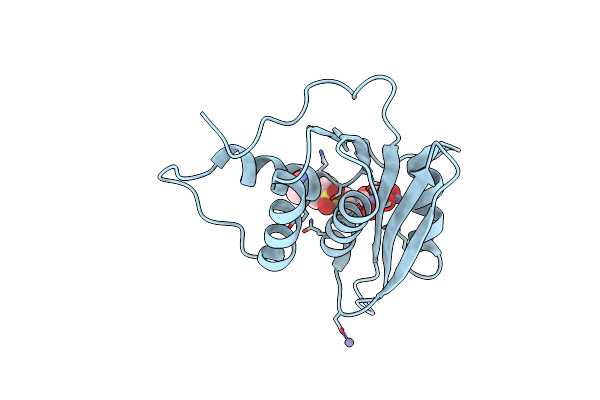
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2025-09-03 Classification: HYDROLASE Ligands: A1EG0 |
 |
Crystal Structure Of Hiv-1 Reverse Transcriptase Rnase H Domain Complexed With A Galloyl Inhibitor
Organism: Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-05-14 Classification: VIRAL PROTEIN Ligands: MN, ZN, A1L9C |
 |
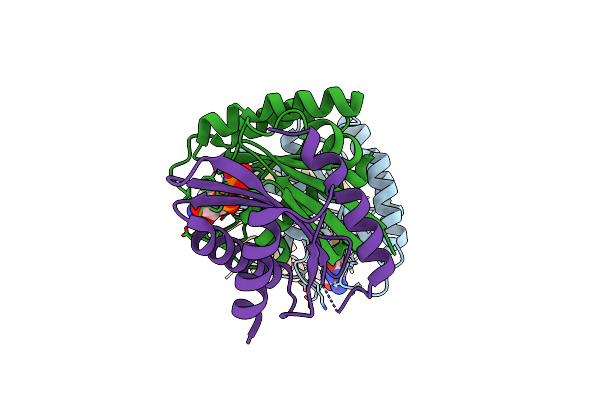
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2024-12-11 Classification: METAL BINDING PROTEIN |
 |
Organism: Synthetic construct, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2024-10-09 Classification: DE NOVO PROTEIN Ligands: GNP, MG |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-10-09 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2024-09-11 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2024-08-14 Classification: DE NOVO PROTEIN Ligands: GOL |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2024-08-14 Classification: DE NOVO PROTEIN |
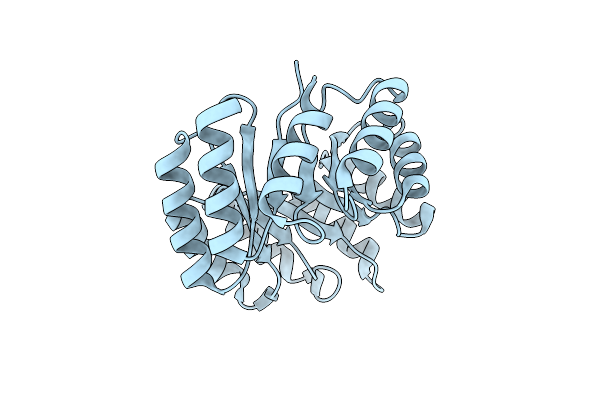 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-08-14 Classification: DE NOVO PROTEIN |
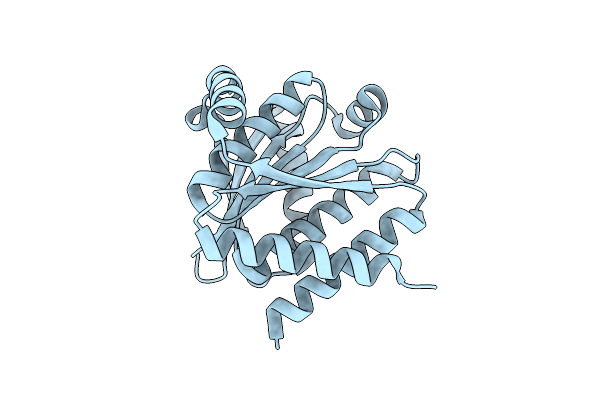 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2024-08-07 Classification: DE NOVO PROTEIN |
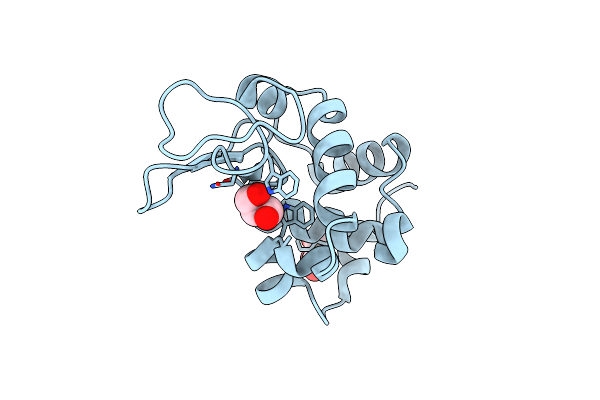 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-08-07 Classification: ANTIMICROBIAL PROTEIN Ligands: GOL |
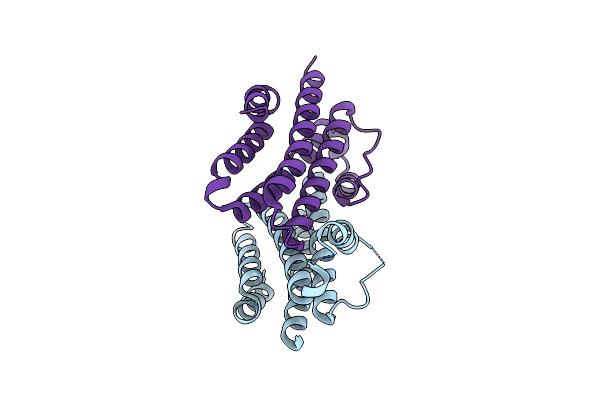 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2024-08-07 Classification: DE NOVO PROTEIN |
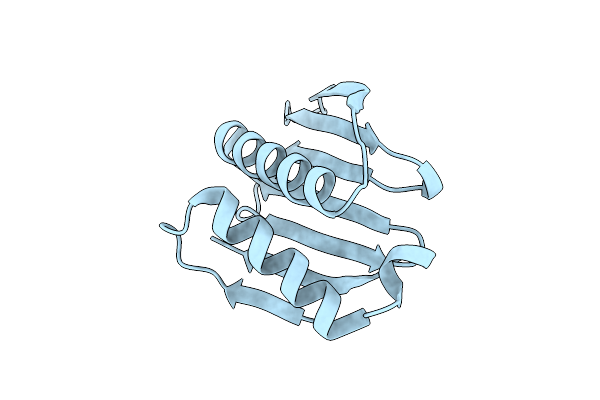 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-08-07 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-08-07 Classification: DE NOVO PROTEIN |
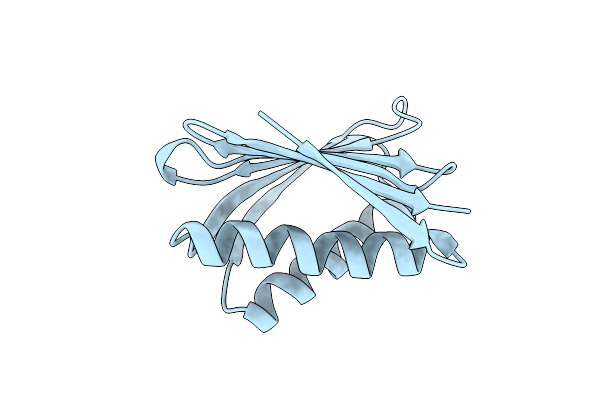 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-08-07 Classification: DE NOVO PROTEIN |

