Search Count: 109
All
Selected
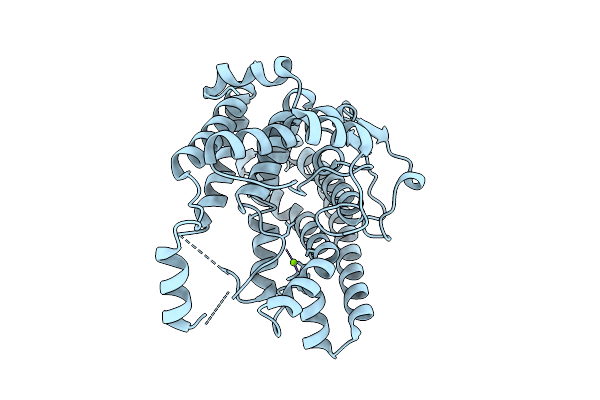 |
Crystal Structure Of The Tc Domain Of A Bifunctional Sesterterpene Synthase
Organism: Aspergillus stellatus
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-03-11 Classification: BIOSYNTHETIC PROTEIN Ligands: MG |
 |
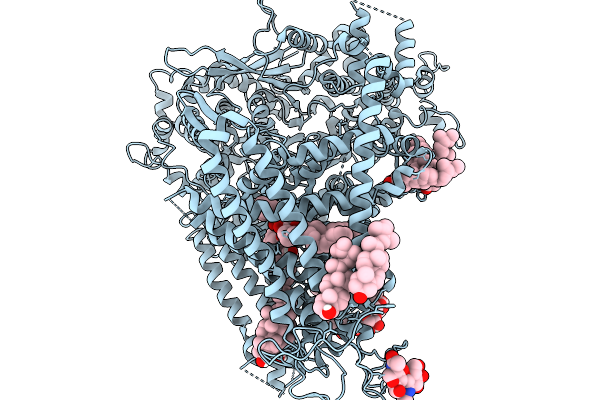
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: POV, AV0, ERG |
 |
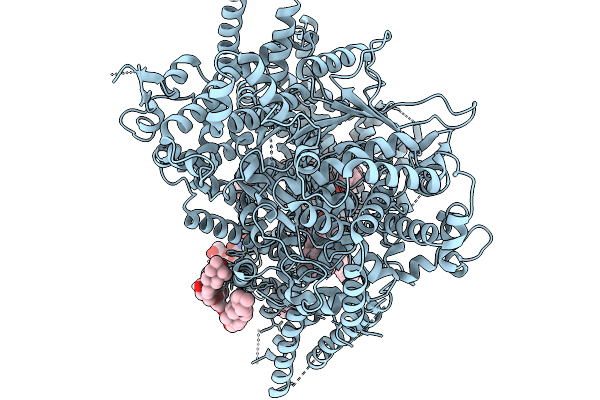
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
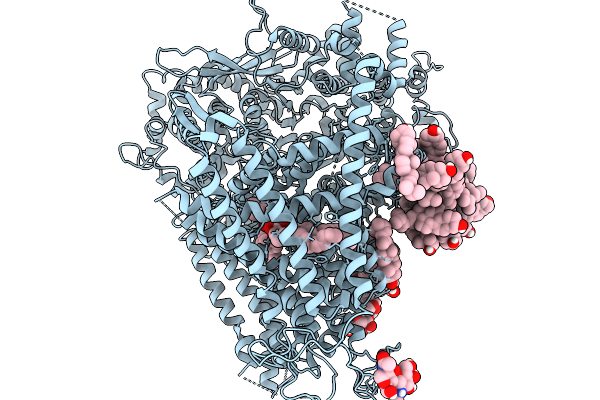
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: AV0, ERG, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Penicillium brasilianum
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
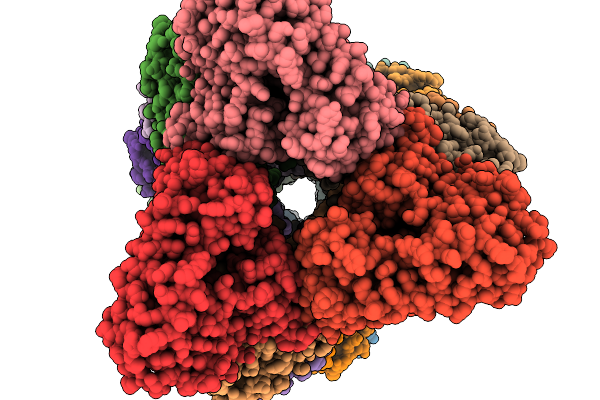 |
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Resolution:3.19 Å Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
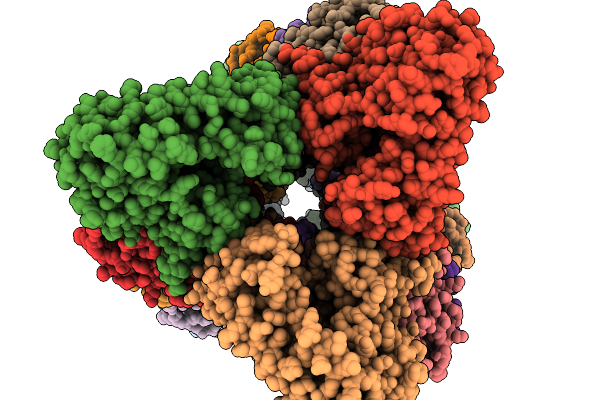 |
Organism: Aspergillus stellatus
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2026-02-11 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-14 Classification: PROTEIN BINDING Ligands: A1EJ5 |
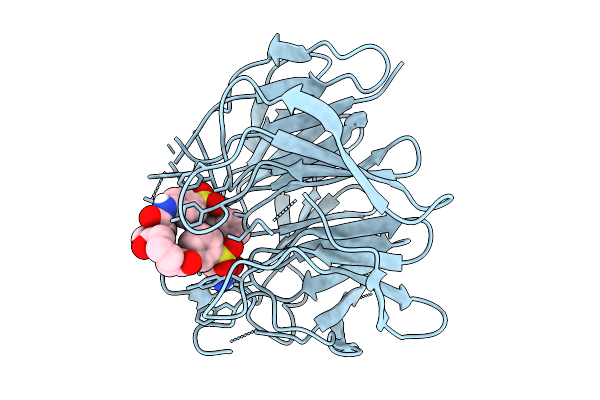 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-12-31 Classification: PROTEIN BINDING Ligands: A1EJB |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-12-10 Classification: PROTEIN BINDING Ligands: A1EHH |
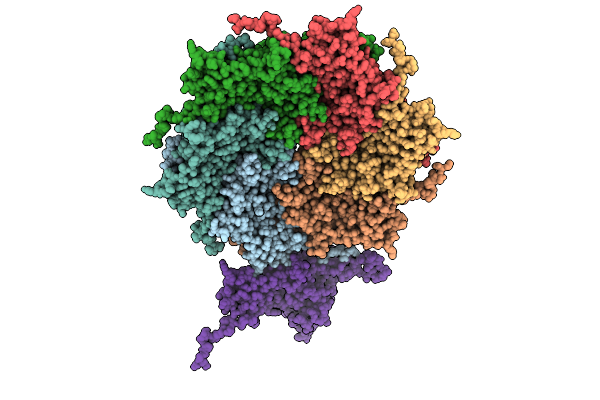 |
Organism: Pseudanabaena phage pan3
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-11-26 Classification: VIRUS |
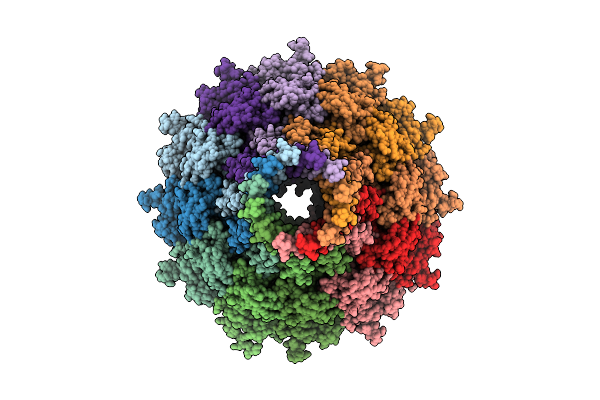 |
Organism: Pseudanabaena phage pan3
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN |
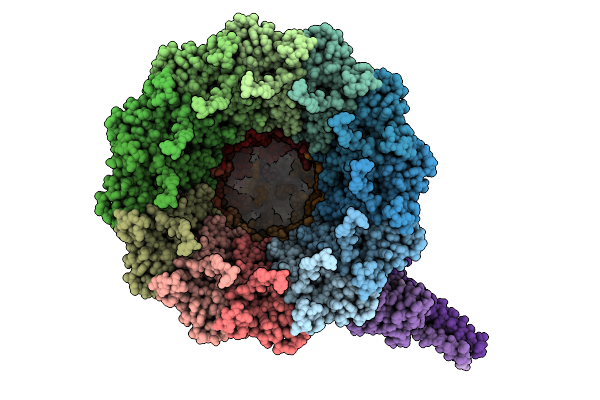 |
Organism: Pseudanabaena phage pan3
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN |
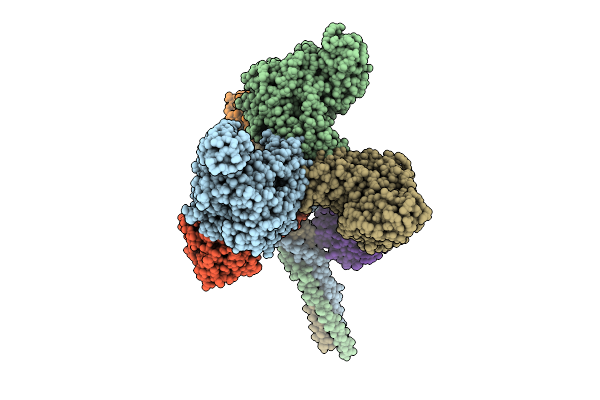 |
Organism: Pseudanabaena phage pan3
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-11-26 Classification: VIRAL PROTEIN |
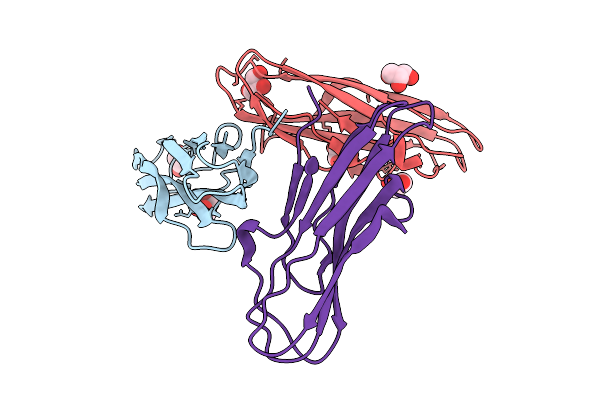 |
Organism: Pseudanabaena phage pan3
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-11-26 Classification: VIRUS Ligands: IOD, GOL |

