Search Count: 1,001
All
Selected
 |
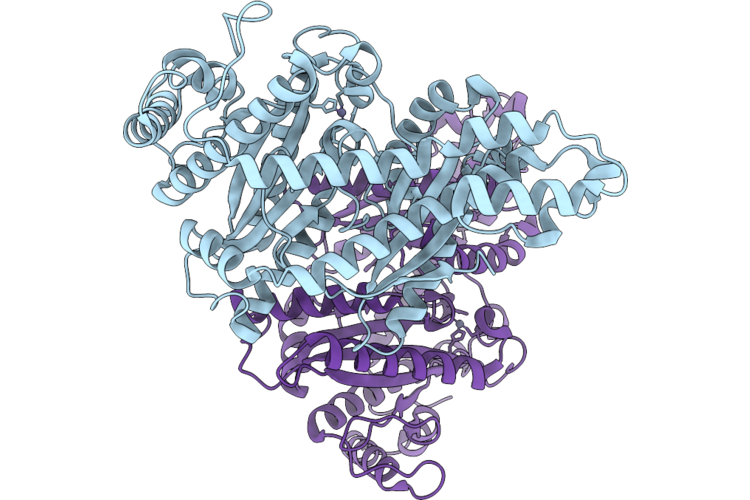
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Major State, Inactive Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.12 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
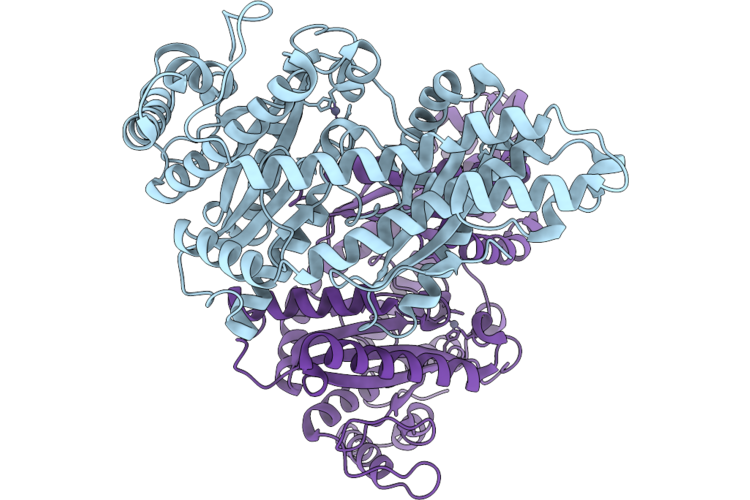
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.08 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 1
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.18 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca C283A/C284A Inactive Mutant, Dimer, State 2
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.22 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
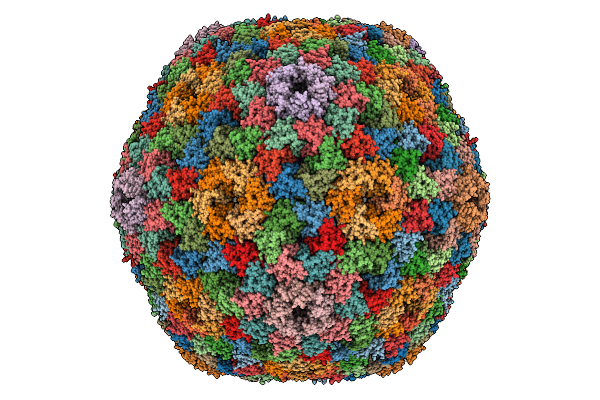 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
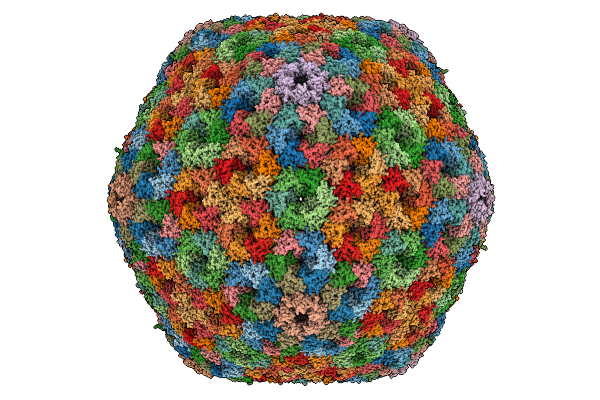 |
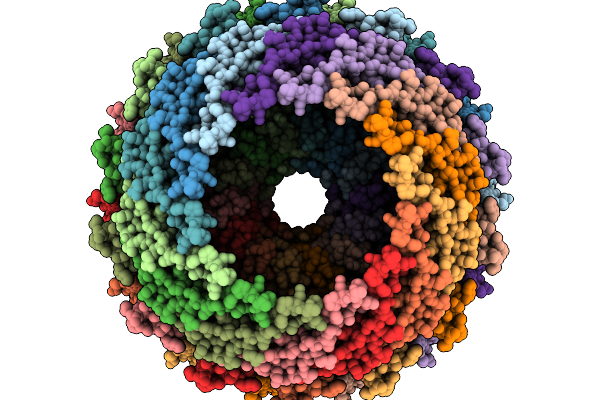
Cryo-Em Structure Of Carboxysomal Mid-Shell: T = 16 Shell Under C1 Symmetry.
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2025-12-31 Classification: STRUCTURAL PROTEIN |
 |
Cryo-Em Structure Of The Type Ii Secretion System Protein From Acidithiobacillus Caldus
Organism: Acidithiobacillus caldus (strain sm-1)
Method: ELECTRON MICROSCOPY Resolution:2.39 Å Release Date: 2025-11-19 Classification: TOXIN |
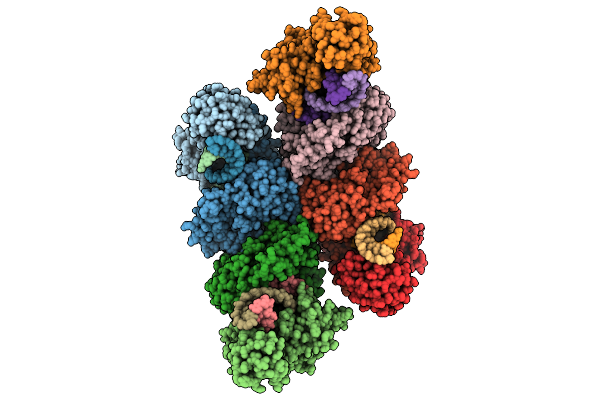 |
Cryo-Em Structure Of Filament Form Acidithiobacillus Caldus (Aca) Short Prokaryotic Argonautes, Hnh-Associated (Sparha) With Grna And Tdna
Organism: Acidithiobacillus caldus
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: IMMUNE SYSTEM/DNA/RNA Ligands: MN |
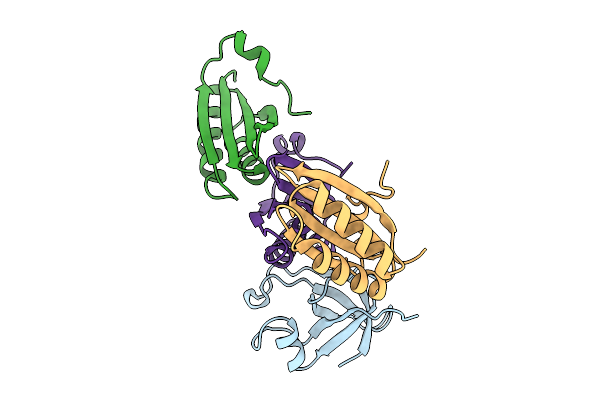 |
Cryo-Em Structure Of Halothiobacillus Neapolitanus Alpha-Carboxysome T=4 Mini-Shell Containing Ctd Truncated Mutant Of Csosca
Organism: Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: STRUCTURAL PROTEIN |
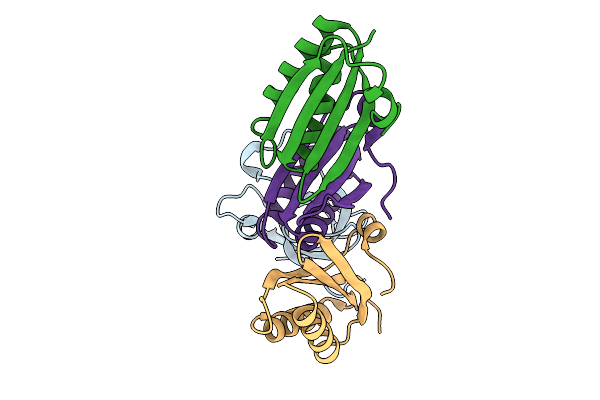 |
Cryo-Em Structure Of Alpha-Carboxysome T=4 Mini-Shell Containing Ctd Only Mutant Of Csosca
Organism: Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2025-10-08 Classification: STRUCTURAL PROTEIN |
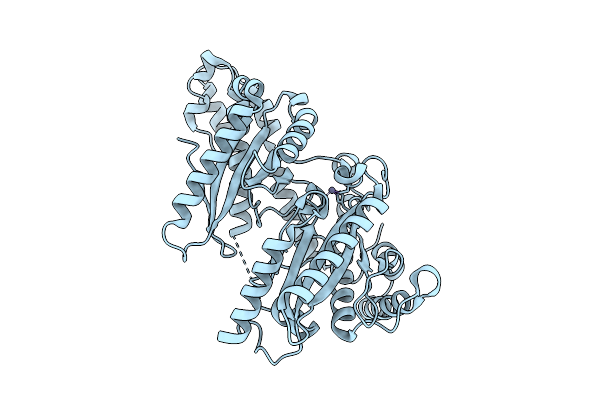 |
Beta Carbonic Anhydrase Csosca From The Halothiobacillus Neapolitanus Alpha-Carboxysome
Organism: Streptococcus sp. 'group g', Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: LYASE Ligands: ZN |
 |
Organism: Acidithiobacillus
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2025-05-14 Classification: LYASE Ligands: PLP |
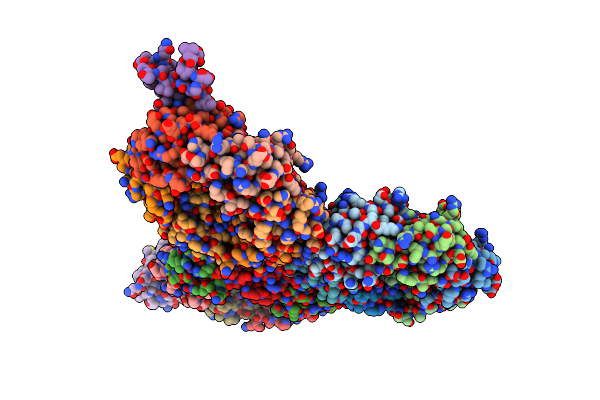 |
Cryo-Em Structure Of Carboxysomal Midi-Shell:Icosahedral Assembly From Csos4A/4B/1A/1B/1C/1D And Csos2 C-Terminal Co-Expression (T = 19)
Organism: Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: STRUCTURAL PROTEIN |
 |
Cryo-Em Structure Of Carboxysomal Midi-Shell: Icosahedral Assembly From Csos4A/4B/1A/1B/1C/1D And Csos2 C-Terminal Co-Expression (T = 16)
Organism: Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: STRUCTURAL PROTEIN |
 |
Cryo-Em Structure Of Carboxysomal Midi-Shell: Icosahedral Assembly From Csos4A/4B/1A/1B/1C/1D And Csos2 C-Terminal Co-Expression (T = 9)
Organism: Halothiobacillus neapolitanus
Method: ELECTRON MICROSCOPY Release Date: 2024-12-11 Classification: STRUCTURAL PROTEIN |

