Search Count: 5,597
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: A1EIW |
 |
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: ATP, MG, A1E61 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.47 Å Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: A1EIX |
 |
Apo Multidrug Resistance-Associated Protein 2 In Complex With Amp-Pnp In Rest State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: MG, ANP |
 |
Multidrug Resistance-Associated Protein 2 In Complex With Amp-Pnp In Active State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: MG, ANP |
 |
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: MG |
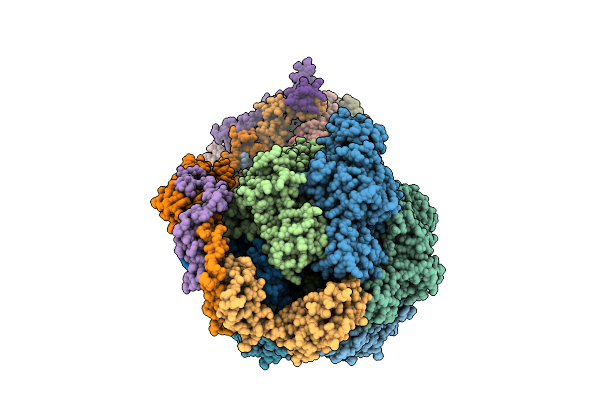 |
Cryo-Em Structure Of Fof1-Atpase Monomer State 1 On The Bovine Heart Submitochondrial Particles (Fof1-1)
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2026-04-01 Classification: MOTOR PROTEIN Ligands: ATP, MG, ADP |
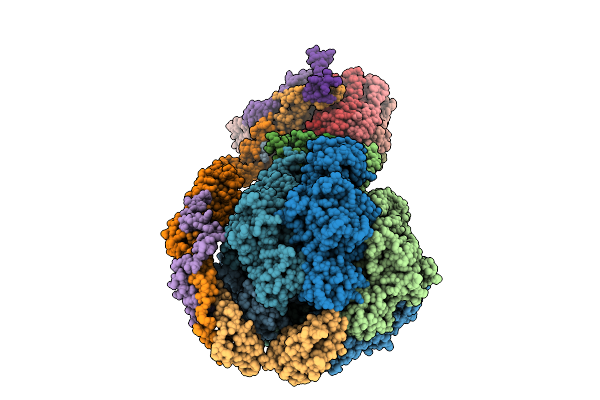 |
Cryo-Em Structure Of Fof1-Atpase Monomer State 3 On The Bovine Heart Submitochondrial Particles (Fof1-2)
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2026-04-01 Classification: MOTOR PROTEIN |
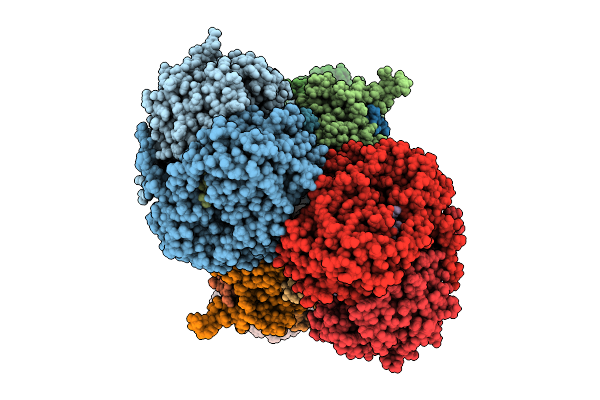 |
Cryo-Em Structure Of Complex Iii On The Bovine Heart Submitochondrial Particles, Iii-1
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MOTOR PROTEIN Ligands: HEM, HEC, FES |
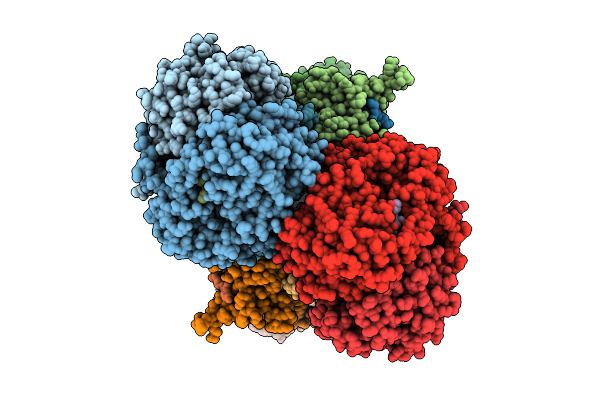 |
Cryo-Em Structure Of Complex Iii On The Bovine Heart Submitochondrial Particles, Iii-2
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, HEC, FES |
 |
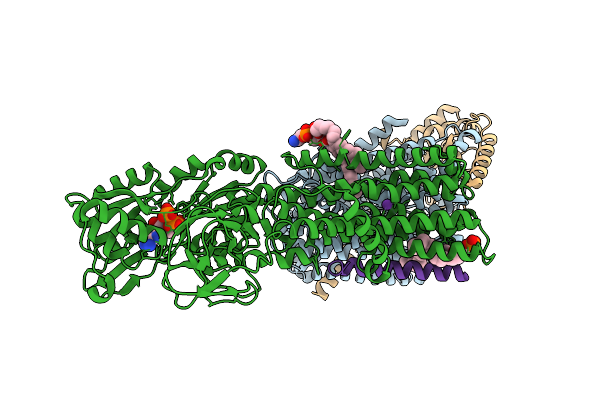
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1 State In Lipid Nanodisc
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.34 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1-Atp State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, ATP |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, MG |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.58 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2 State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
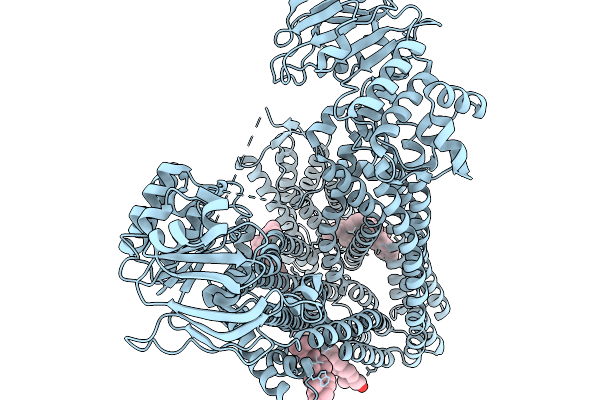
Structure Of The Ybjp Lipoprotein Bound To The Macab-Tolc Tripartite Efflux Pump
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.48 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN Ligands: CL, Z41 |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: CLR |
 |
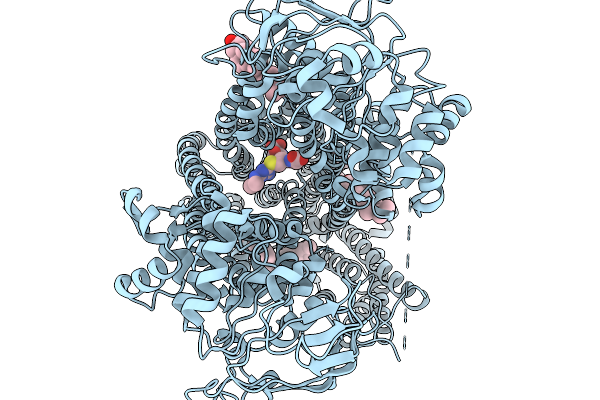
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: CLR, R16 |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: TRANSPORT PROTEIN Ligands: ATA, CLR |

