Search Count: 195
All
Selected
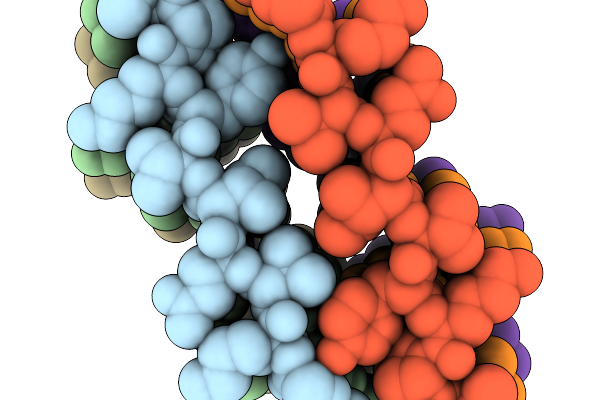 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: PROTEIN FIBRIL |
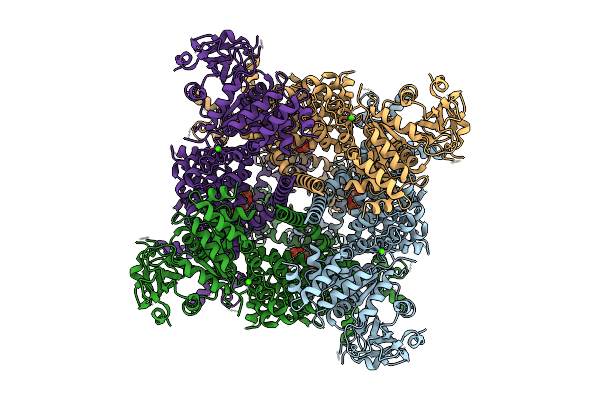 |
Cryo-Em Structure Of The Human Trpm4 Channel In A Calcium And Pi(4,5)P2 Bound Open State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: TRANSPORT PROTEIN Ligands: PT5, CA |
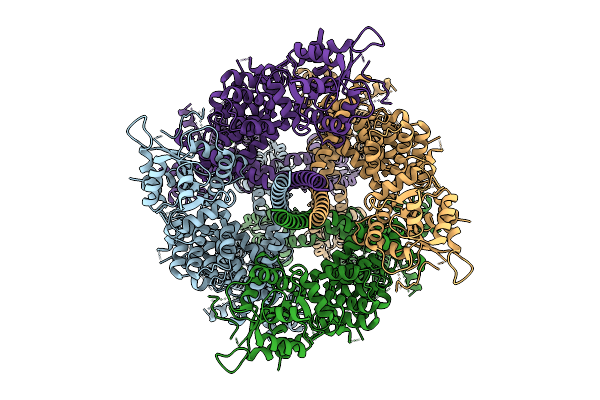 |
Cryo-Em Structure Of The Human Trpm4 Channel In A Calcium Bound Putative Desensitized State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: TRANSPORT PROTEIN Ligands: CA |
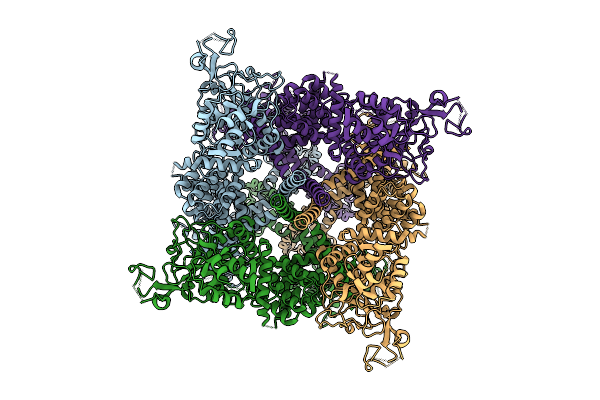 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: TRANSPORT PROTEIN |
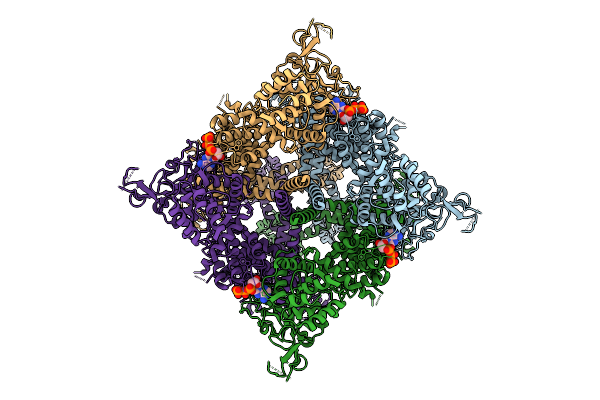 |
Cryo-Em Structure Of The Human Trpm4 Channel In A Atp Bound Inhibited State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: TRANSPORT PROTEIN Ligands: ATP |
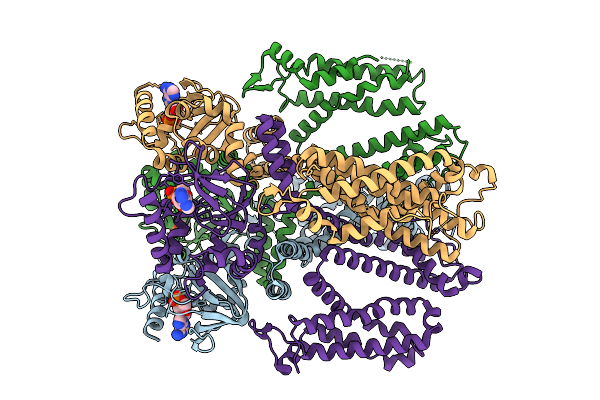 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.59 Å Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: CA, CMP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: CA |
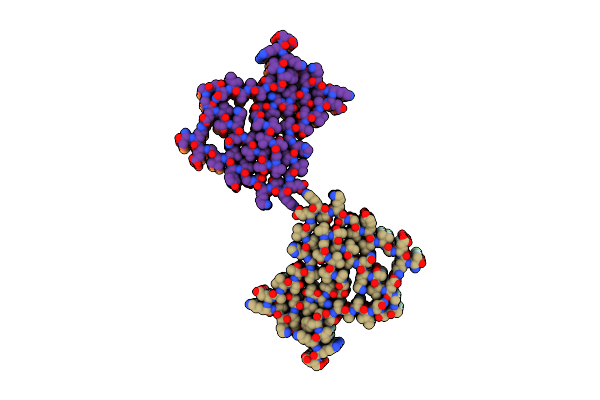 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: PROTEIN FIBRIL |
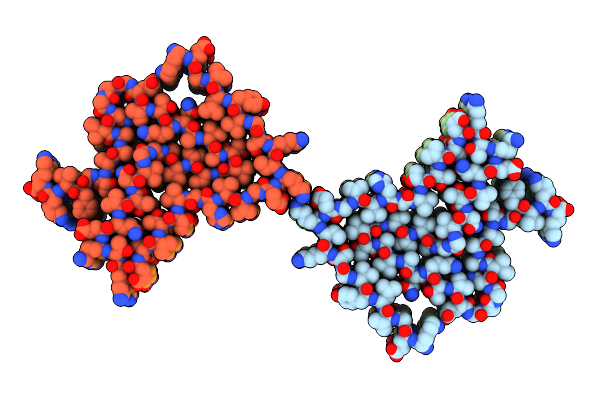 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: PROTEIN FIBRIL |
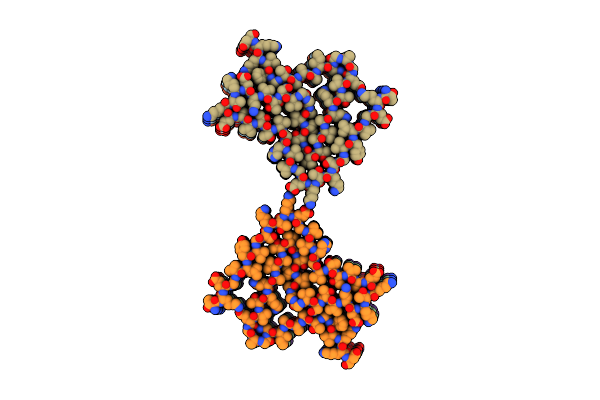 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: PROTEIN FIBRIL |
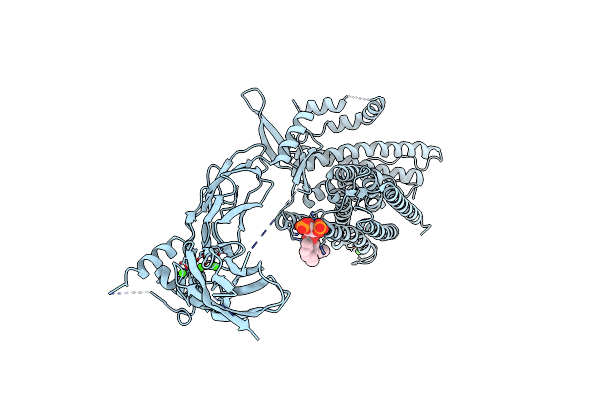 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-03-19 Classification: TRANSPORT PROTEIN Ligands: CA, PIO |
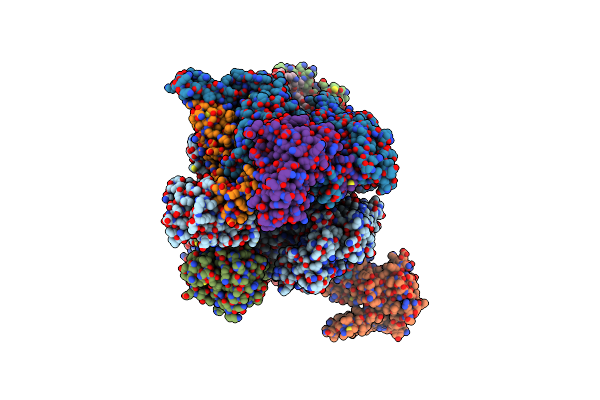 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-29 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSCRIPTION/RNA/DNA Ligands: ZN, MG |
 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSCRIPTION Ligands: ZN, MG |
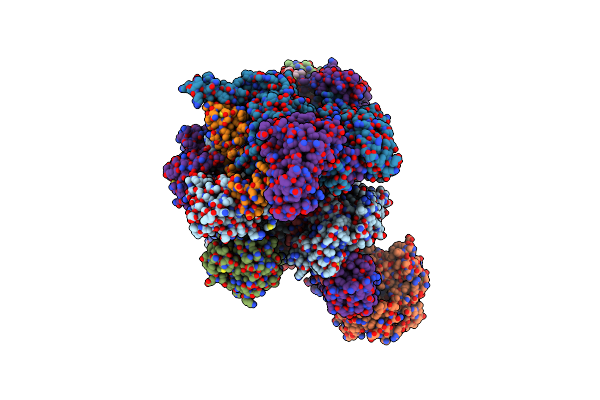 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Organism: African swine fever virus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-22 Classification: TRANSCRIPTION Ligands: ZN, MG |
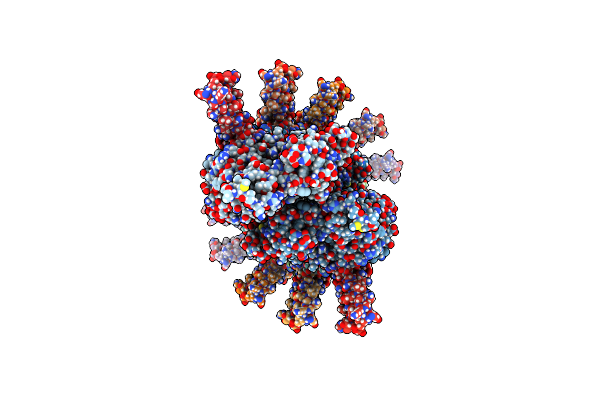 |
Actin-Binding Domain Of Legionella Pneumophila Effector Lfat1 (Lpg1387) Bound To F-Actin
Organism: Legionella pneumophila, Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-18 Classification: TOXIN Ligands: ADP, MG |
 |
Cryo-Em Structure Of Mouse Trpml1 Channel Y404W At 2.86 Angstrom Resolution
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: MEMBRANE PROTEIN |
 |
Cryo-Em Structure Of Mouse Pi(4,5)P2-Bound Trpml1 Channel At 2.46 Angstrom Resolution
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: MEMBRANE PROTEIN Ligands: PIO, FO4 |

