Search Count: 177
 |
Cryo-Em Structure Of Self-Assembled Zymomonas Mobilis Levansucrase Nanotube
Organism: Zymomonas mobilis subsp. mobilis atcc 10988
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: TRANSFERASE |
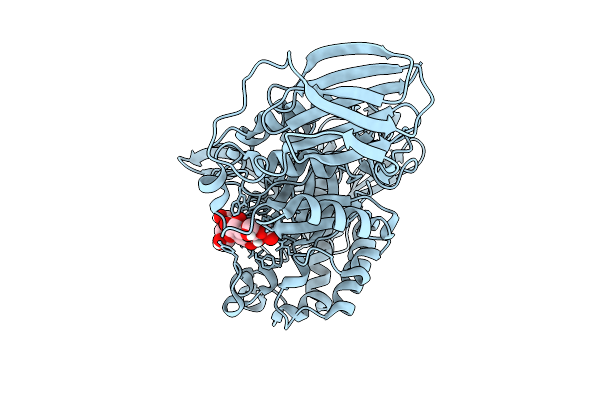 |
Mandh5 E303Q In Complex With Mannotriose After Co-Crystalliztion With Mannohexaose At 1.36 Angstroms Resolution A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2025-12-10 Classification: HYDROLASE |
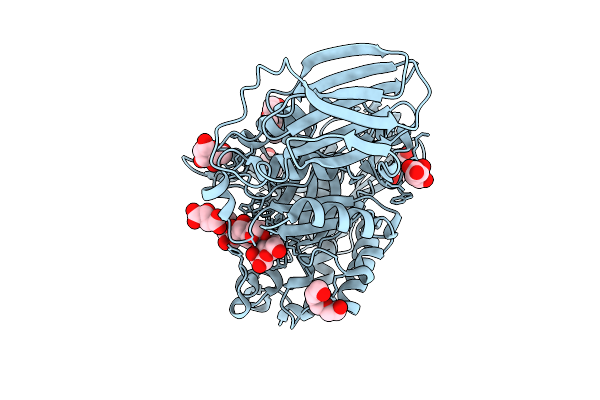 |
Mandh5 E303Q In Complex With Mannotetraose After Co-Crystalliztion With Mannotetraose At 1.6 Angstroms Resolution A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-12-03 Classification: HYDROLASE Ligands: BMA, MLT, PG4, PGE |
 |
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: P6G, PGE, PG4, TRS, MLT, 1PE |
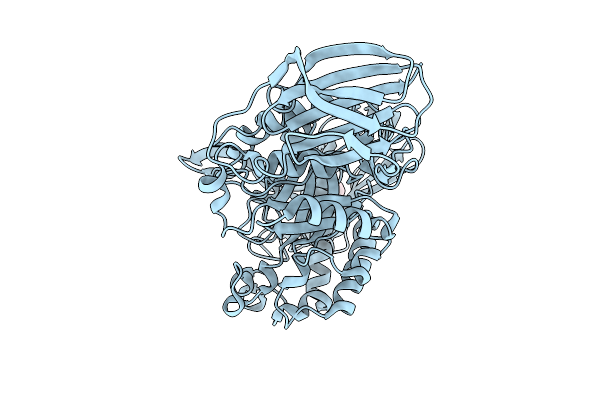 |
The Crystal Structure Of Mandh5 E303Q Mutant At 1.4 Angstroms Resolution - A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: GOL |
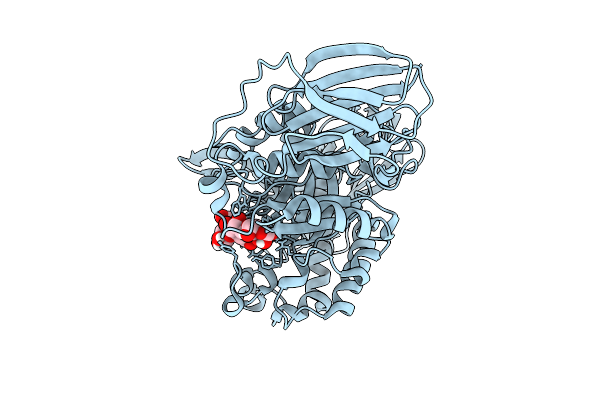 |
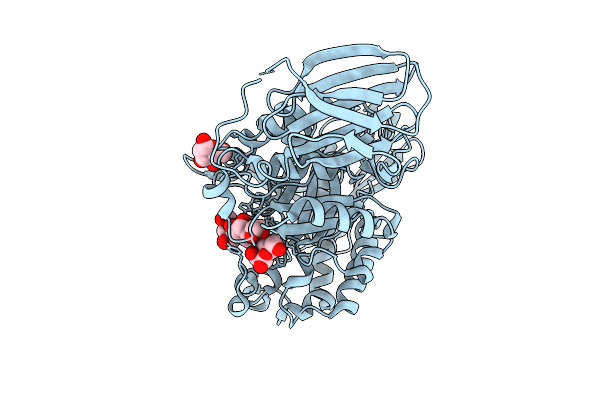
Mandh5 E303Q Mutant In Complex With Mannotriose A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2025-09-17 Classification: HYDROLASE |
 |
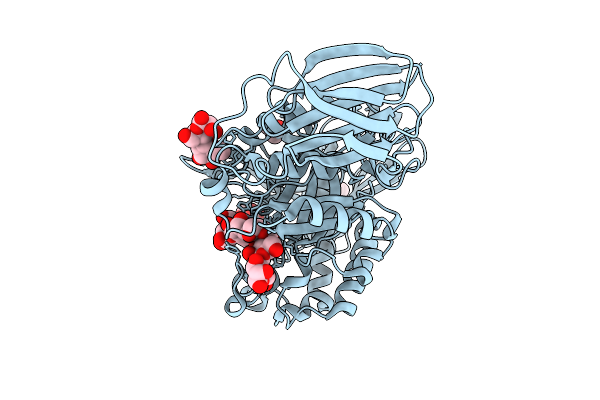
Mandh5 E303Q In Complex With Mannotetraose After Co-Crystalliztion With Mannotetraose At 1.6 Angstroms Resolution A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: BMA |
 |
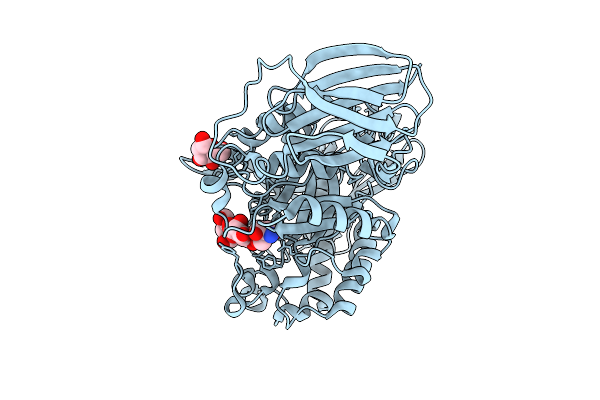
Mandh5 E303Q In Complex With Mannopentaose After Co-Crystalliztion With Mannopentaose At 1.6 Angstroms Resolution A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: GOL |
 |
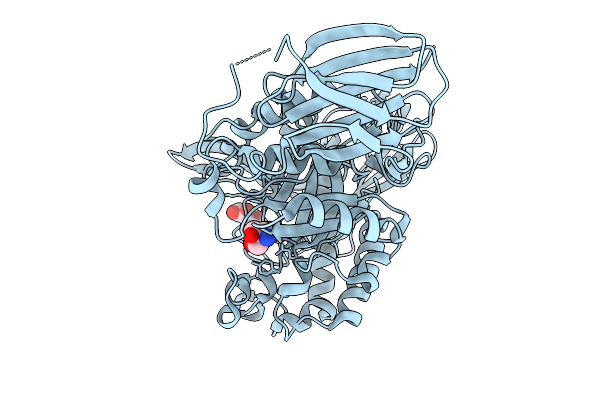
Complex Of Mandh5 Native Enzyme With Mannobiose After Co Crystallization With Mannopentaose At 1.7 Angstroms Resolution- A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-09-17 Classification: HYDROLASE Ligands: BMA, TRS |
 |
The Crystal Structure Of Mandh5 Selenomethionine Derivative At 1.6 Angstroms Resolution - A Beta-D-Mannanase Of Gh5 Family From Dictyoglomus Thermophilium
Organism: Dictyoglomus thermophilum
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-09-10 Classification: HYDROLASE Ligands: GOL, TRS |
 |
The Structure Of Xynx, A Nif3 Family Protein From Geobacillus Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-08-06 Classification: UNKNOWN FUNCTION Ligands: EDO, ZN |
 |
The Rec Domain (In The Non-Phosphorylated State) Of Xync, A Response Regulator From G.Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2025-06-25 Classification: DNA BINDING PROTEIN Ligands: MG |
 |
Organism: Acetivibrio thermocellus dsm 1313
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-06-25 Classification: SUGAR BINDING PROTEIN |
 |
The Structure Of Cbm42 From Clostridium Thermocellum In Complex With Arabinofuranosyl Xylobiose (A3X) Substrate Molecules
Organism: Acetivibrio thermocellus dsm 1313
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-06-25 Classification: SUGAR BINDING PROTEIN Ligands: CL, GOL |
 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: ZN, CL |
 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: PG4, CL, ZN |
 |
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: ZN, CL |
 |
The Structure Of Xt6 From G.Proteiniphilus T-6: The E265G/Q238A/W241A Mutant
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2025-06-18 Classification: HYDROLASE Ligands: CL, ZN |
 |
The Structure Of Glycosynthase Ixt6 (E241G Mutant), The Intracellular Xylanase Of G.Proteiniphilus T-6
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-06-18 Classification: HYDROLASE |
 |
The Structure Of Glycosynthase Ixt6 (E241G Mutant), The Intracellular Xylanase Of G.Proteiniphilus T-6 In Complex With Xylobiose-F Molecule
Organism: Geobacillus proteiniphilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-06-18 Classification: HYDROLASE |

