Search Count: 542
All
Selected
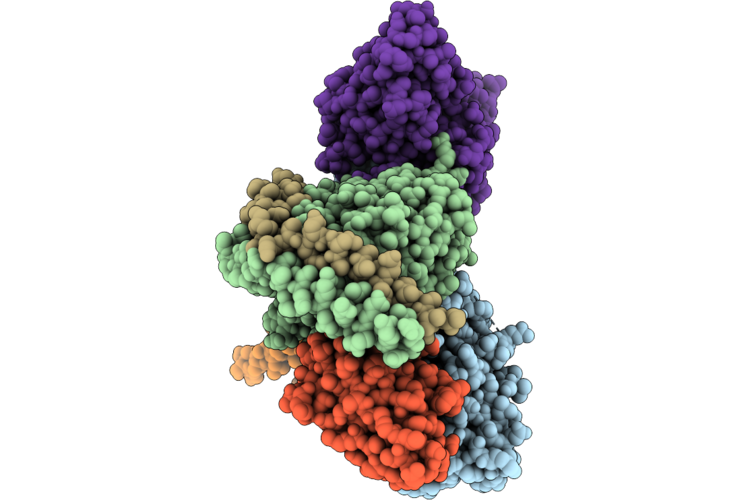 |
Organism: Petromyzon marinus, Homo sapiens, Lama glama, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: SPM |
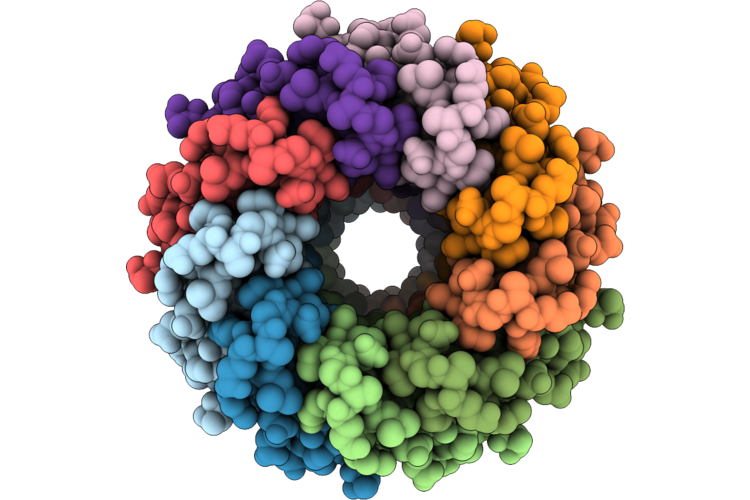 |
Organism: Pseudanabaena phage pam5
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-01 Classification: ANTITOXIN |
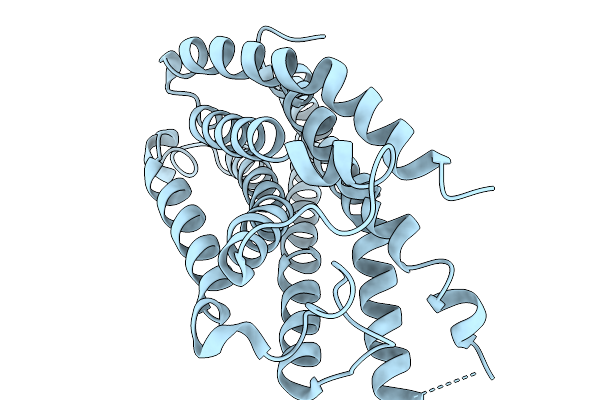 |
Organism: Homo sapiens, Escherichia coli o157:h7
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
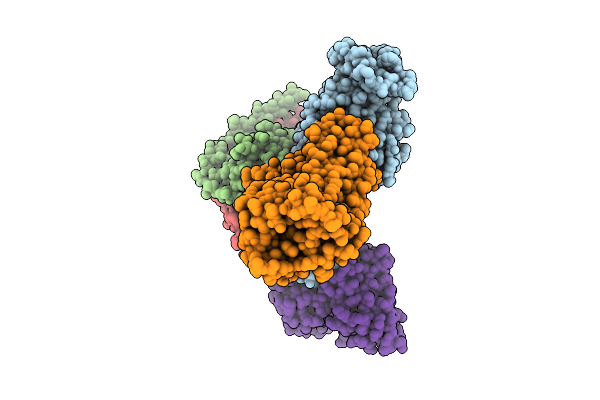 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: AKG |
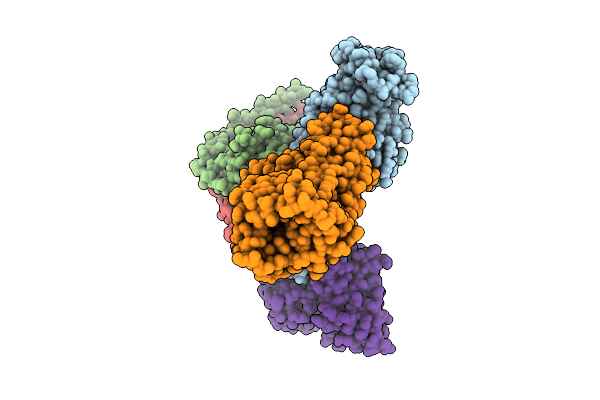 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: ITN |
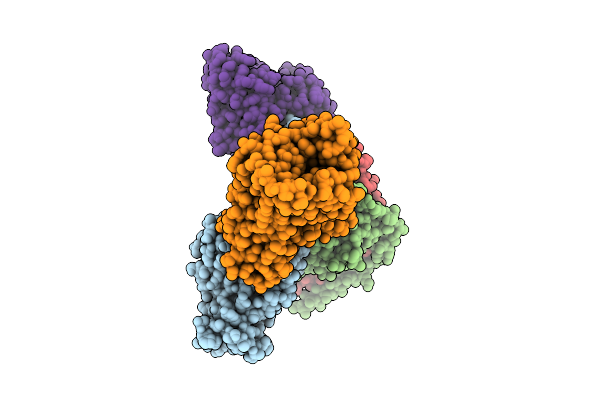 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E2R |
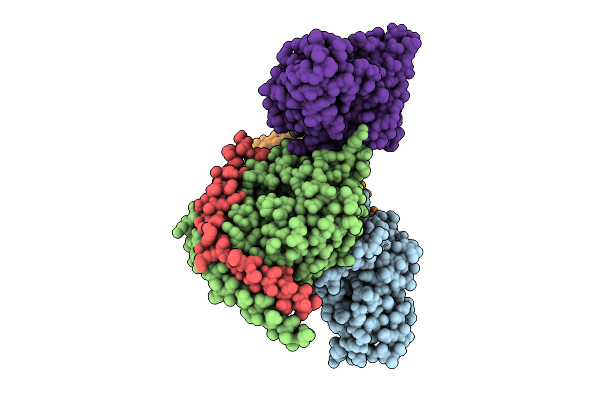 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
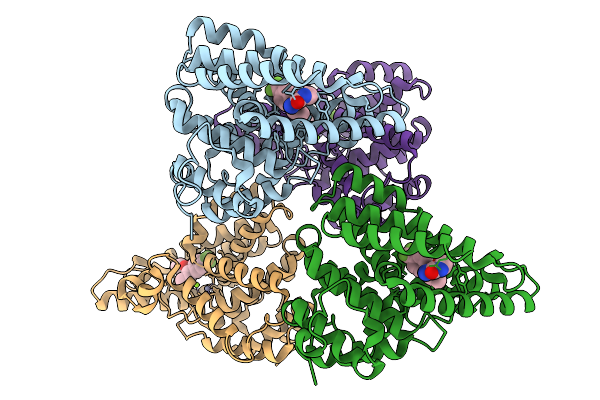 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: ZN, MG, A1E46 |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: PROTEIN BINDING |
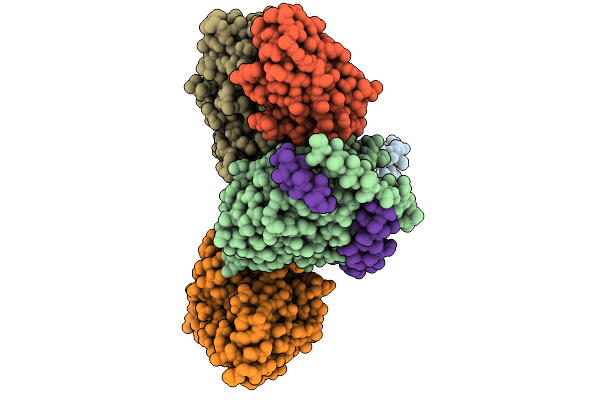 |
Organism: Synthetic construct, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
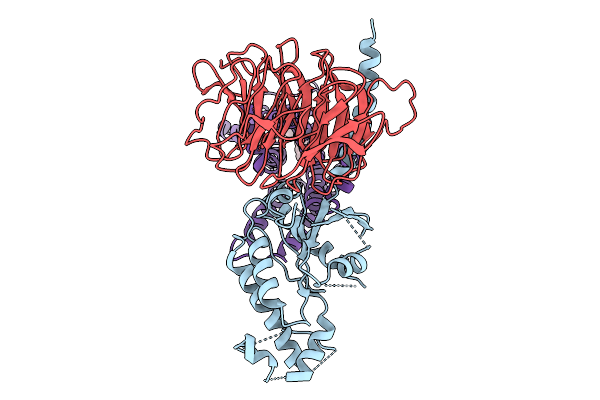 |
Organism: Synthetic construct, Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1EKK |
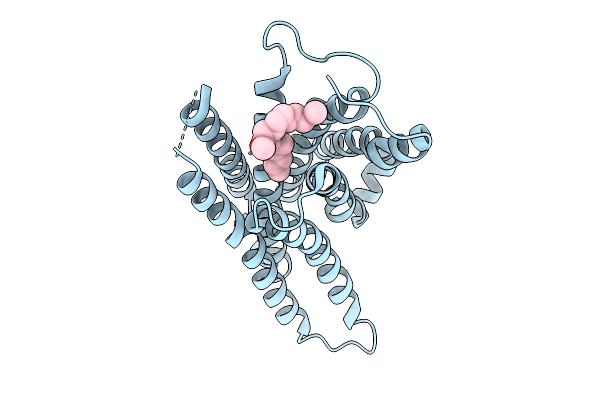 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1EKK |
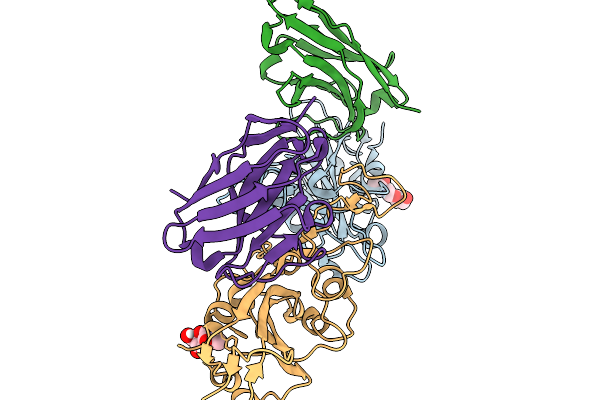 |
Crystal Structure Of Sars-Cov-2 Spike Receptor-Binding Domain (Delta) In Complex With Ph-Dependent Nanobody Mnb-11.
Organism: Severe acute respiratory syndrome coronavirus 2, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-01-14 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
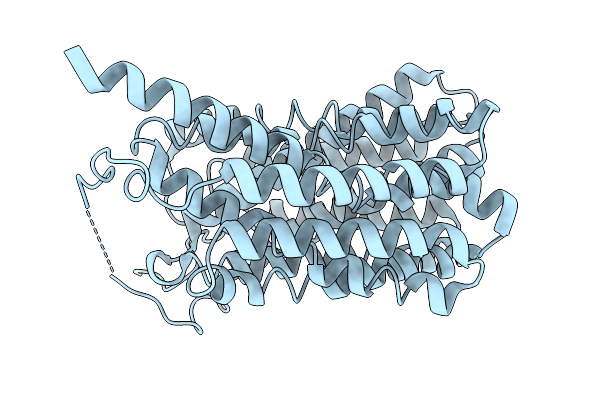 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
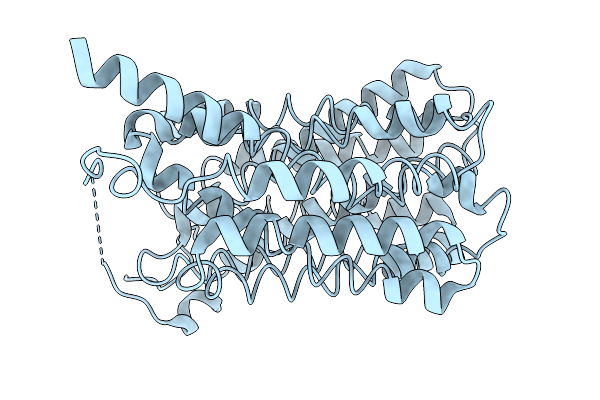 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
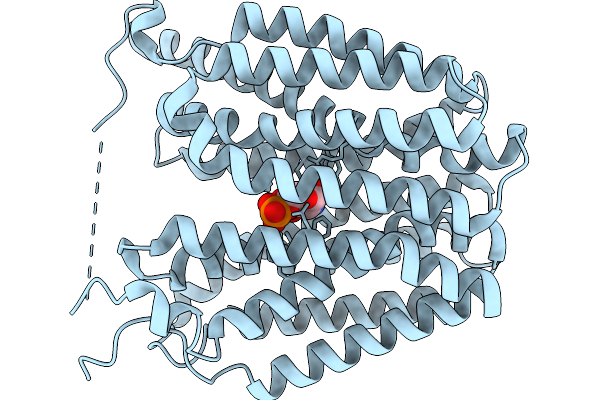 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: GLP |
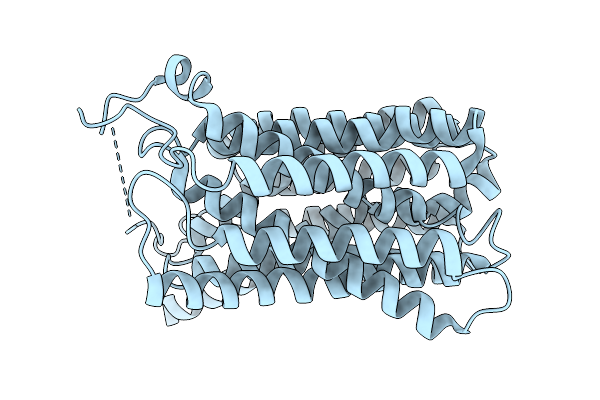 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.47 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
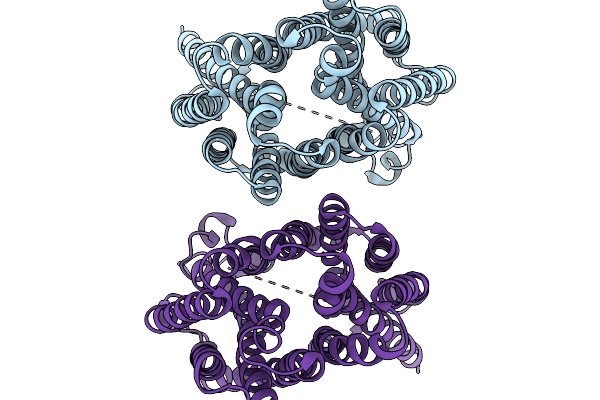 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.89 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: PO4 |

