Search Count: 76
All
Selected
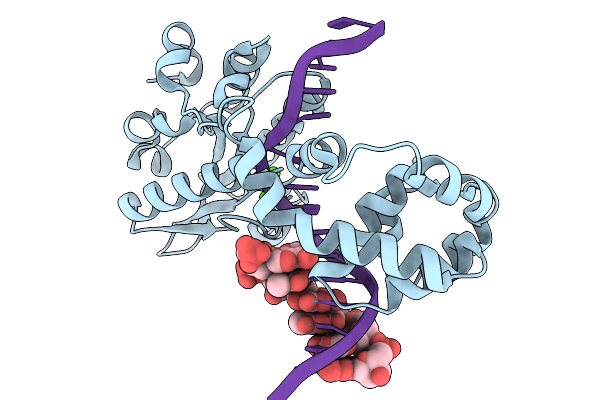 |
Organism: Geobacillus stearothermophilus, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: CA |
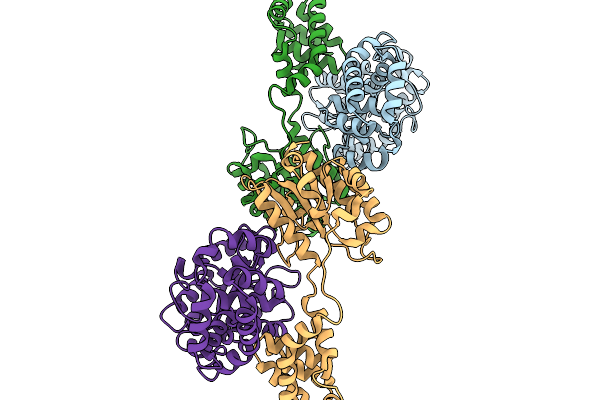 |
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.72 Å Release Date: 2026-03-04 Classification: HYDROLASE |
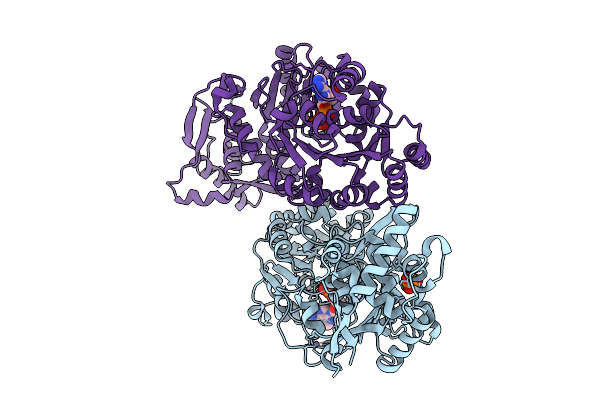 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: ZN, ADP, MG, PO4 |
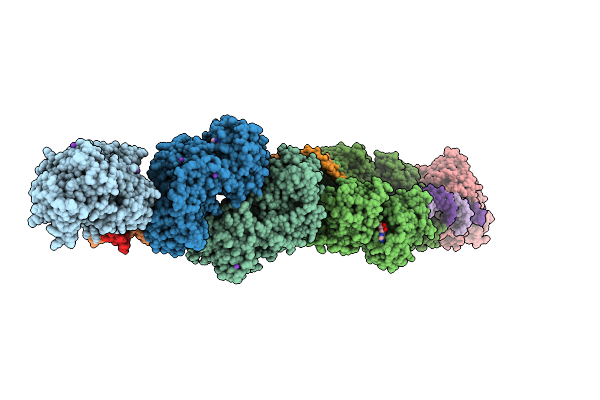 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: ZN, ADP, MG, PO4, K |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: ZN, ADP, MG, PO4, K |
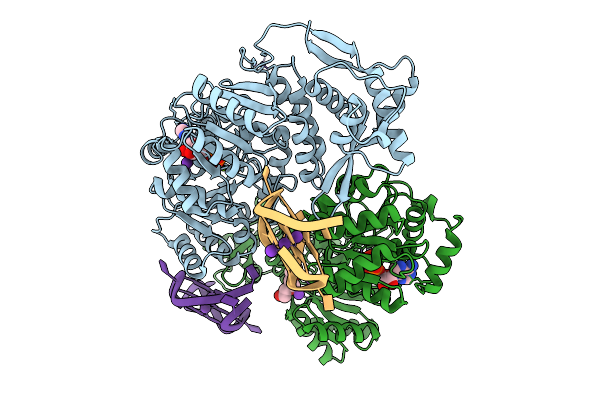 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: ZN, ADP, MG, K, PGE |
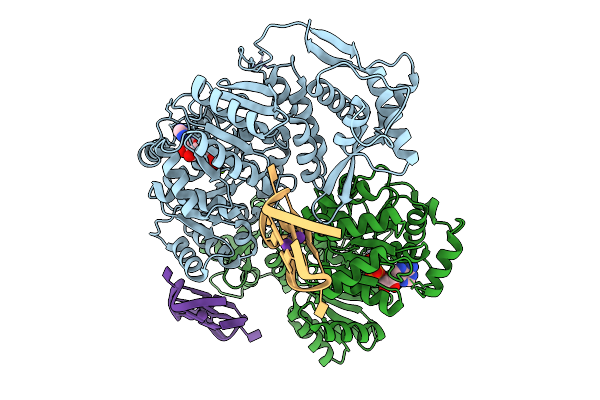 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.69 Å Release Date: 2025-10-01 Classification: HYDROLASE Ligands: ZN, ADP, MG, K |
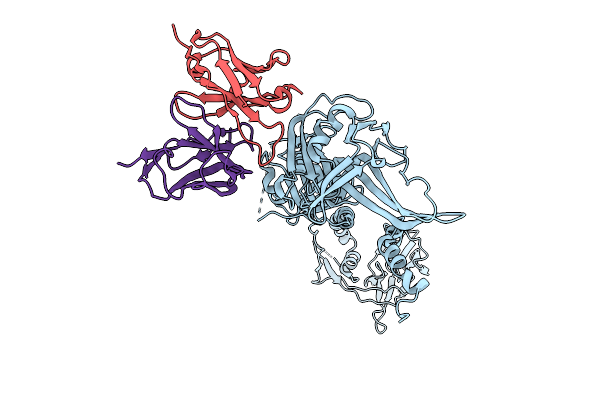 |
Organism: Human herpesvirus 2, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Mus musculus, Cercopithecine herpesvirus 1
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
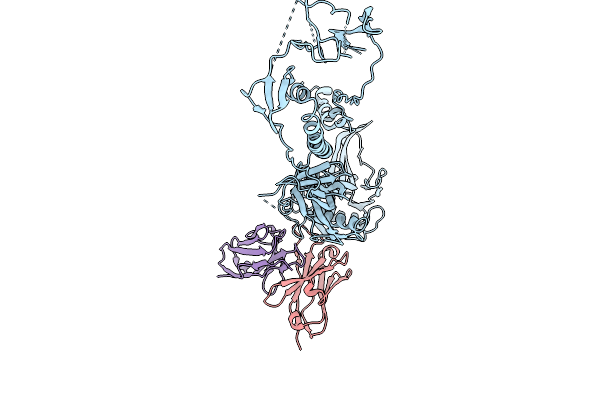 |
Organism: Suid alphaherpesvirus 1, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Human herpesvirus 3, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.66 Å Release Date: 2024-05-08 Classification: HYDROLASE Ligands: PO4 |
 |
Structure Of Escherichia Coli Hrpa In Complex With Adp And Dinucleotide Dcdc
Organism: Escherichia coli k-12, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2024-05-08 Classification: HYDROLASE Ligands: ADP, MG, PO4 |
 |
Structure Of Escherichia Coli Hrpa In Complex With Adp And Oligonucleotide Poly(Dc)11 Forming An I-Motif
Organism: Escherichia coli k-12, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2024-05-08 Classification: HYDROLASE Ligands: ADP, MG, SIN |
 |
Organism: Oryza sativa indica group
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG, SIN |
 |
Organism: Oryza sativa indica group
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG, PO4, NA |
 |
Structure Of Nal1 Indica Cultivar Ir64, Construct 36-458 In Presence Of Peptide From Fzp Protein
Organism: Oryza sativa indica group
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG, SO4, NH4 |
 |
Structure Of Nal1 Protein , Spike Allele From Japonica Rice, Construct 46-458
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG |
 |
Structure Of Nal1 Protein, Allele Spike From Japonica Rice, Construct 31-458
Organism: Oryza sativa japonica group
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG, PO4 |
 |
Cryoem Structure Of Nal1 Protein, Allele Spike, From Oryza Sativa Japonica Group
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2024-04-17 Classification: PLANT PROTEIN Ligands: ATP, MG |

