Search Count: 36
All
Selected
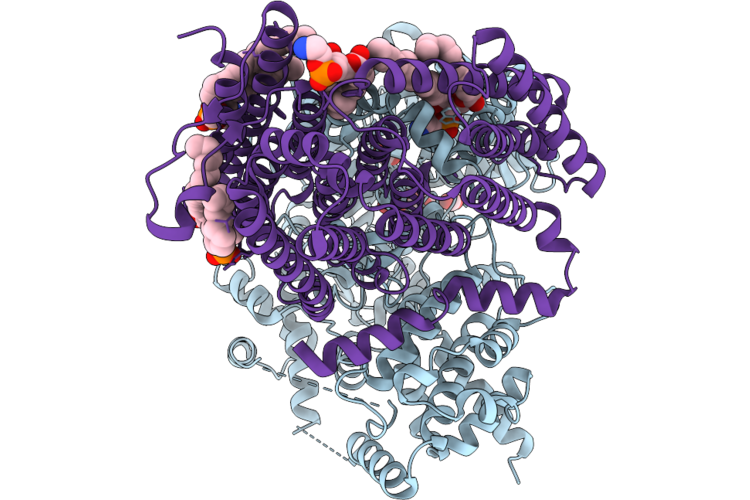 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
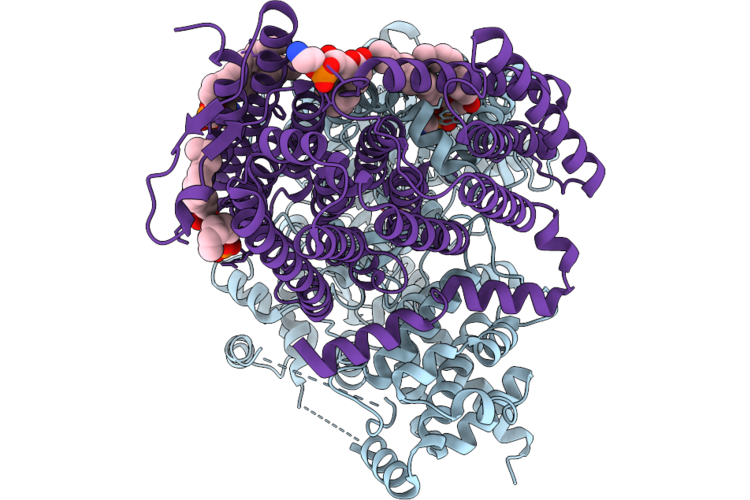 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, BCT, CO2, 6OU |
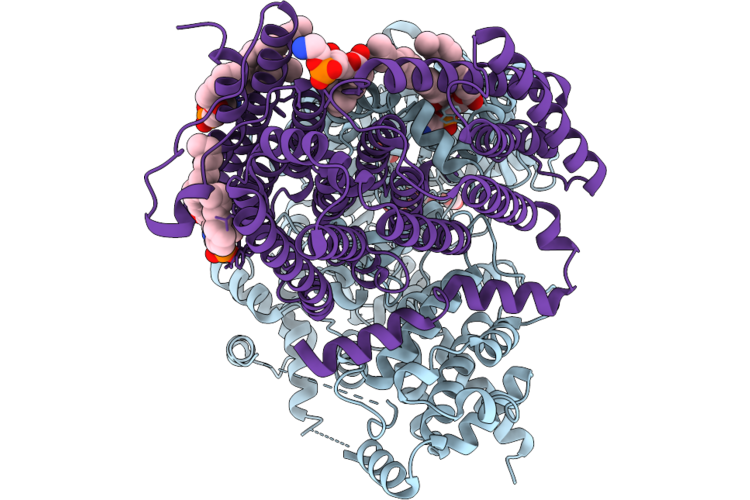 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
 |
Organism: Clostridium pasteurianum
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: 402, SF4, FES, MG, GOL, CL |
 |
Organism: Clostridium pasteurianum
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: 402, SF4, FES, MG, GOL, CL |
 |
Organism: Clostridium pasteurianum
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: 402, SF4, FES, GOL, MG, CL |
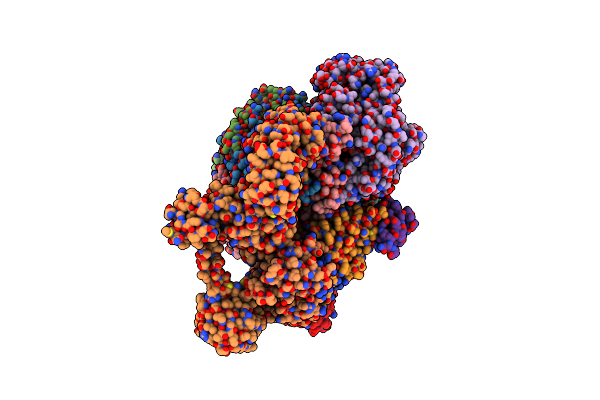 |
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: SHT, TP7, COM, F43, S5Q, ZN, ATP, MG |
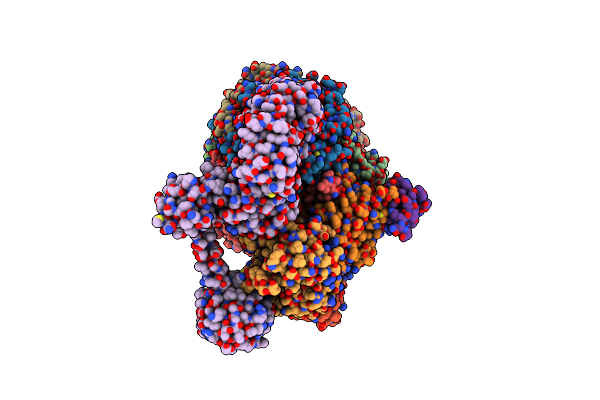 |
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: F43, SHT, COM, TP7, S5Q |
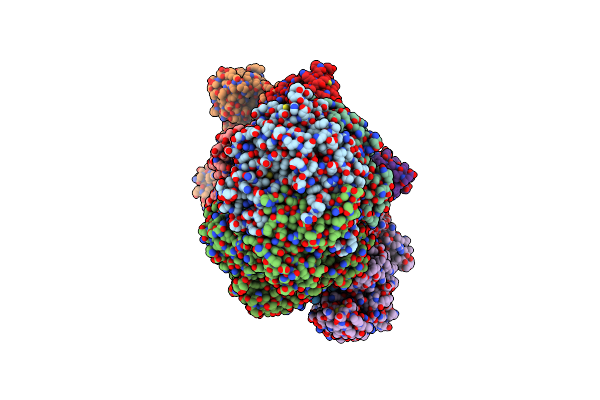 |
Methyl-Coenzyme M Reductase Activation Complex Binding To The A2 Component After Incubation With Atp
Organism: Methanococcus maripaludis
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: OXIDOREDUCTASE Ligands: F43, COM, TP7, SHT, S5Q, ZN, ATP, MG |
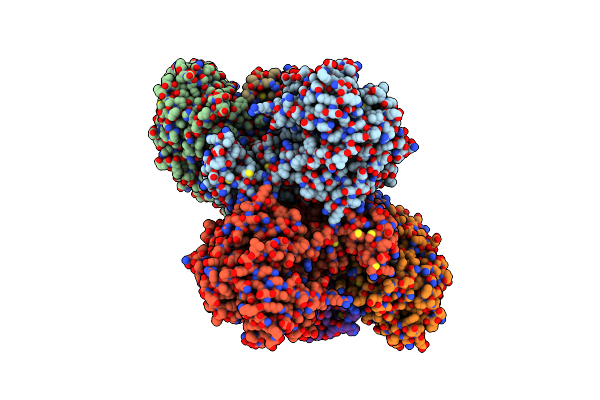 |
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2023-02-22 Classification: ELECTRON TRANSPORT Ligands: SF4, FES, ZN, FMN, NAD |
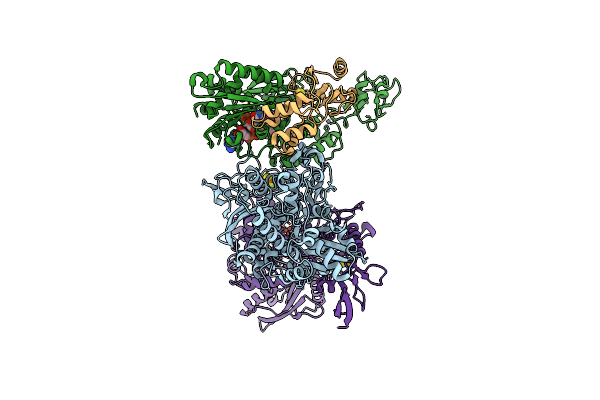 |
Cryo-Em Structure Of The Electron Bifurcating Fe-Fe Hydrogenase Hydabc Complex From Acetobacterium Woodii In The Reduced State
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2023-02-22 Classification: ELECTRON TRANSPORT Ligands: SF4, FES, HC1, ZN, FMN, NAI |
 |
Cryoem Structure Of Electron Bifurcating Fe-Fe Hydrogenase Hydabc Complex A. Woodii In The Oxidised State
Organism: Acetobacterium woodii dsm 1030
Method: ELECTRON MICROSCOPY Release Date: 2023-02-15 Classification: ELECTRON TRANSPORT Ligands: SF4, FES, FMN, ZN |
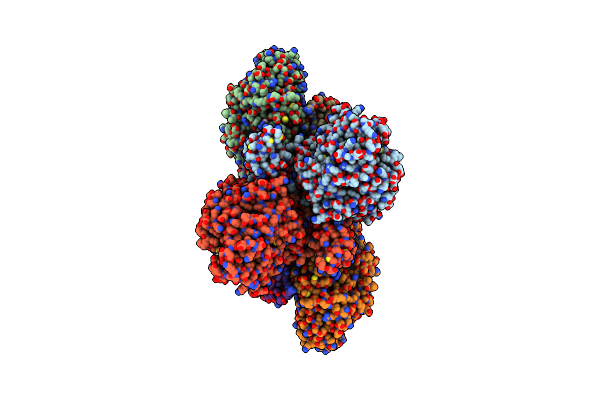 |
Cryo-Em Structure Of The Electron Bifurcating Fe-Fe Hydrogenase Hydabc Complex From Thermoanaerobacter Kivui In The Reduced State
Organism: Thermoanaerobacter kivui
Method: ELECTRON MICROSCOPY Release Date: 2023-02-15 Classification: ELECTRON TRANSPORT Ligands: SF4, HC1, FES, FMN, ZN, NAP |
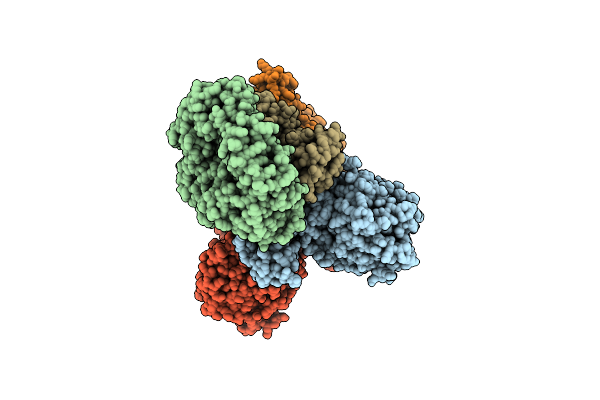 |
Cryo-Em Structure Of The Electron Bifurcating Fe-Fe Hydrogenase Hydabc Complex From Thermoanaerobacter Kivui In The Oxidised State
Organism: Thermoanaerobacter kivui
Method: ELECTRON MICROSCOPY Release Date: 2023-02-15 Classification: ELECTRON TRANSPORT Ligands: SF4, FES, FMN, ZN |
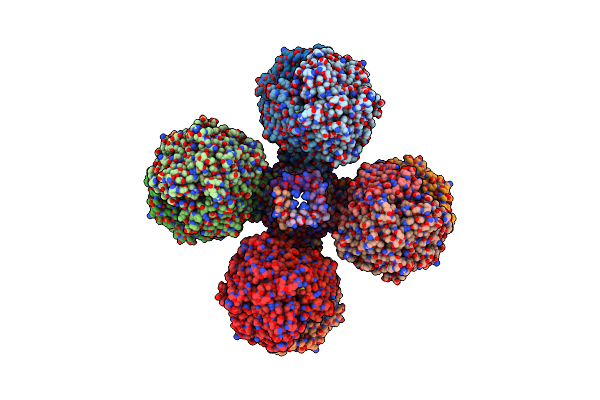 |
The 2.19-Angstrom Cryoem Structure Of The [Nife]-Hydrogenase Huc From Mycobacterium Smegmatis - Complex Minus Stalk
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2023-01-04 Classification: OXIDOREDUCTASE Ligands: 3NI, FCO, MG, MQ9, F3S |
 |
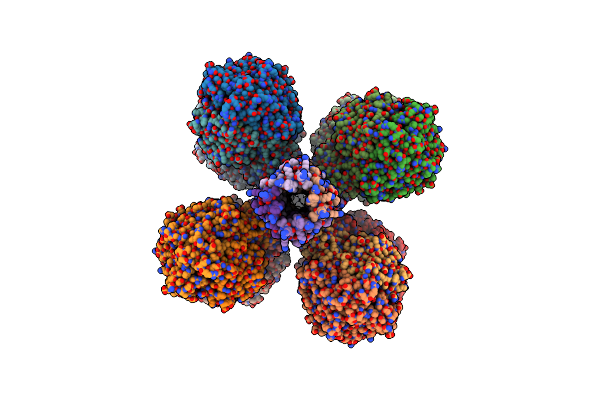
The 1.67 Angstrom Cryoem Structure Of The [Nife]-Hydrogenase Huc From Mycobacterium Smegmatis - Catalytic Dimer (Huc2S2L)
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2023-01-04 Classification: OXIDOREDUCTASE Ligands: 3NI, FCO, OH, MG, VK3, F3S |
 |
The Cryoem Structure Of The [Nife]-Hydrogenase Huc From Mycobacterium Smegmatis - Full Complex Focused Refinement Of Stalk
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2023-01-04 Classification: OXIDOREDUCTASE Ligands: 3NI, FCO, MG, MQ9, F3S |
 |
The 1.52 Angstrom Cryoem Structure Of The [Nife]-Hydrogenase Huc From Mycobacterium Smegmatis - Catalytic Dimer (Huc2S2L)
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2023-01-04 Classification: ELECTRON TRANSPORT Ligands: O, MG, FCO, 3NI, VK3, F3S |
 |
Organism: Clostridium pasteurianum
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2022-12-14 Classification: OXIDOREDUCTASE Ligands: MHX, SF4, FES, GOL, MG, CL |
 |
Organism: Clostridium pasteurianum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2022-12-14 Classification: OXIDOREDUCTASE Ligands: SF4, FES, MG, GOL, VHR, CL |

