Search Count: 265
All
Selected
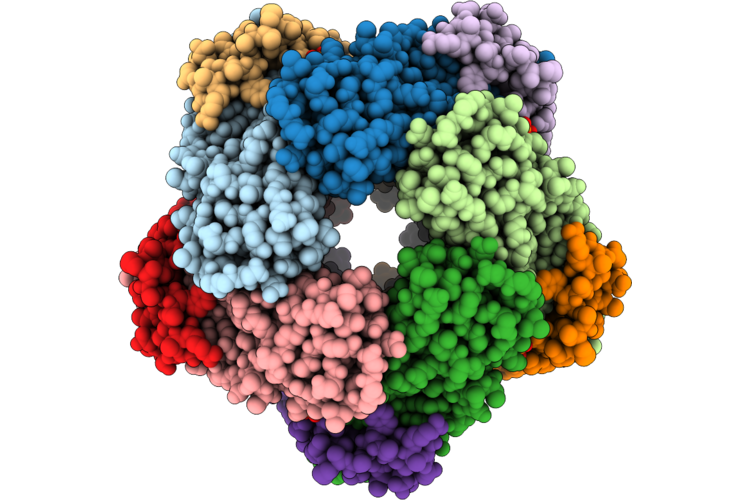 |
Organism: Streptomyces mobaraensis
Method: ELECTRON MICROSCOPY Resolution:2.01 Å Release Date: 2026-04-22 Classification: BIOSYNTHETIC PROTEIN Ligands: ZN, GTP |
 |
Crystal Structure Of 1L-Myo-Inositol 1-Phosphate Synthase 1 From Oryza Sativa
Organism: Oryza sativa
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-02-04 Classification: ISOMERASE Ligands: KPG, NAI, EDO, K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: PIO, A1LVR |
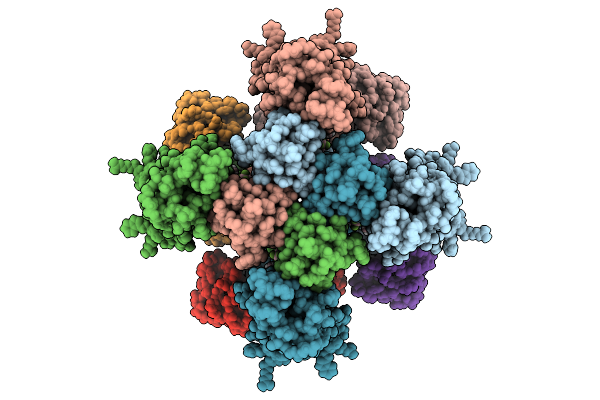 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: PIO, A1LWZ |
 |
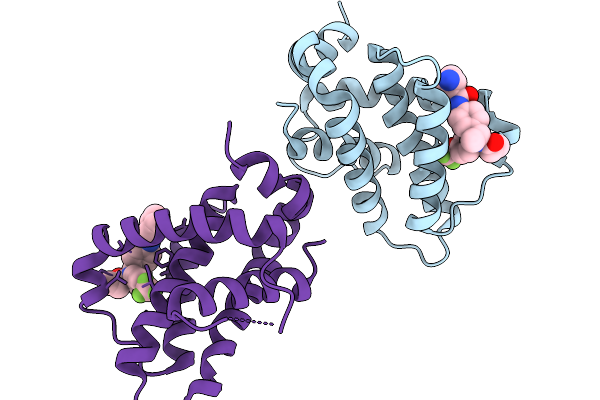
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1LVR |
 |
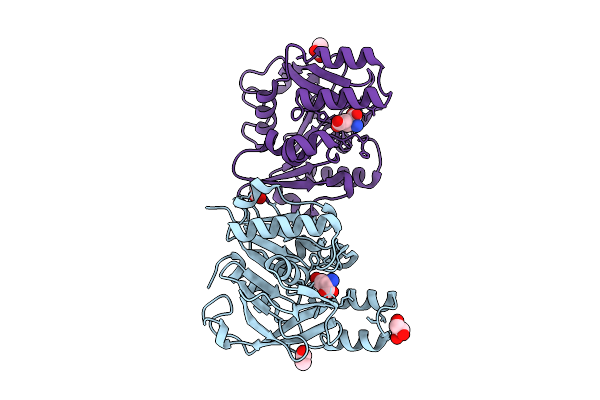
Structure Of Bfl1 In Complex With A Covalent Inhibitor, Alternative Series, Cmpd25
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-01-14 Classification: APOPTOSIS Ligands: A1JL2 |
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: GLU, ACT, GOL |
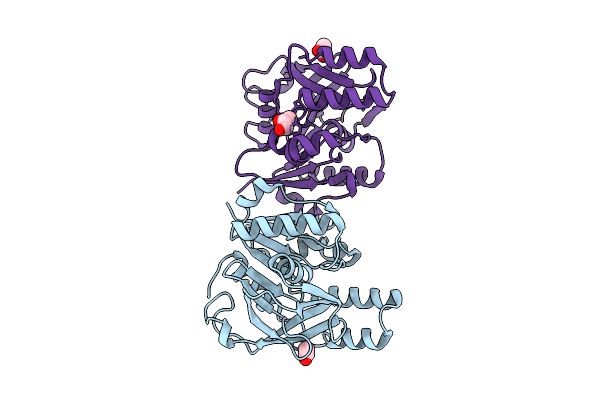 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: ACT, GOL |
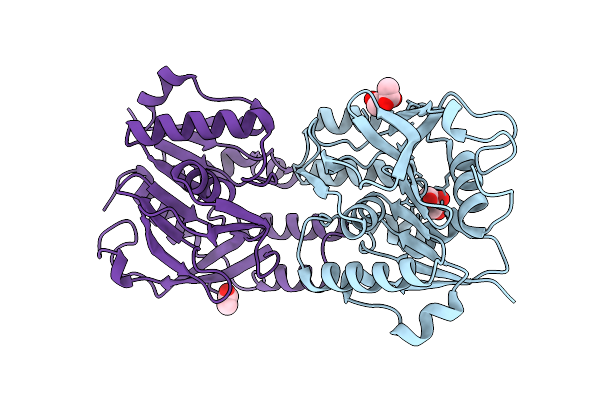 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: ACT, GOL |
 |
Gamma-Glutamyl Peptidase 1 From Arabidopsis Thaliana (H192N Gamma-Glu Intermediate)
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: GLU, ACT, GOL |
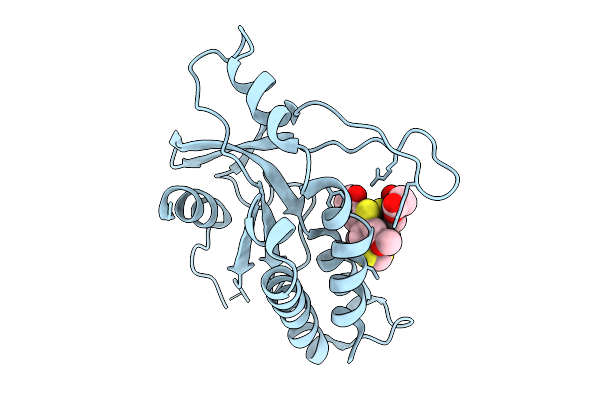 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2025-09-24 Classification: SIGNALING PROTEIN Ligands: A1JA0 |
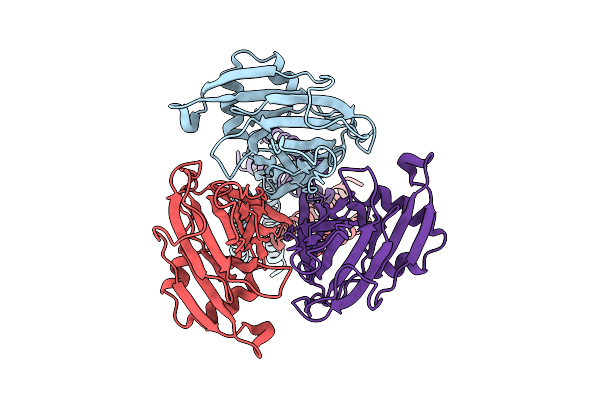 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.43 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: DXC |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: IZ8 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: CLR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN Ligands: PIO |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN Ligands: 9MF, PIO |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: MEMBRANE PROTEIN Ligands: PIO, 9MF |
 |
Crystal Structure Of The Vhl-Eloc-Elob Complex With A Covalent Compound Bound To C77 Of Vhl.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-03-26 Classification: LIGASE Ligands: 3JF, A1IMD |

