Search Count: 436
All
Selected
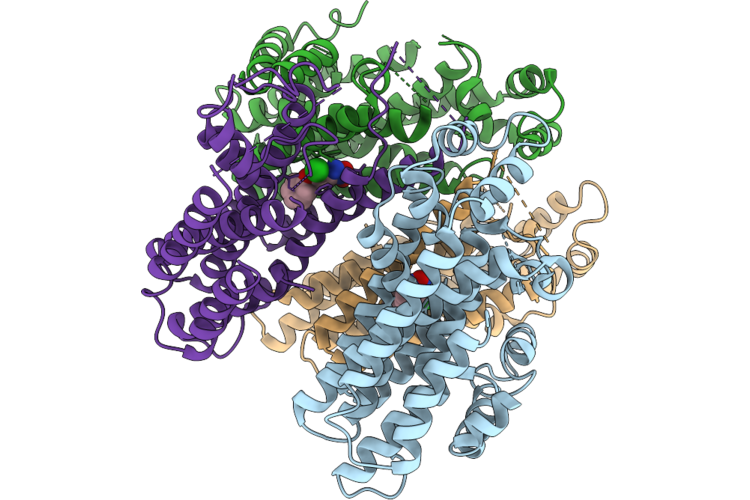 |
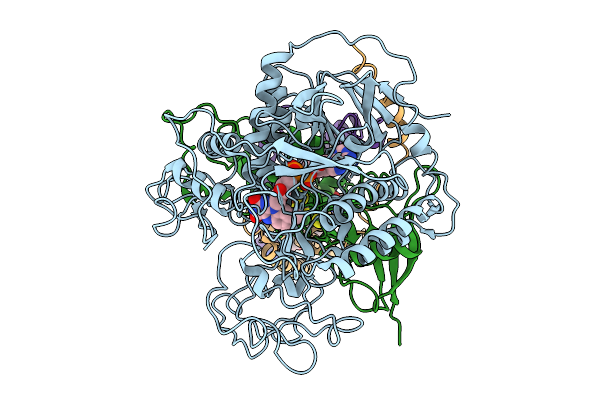
Crystal Structure Of Setaria Viridis Sps Complexed With Aclonifen (Surface Polar Residue Mutant)
Organism: Setaria viridis
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2026-04-29 Classification: TRANSFERASE Ligands: A1L38 |
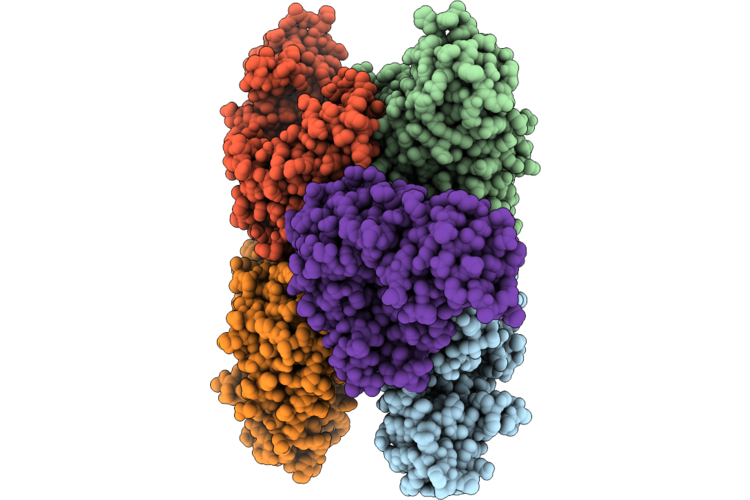 |
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-04-29 Classification: TRANSFERASE |
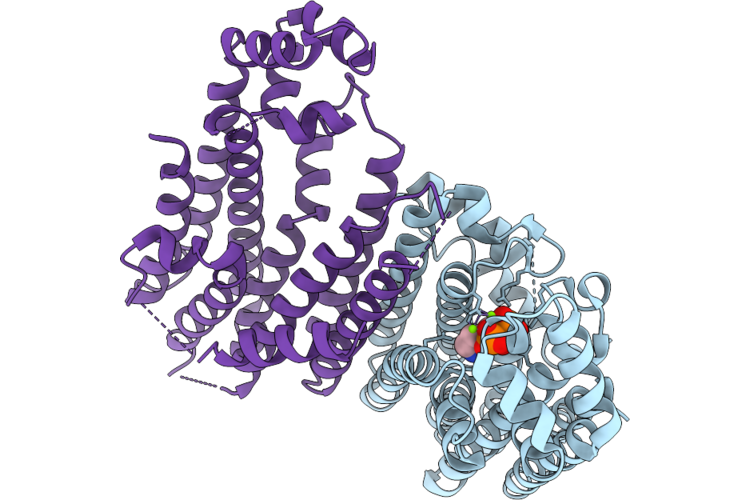 |
Organism: Sorghum bicolor
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-04-29 Classification: TRANSFERASE Ligands: ZOL, MG |
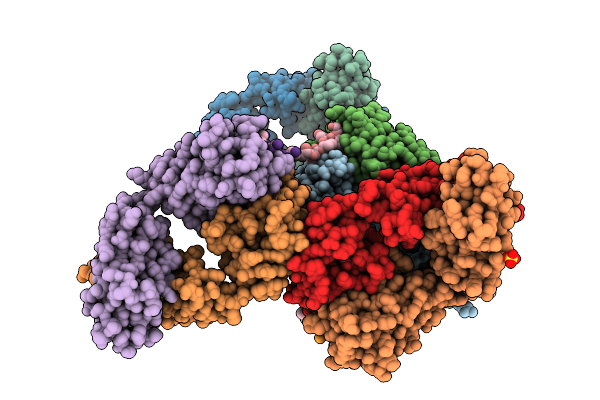 |
Organism: Homo sapiens, Coroavirus
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: SO4 |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDE |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDF |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDG |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDH |
 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDI |
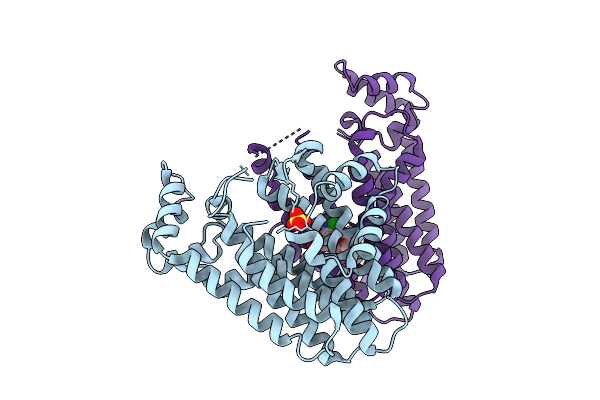 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: SO4, A1EDT |
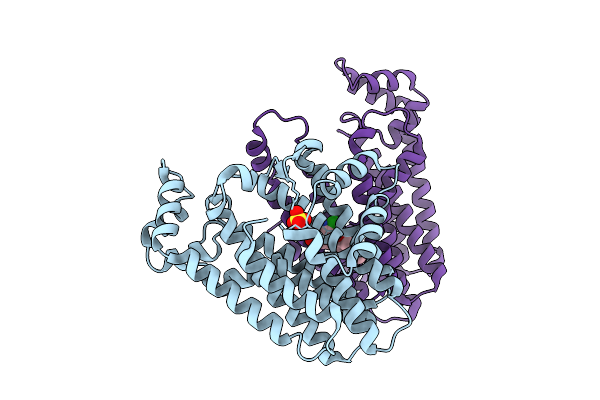 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC1 |
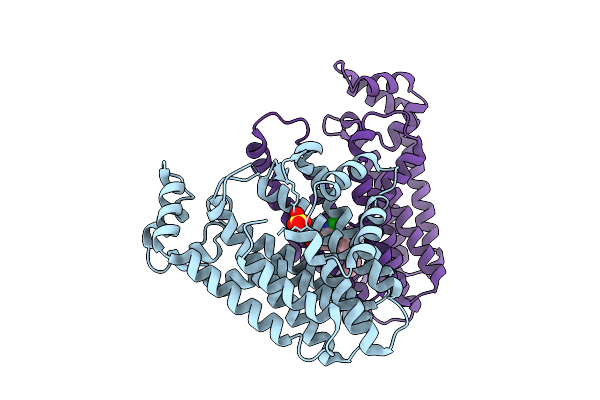 |
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1EC2 |
 |
Organism: Henipavirus nipahense
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN/RNA |
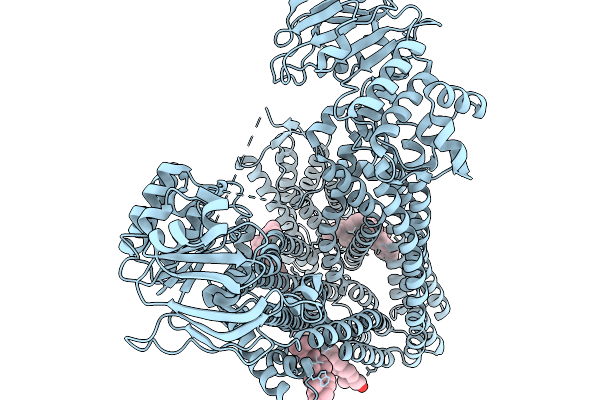 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: CLR |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: CLR, R16 |
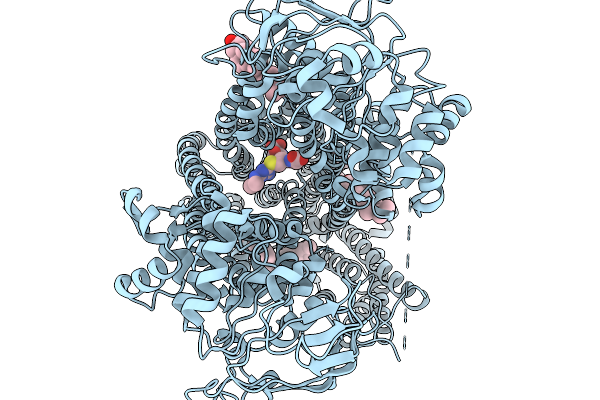 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: ATA, CLR |
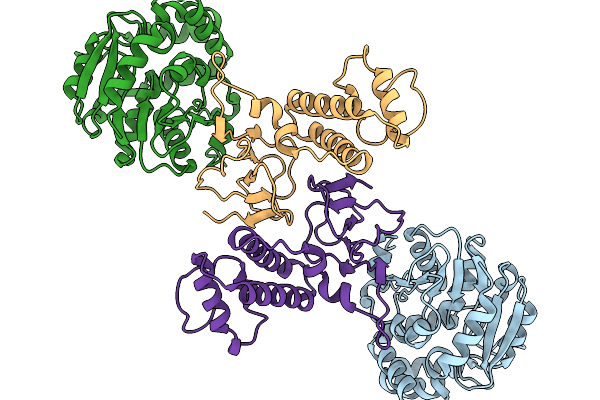 |
Organism: Oryza sativa japonica group, Tenuivirus oryzabrevis
Method: ELECTRON MICROSCOPY Resolution:3.67 Å Release Date: 2026-03-04 Classification: PLANT PROTEIN |
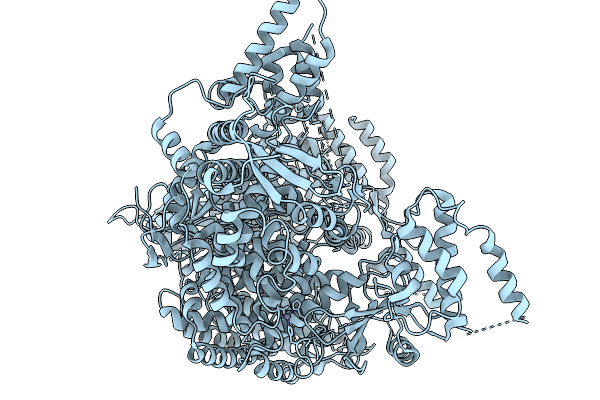 |
2.62A Cryo-Em Structure Of Rna-Directed Rna Polymerase L Of Crimean-Congo Hemorrhagic Fever Virus (Apo State)
Organism: Crimean-congo hemorrhagic fever virus
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2026-03-04 Classification: TRANSFERASE Ligands: ZN |
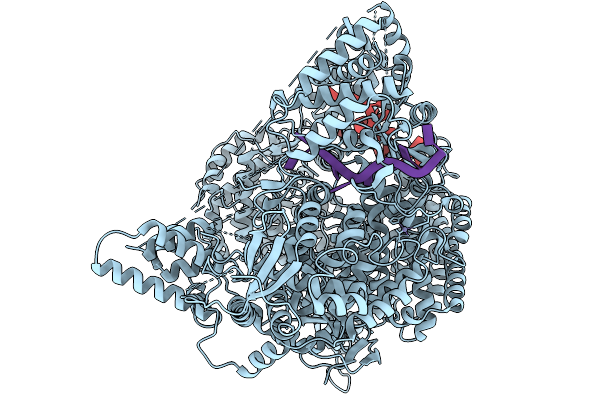 |
2.53A Cryo-Em Structure Of Rna-Directed Rna Polymerase L Of Crimean-Congo Hemorrhagic Fever Virus (Rna Bound)
Organism: Crimean-congo hemorrhagic fever virus, Crimean-congo hemorrhagic fever virus strain ibar10200
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-03-04 Classification: TRANSFERASE/RNA Ligands: ZN |
 |
Cryo-Em Structure Of Saccharomyces Cerevisiae Mitochondrial Respiratory Complex Ii In Cyclobutrifluram-Bound State
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: FAD, A1EKU, FES, SF4, F3S, PEE |

