Search Count: 931
All
Selected
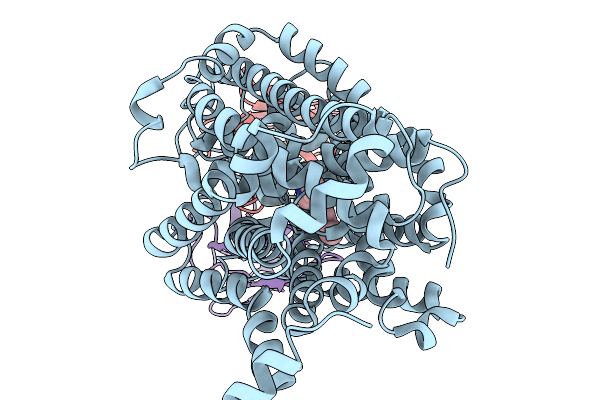 |
Structure Of Human Serotonin Transporter Bound To Small Molecule Zpzd In Lipid Nanodisc And Nacl
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM/INHIBITOR Ligands: CL, A1CIZ |
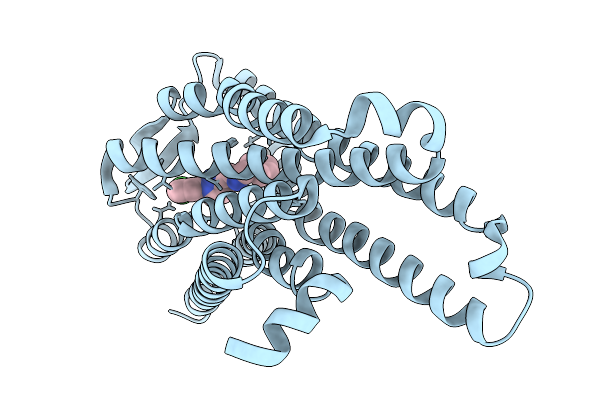 |
Locally-Refined Mu-Opioid Receptor Bound With Novel Compound 0505 (3-[({[(1P)-1-(3-Chlorophenyl)-1H-Pyrazol-3-Yl]Methyl}Amino)Methyl]Phenol)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: A1CIX |
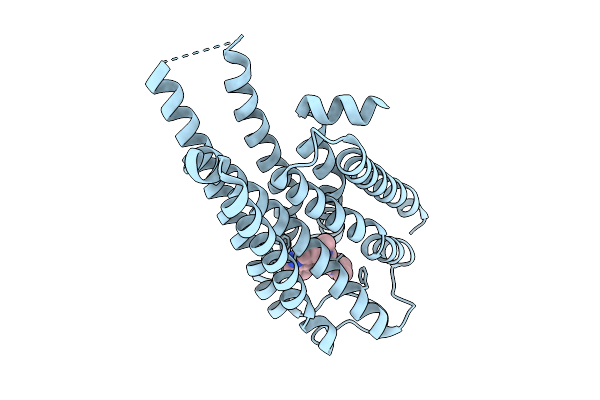 |
Local Refinement Of Drd2 Bound To Lsd In Complex With A Mini-Goa And Scfv16 Obtained By Cryo-Electron Microscopy (Cryoem)
Organism: Escherichia coli, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2025-09-17 Classification: MEMBRANE PROTEIN Ligands: 7LD |
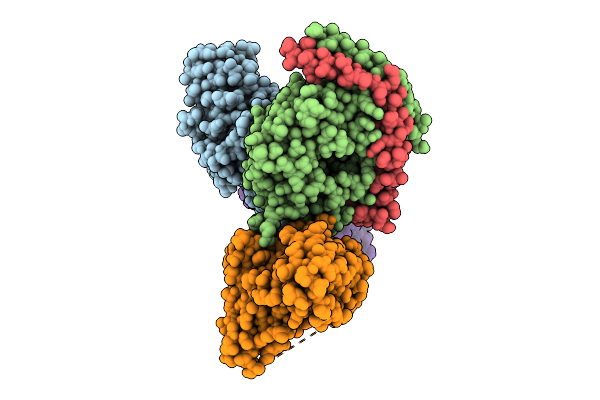 |
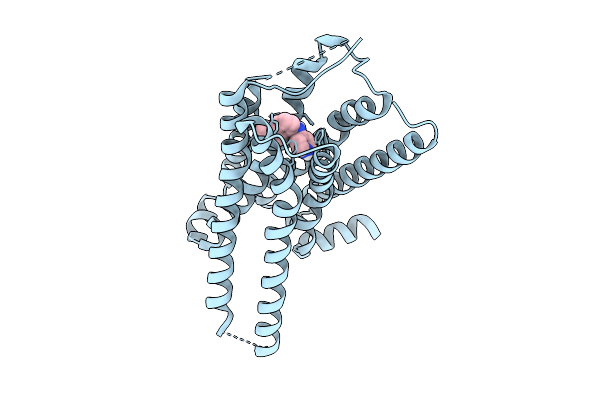
Global Reconstruction Of Drd2 Bound To Lsd In Complex With A Mini-Goa And Scfv16 Obtained By Cryo-Electron Microscopy (Cryoem)
Organism: Homo sapiens, Escherichia coli, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.32 Å Release Date: 2025-09-17 Classification: MEMBRANE PROTEIN Ligands: 7LD |
 |
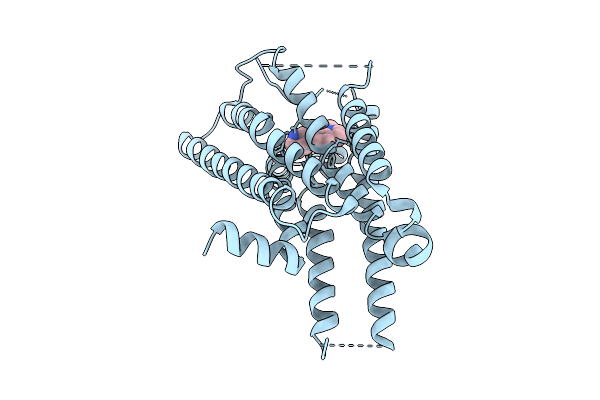
Locally-Refined Structure Of Alpha2A Adrenergic Receptor In Complex With Go Heterotrimer, Scfv16, And N-(5-Methylnaphthalen-1-Yl)Pyridin-4-Amine (Compound 4905)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2025-08-20 Classification: MEMBRANE PROTEIN Ligands: A1CIU |
 |
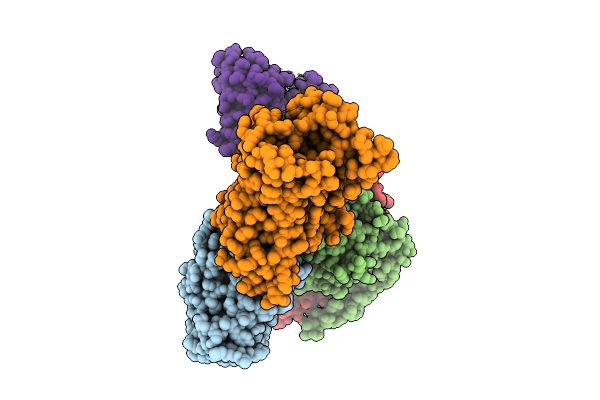
Locally-Refined Structure Of Alpha2A Adrenergic Receptor In Complex With Go Heterotrimer, Scfv16, And Compound Z7149
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2025-08-20 Classification: MEMBRANE PROTEIN Ligands: A1CIZ |
 |
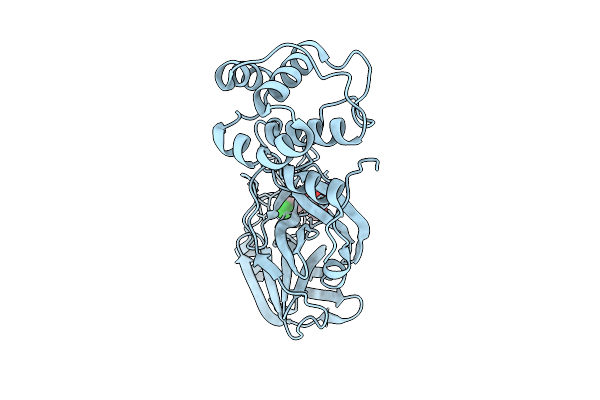
Structure Of Alpha2A Adrenergic Receptor In Complex With Go Heterotrimer, Scfv16, And N-(5-Methylnaphthalen-1-Yl)Pyridin-4-Amine (Compound 4905)
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.74 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: A1CIU |
 |
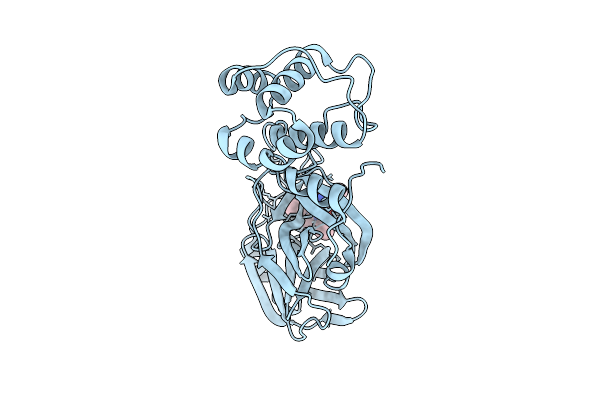
Crystal Structure Of Sars-Cov-2 Main Protease (Mpro)In Complex With Inhibitor Avi-3318
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2025-07-23 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: A1BTM |
 |
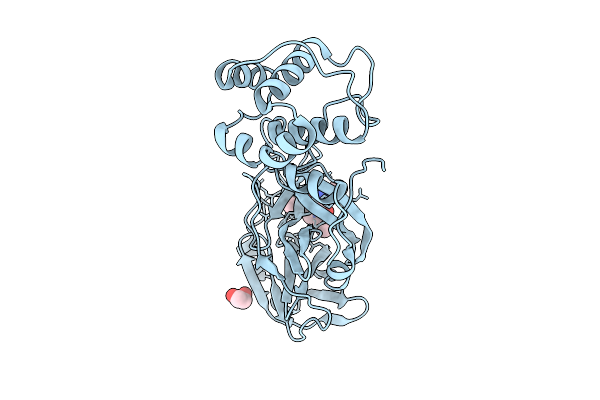
Crystal Structure Of Sars-Cov-2 Main Protease (Mpro) In Complex With Inhibitor Avi-4692
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2025-07-23 Classification: HYDROLASE Ligands: A1BVT, A1BTO |
 |
Crystal Structure Of Sars-Cov-2 Main Protease (Mpro)In Complex With Inhibitor Avi-4516
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-07-23 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: A1BTP, EDO |
 |
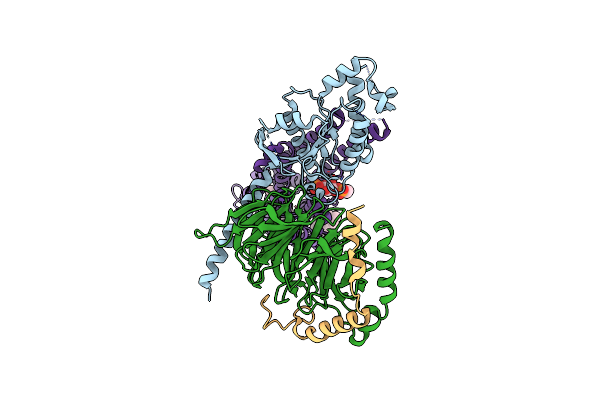
Crystal Structure Of Sars-Cov-2 Main Protease (Mpro) Variant Q192T In Complex With Inhibitor Avi-4303
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2025-07-23 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: EDO, A1BTN, CL |
 |
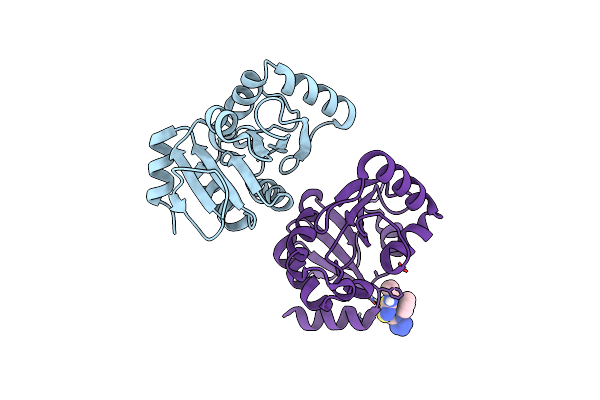
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: MEMBRANE PROTEIN Ligands: CLR, T7M, ZKV |
 |
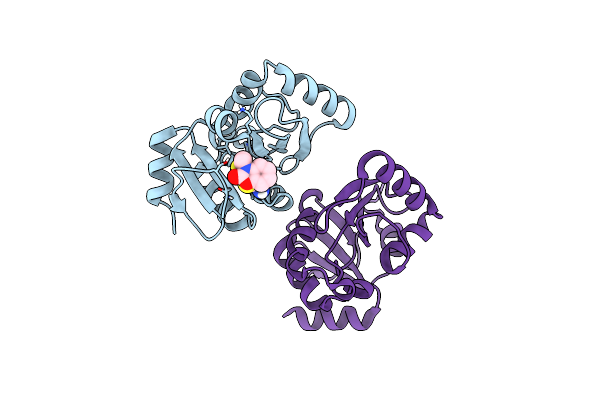
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000286
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A2H |
 |
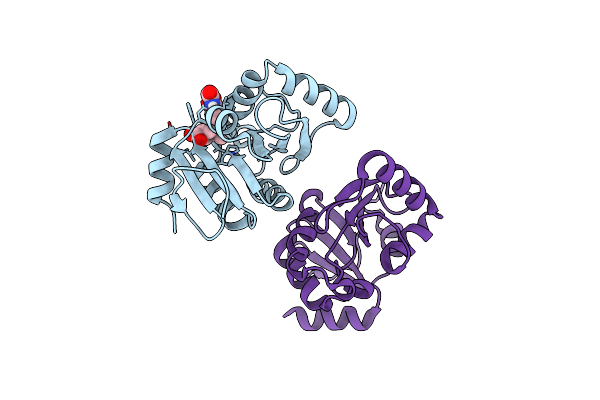
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000296
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A41 |
 |
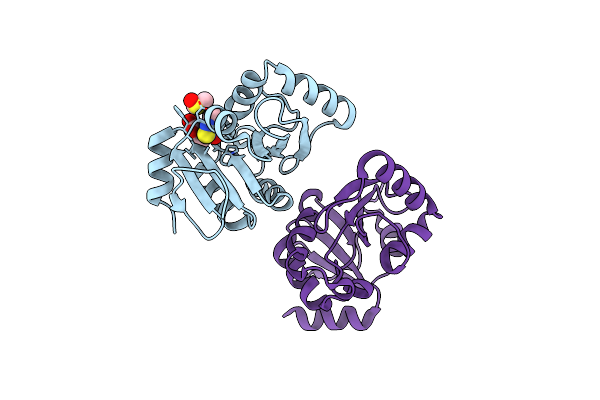
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000303
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A42 |
 |
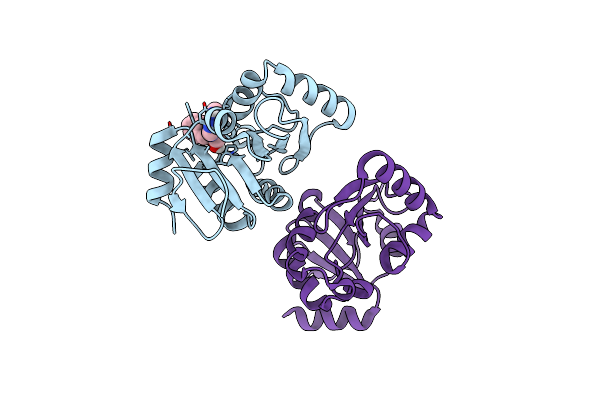
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000002
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: DMS, A1A47 |
 |
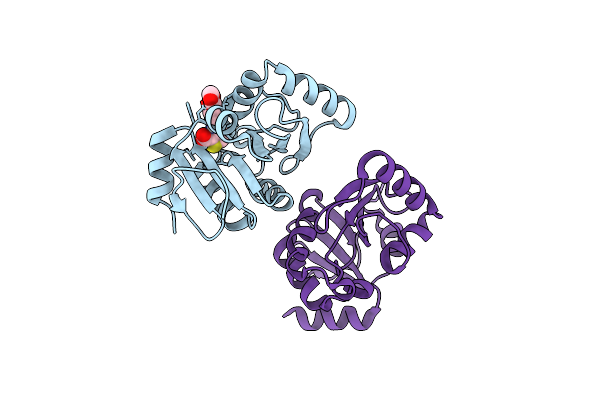
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000313
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A49, A1A5A, CL |
 |
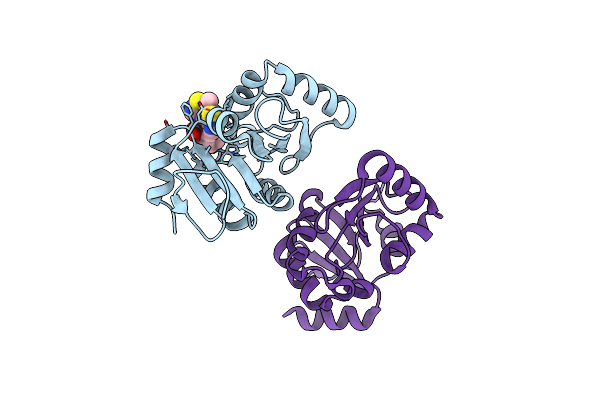
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000316
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A5B |
 |
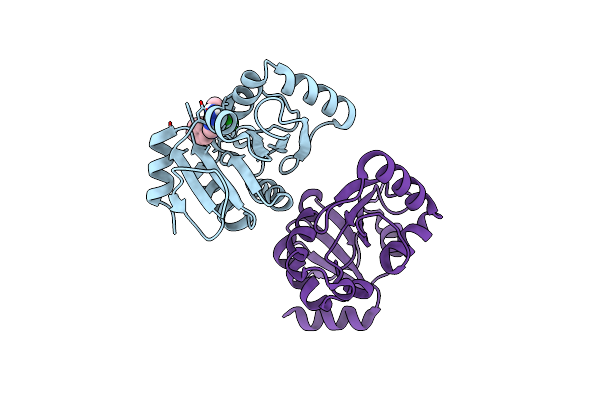
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000317
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A5C |
 |
Pandda Analysis Group Deposition -- Crystal Structure Of Sars-Cov-2 Nsp3 Macrodomain In Complex With Avi-0000321
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2025-06-11 Classification: VIRAL PROTEIN, Hydrolase Ligands: A1A19, CL |

