Search Count: 166
All
Selected
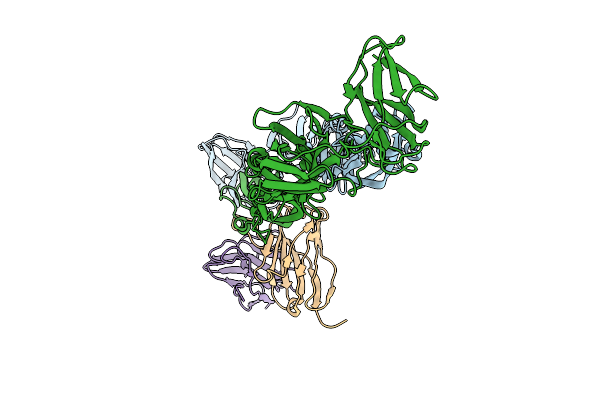 |
Helical Reconstruction Of Denv2 Thsti/Trc/01 Tubular Particles Bound With D14.F25.S02 Fab
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
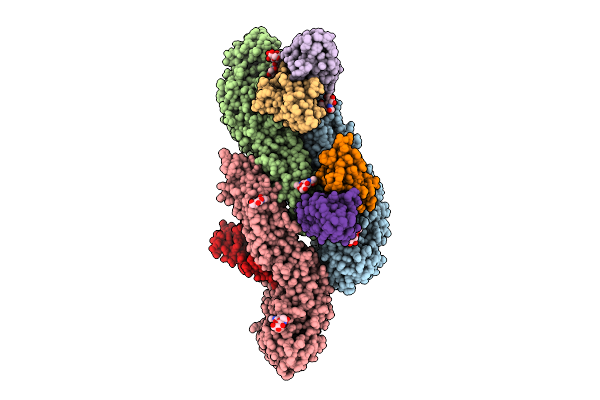 |
Cryo-Em Structure Of Dengue Virus Serotype2 Us/Bid-V594/2006 Strain Bound With J9 Fab.
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: VIRUS Ligands: NAG |
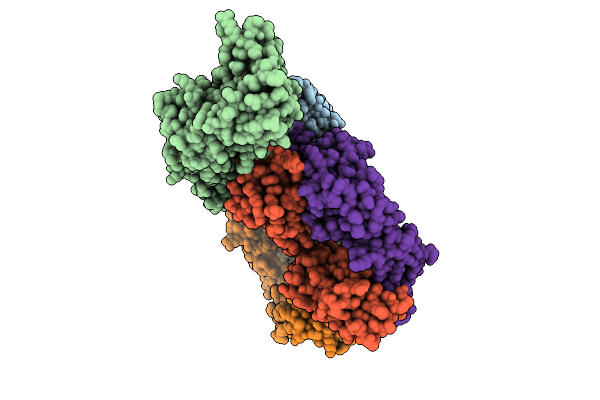 |
Helical Reconstruction Of Denv2 Us/Bid-V594/2006 Tubular Particles Bound With J9 Fab
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
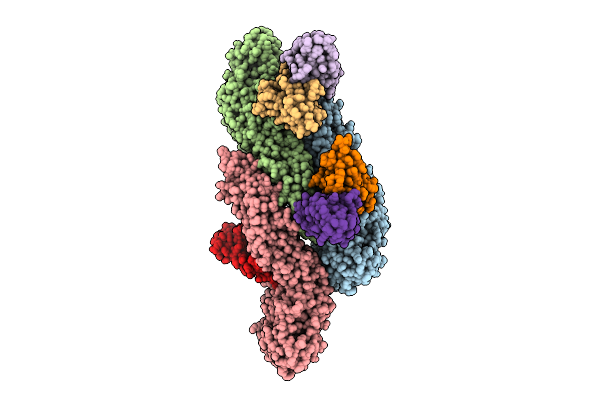 |
Cryo-Em Structure Of Dengue Virus Serotype2 Thsti/Trc/01 Strain Bound With J9 Fab
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: VIRUS |
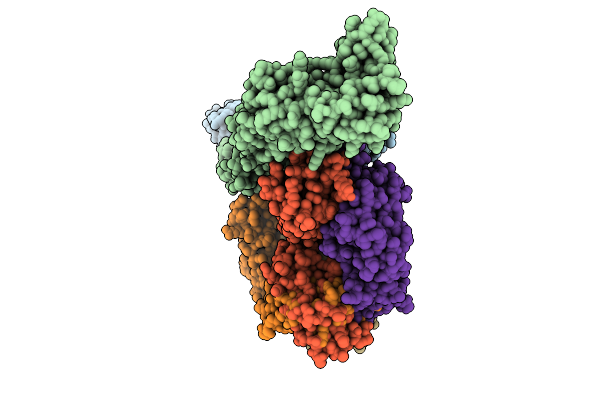 |
Helical Reconstruction Of Denv2 Thsti/Trc/01 Tubular Particles Bound With J9 Fab
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
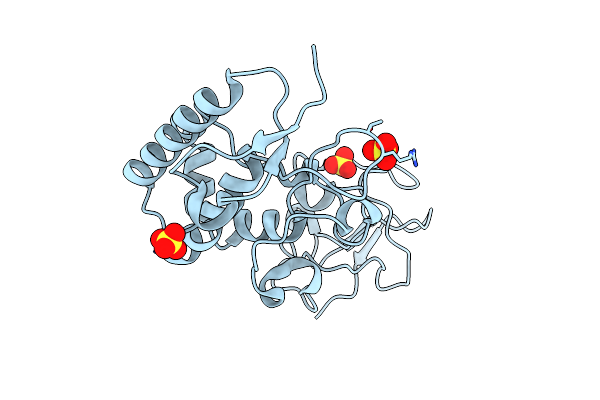 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.48 Å Release Date: 2024-07-10 Classification: SIGNALING PROTEIN Ligands: SO4 |
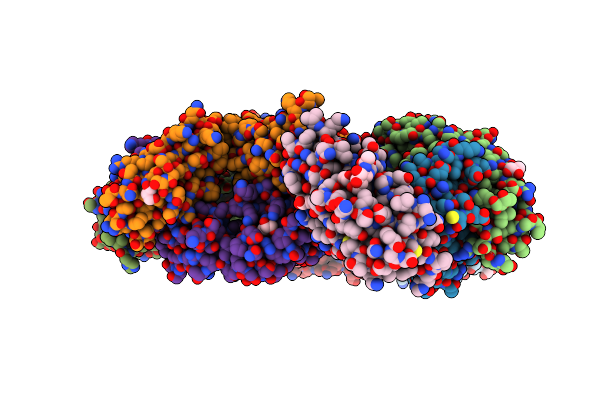 |
Crystal Structure Of The Sars-Cov-2 Receptor Binding Domain In Complex With Neutralizing Antibody Wrair-5021
Organism: Severe acute respiratory syndrome coronavirus 2, Macaca mulatta
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2024-01-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: GOL, NAG, MLI |
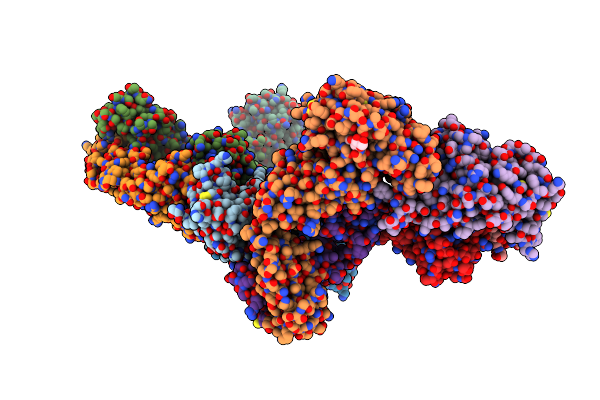 |
Crystal Structure Of Sars-Cov-2 Receptor Binding Domain In Complex With Neutralizing Antibody Wrair-5001
Organism: Severe acute respiratory syndrome coronavirus 2, Macaca mulatta
Method: X-RAY DIFFRACTION Resolution:4.20 Å Release Date: 2024-01-17 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, GOL |
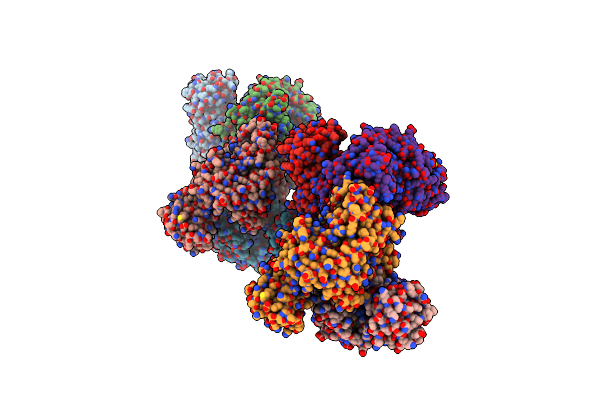 |
Organism: Oryctolagus cuniculus
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2023-05-24 Classification: CYTOSOLIC PROTEIN Ligands: MG, NA, TRS, GOL |
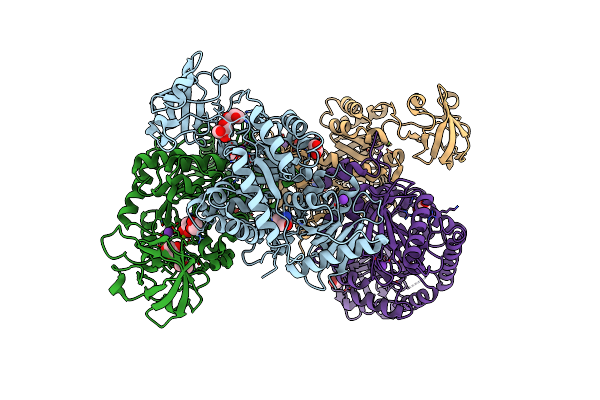 |
Rabbit Muscle Pyruvate Kinase In Complex With Magnesium, Potassium And Pyruvate
Organism: Oryctolagus cuniculus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2023-05-24 Classification: CYTOSOLIC PROTEIN Ligands: GOL, EDO, K, MG, PYR, TRS |
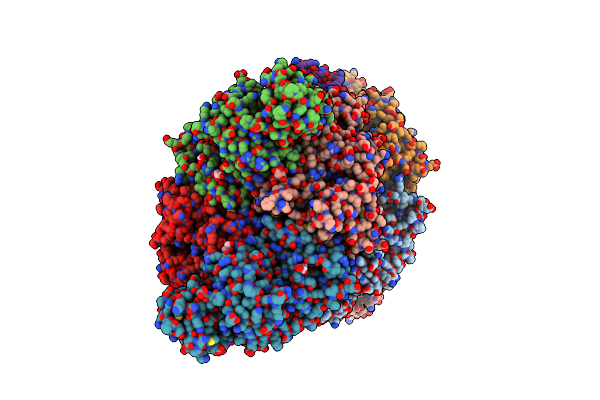 |
Complex Of Rabbit Muscle Pyruvate Kinase With Adp And The Phosphonate Analogue Of Pep Mimicking The Michaelis Complex.
Organism: Oryctolagus cuniculus
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2023-05-24 Classification: CYTOSOLIC PROTEIN Ligands: ADP, SIN, MN, K, ALA, GOL, GZ3 |
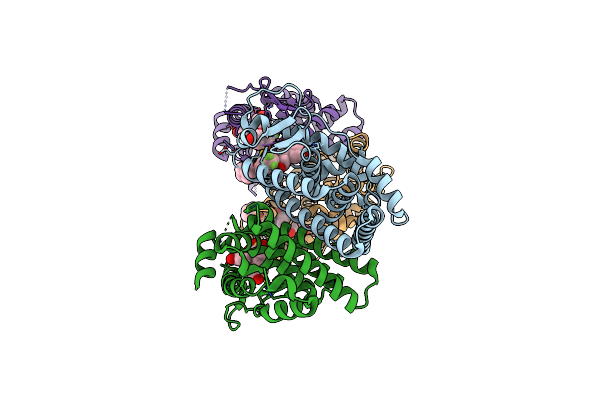 |
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-19
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2021-09-22 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7AI, CL |
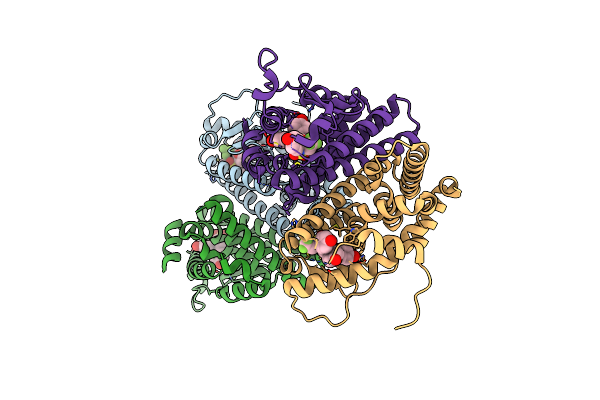 |
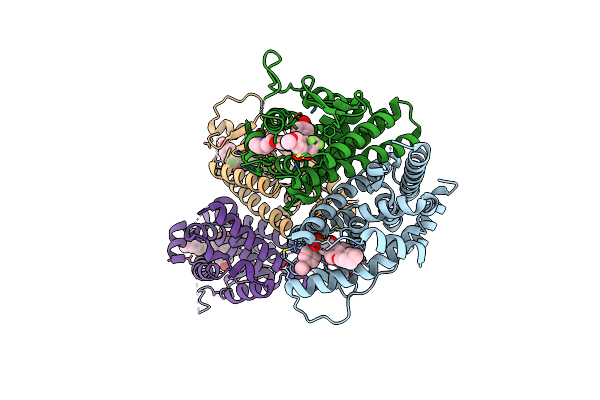
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-30
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2021-09-22 Classification: TRANSCRIPTION/TRANSCRIPTION Inhibitor Ligands: 73I |
 |
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-16
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2021-09-22 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7EI |
 |
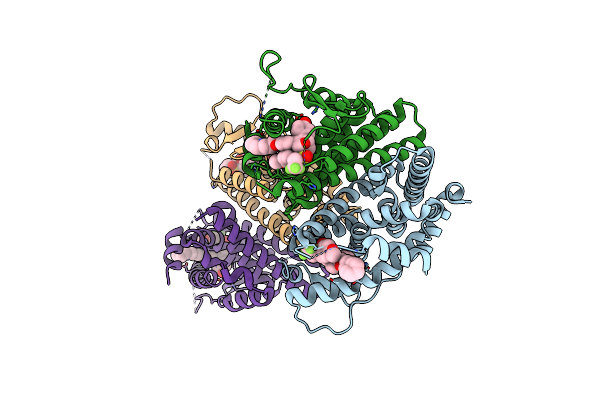
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-20
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: L84, CL |
 |
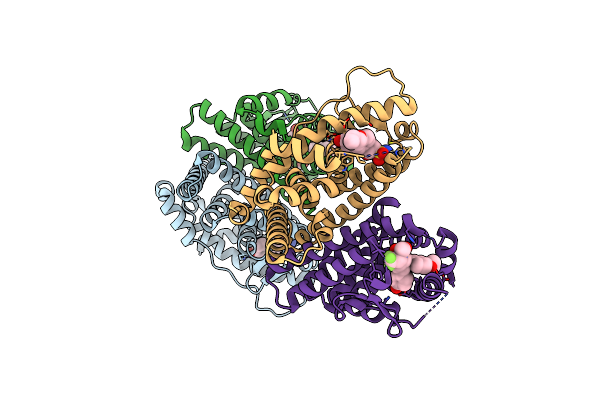
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-30
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 77I |
 |
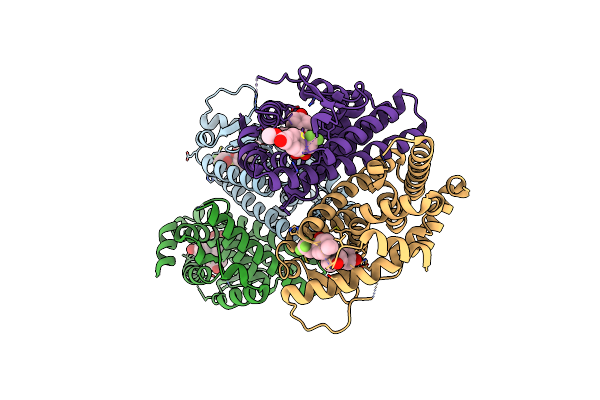
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-18
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7I9 |
 |
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-21
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7Q5 |
 |
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-23
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7I5 |
 |
Crystal Structure Of The Er-Alpha Ligand-Binding Domain (L372S, L536S) In Complex With Dmeri-29
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2021-09-08 Classification: TRANSCRIPTION/TRANSCRIPTION INHIBITOR Ligands: 7OR, TYR, SER, CYS |

