Search Count: 83
All
Selected
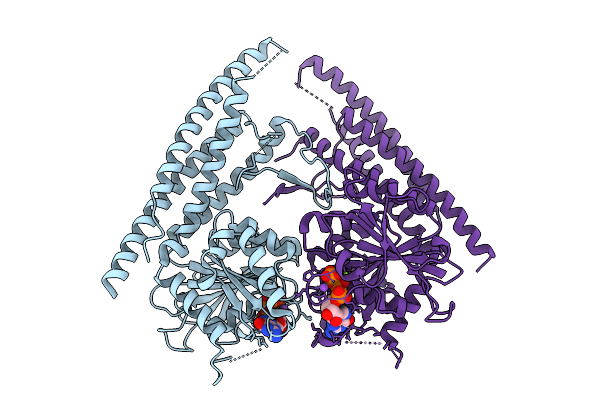 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.73 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: GTP, MG, NA |
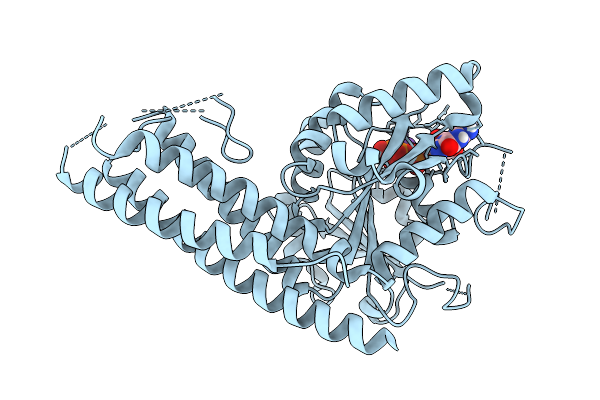 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.81 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: GTP, MG, K |
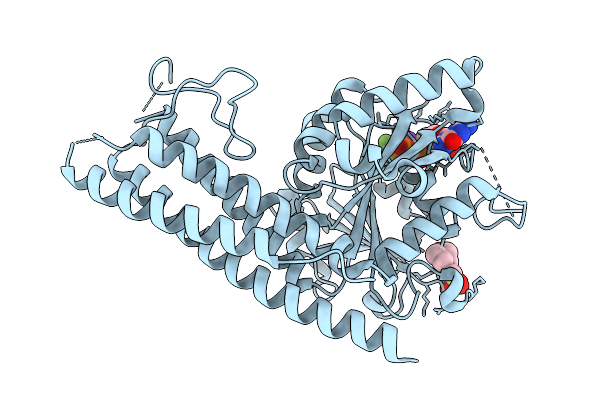 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2025-08-27 Classification: HYDROLASE Ligands: GDP, BEF, MG, K, MES |
 |
Organism: Trypanosoma brucei brucei (strain 927/4 gutat10.1)
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-05-31 Classification: TRANSCRIPTION |
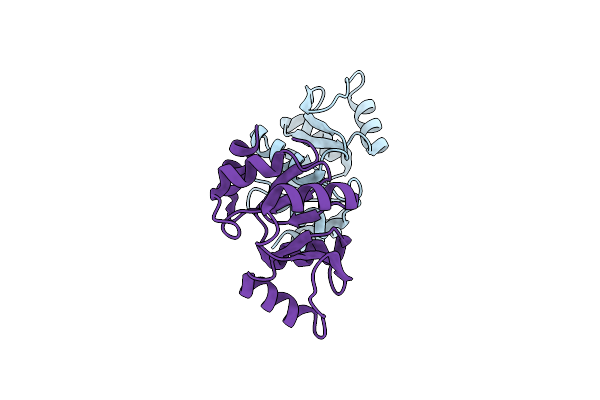 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-11-23 Classification: VIRAL PROTEIN |
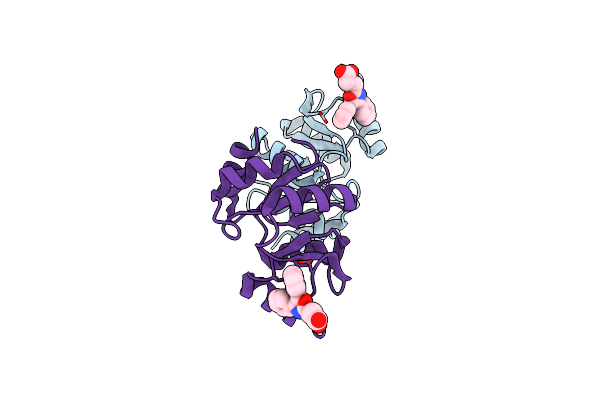 |
Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain 3 (Sars-Unique Domain-M) In Complex With Oxaprozin
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2022-11-23 Classification: VIRAL PROTEIN Ligands: BJ6 |
 |
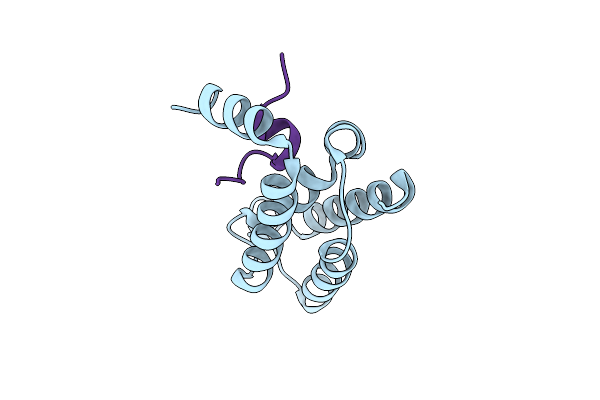
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.76 Å Release Date: 2022-10-19 Classification: IMMUNE SYSTEM Ligands: EDO, NAG |
 |
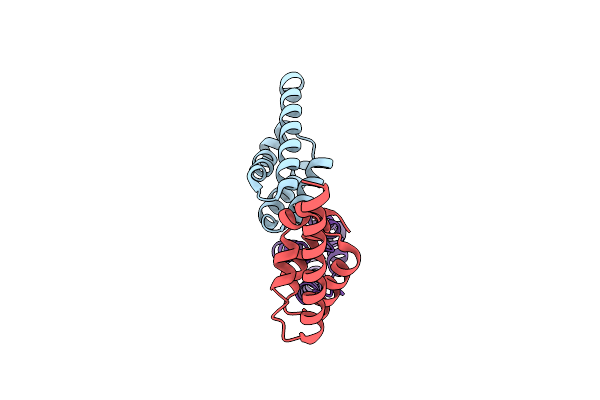
Organism: Trypanosoma brucei brucei treu927
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-07-27 Classification: TRANSCRIPTION |
 |
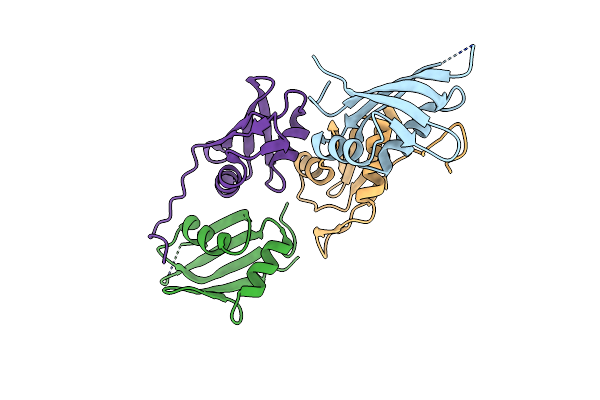
Organism: Saccharomyces cerevisiae s288c
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2022-07-13 Classification: TRANSCRIPTION |
 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-08-25 Classification: PROTEIN BINDING |
 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2021-08-25 Classification: PROTEIN BINDING Ligands: SO4 |
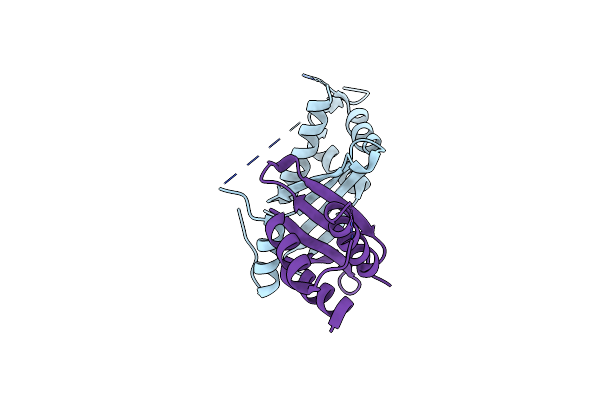 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2021-08-25 Classification: PEPTIDE BINDING PROTEIN |
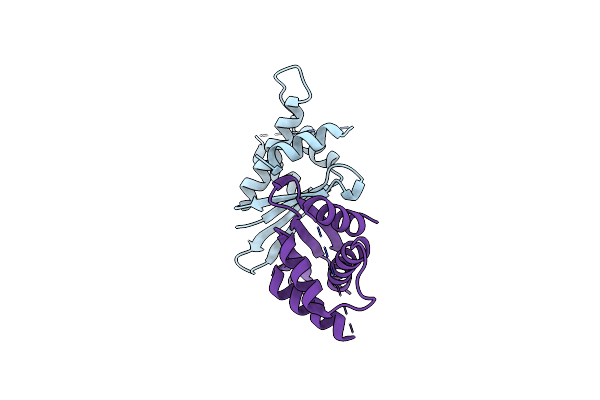 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2021-08-25 Classification: PEPTIDE BINDING PROTEIN |
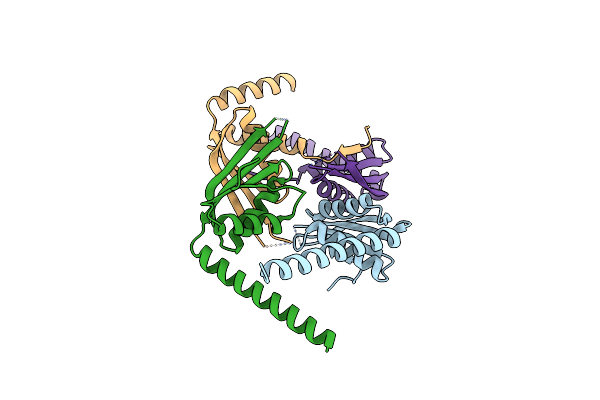 |
Organism: Trypanosoma brucei
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2021-06-23 Classification: RNA BINDING PROTEIN |
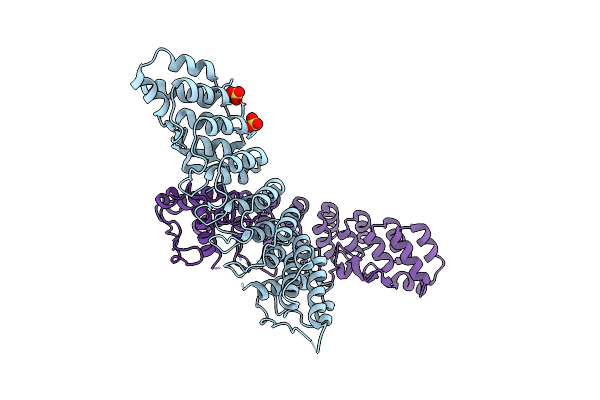 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING Ligands: SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING Ligands: SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING Ligands: SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING Ligands: SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2020-10-21 Classification: PROTEIN BINDING |

