Search Count: 41
All
Selected
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: DQC |
 |
Organism: Mus musculus, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: GGL, A1EA4 |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: TRANSPORT PROTEIN Ligands: L9Q, CDL |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: TRANSPORT PROTEIN Ligands: RFP, CDL |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: ELECTRON MICROSCOPY Release Date: 2024-07-17 Classification: TRANSPORT PROTEIN Ligands: CDL, ANP, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2023-07-12 Classification: HYDROLASE Ligands: ZN, MG, JN0, EDO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-07-12 Classification: HYDROLASE Ligands: ZN, MG, JU3, EDO |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: ELECTRON MICROSCOPY Release Date: 2023-05-31 Classification: MEMBRANE PROTEIN Ligands: CA |
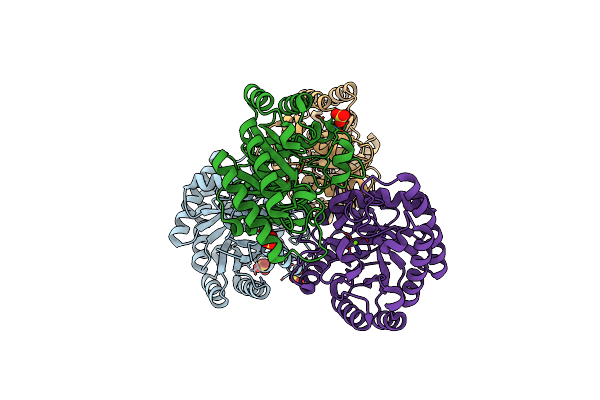 |
Organism: Sinorhizobium fredii ccbau 83666
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2022-08-03 Classification: ISOMERASE Ligands: MG, SO4 |
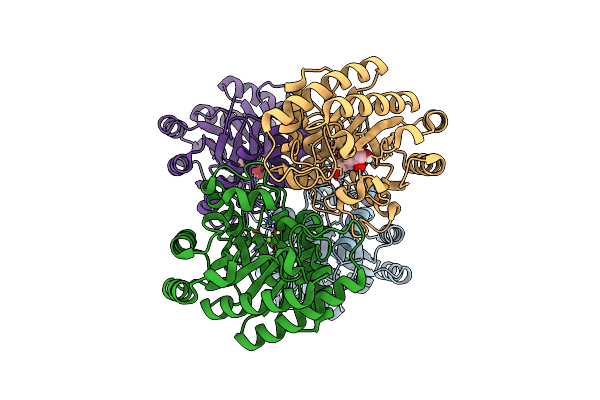 |
Crystal Structures Of D-Allulose 3-Epimerase With D-Fructose From Sinorhizobium Fredii
Organism: Sinorhizobium fredii ccbau 83666
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2022-08-03 Classification: ISOMERASE Ligands: MG, FUD |
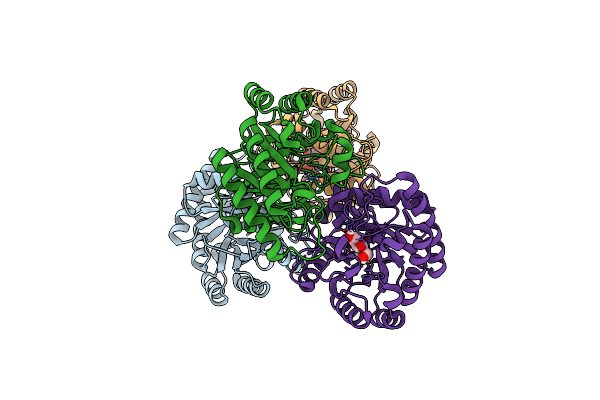 |
Crystal Structures Of D-Allulose 3-Epimerase With D-Tagatose From Sinorhizobium Fredii
Organism: Sinorhizobium fredii ccbau 83666
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2022-08-03 Classification: ISOMERASE Ligands: MG, TAG |
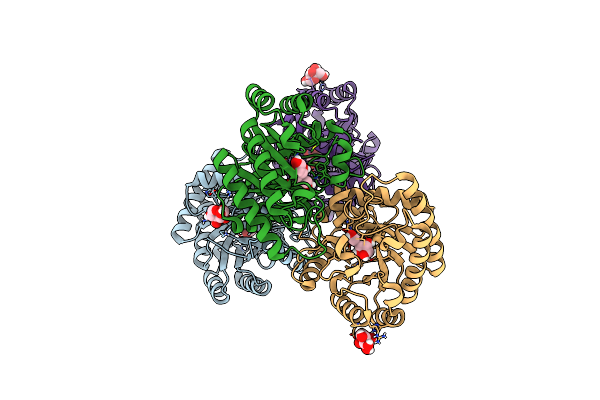 |
Crystal Structures Of D-Allulose 3-Epimerase With D-Sorbose From Sinorhizobium Fredii
Organism: Sinorhizobium fredii ccbau 83666
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-08-03 Classification: ISOMERASE Ligands: SDD, HVC, MG |
 |
Crystal Structures Of D-Allulose 3-Epimerase With D-Allulose From Sinorhizobium Fredii
Organism: Sinorhizobium fredii ccbau 83666
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-08-03 Classification: ISOMERASE Ligands: MG, PSJ |
 |
Organism: Scheffersomyces stipitis (strain atcc 58785 / cbs 6054 / nbrc 10063 / nrrl y-11545)
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2021-05-05 Classification: SPLICING |

