Search Count: 149
All
Selected
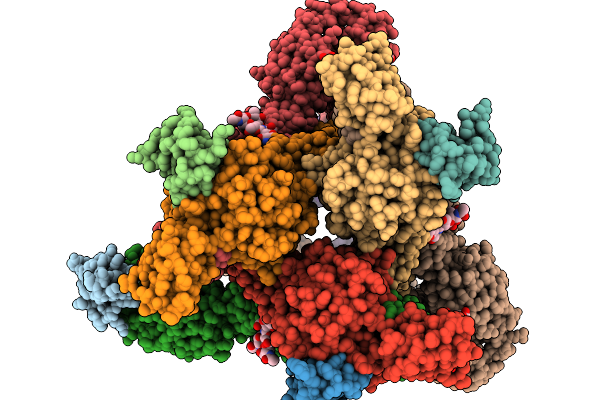 |
Organism: Homo sapiens, Mus musculus, Semliki forest virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS Ligands: CA |
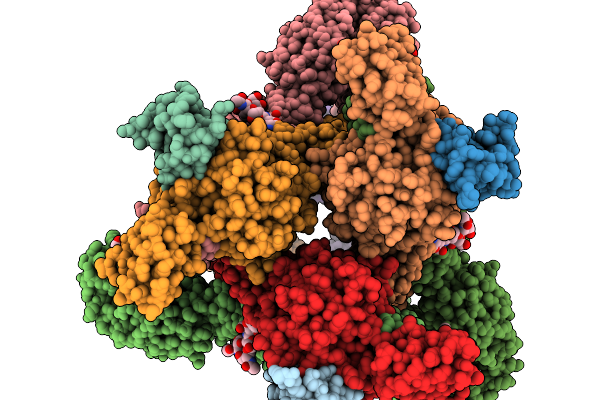 |
Organism: Semliki forest virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: MEMBRANE PROTEIN Ligands: NAG, A1ESQ |
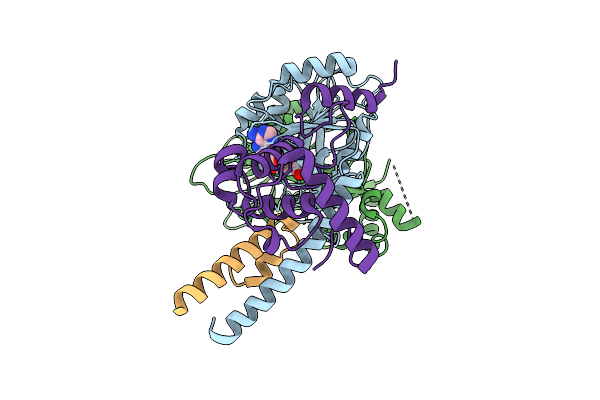 |
Organism: Tetrahymena thermophila sb210
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: DNA BINDING PROTEIN Ligands: SAM |
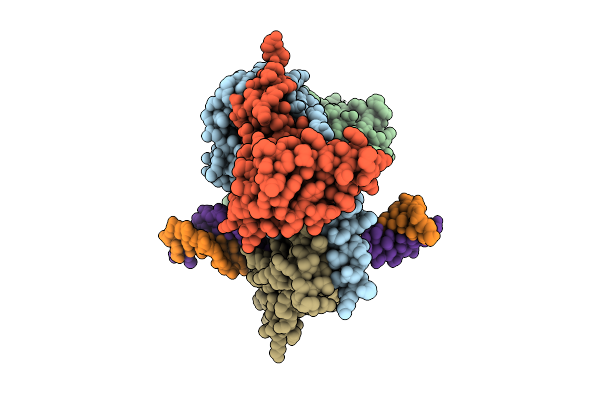 |
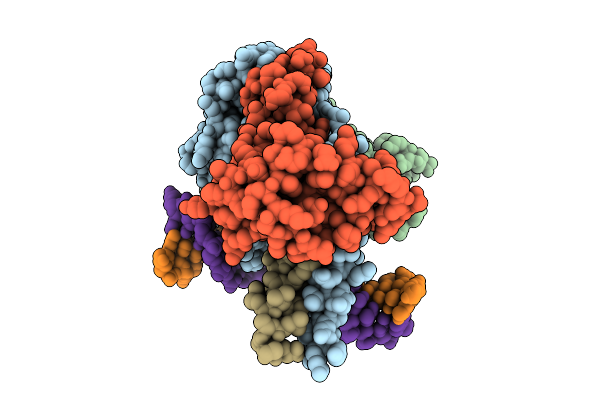
Cryo-Em Structure Of Hemimethylated Dna-Bound Tetrahymena Dna Methyltransferase Complex Mta1C
Organism: Tetrahymena thermophila sb210, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: DNA BINDING PROTEIN/DNA Ligands: SAM |
 |
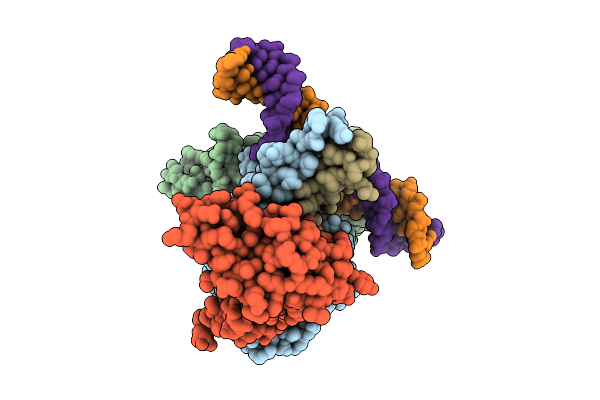
Cryo-Em Structure Of Hemi-Methylated Dna-Bound Tetrahymena Dna Methyltransferase Complex Mta1C (Mta9-B)
Organism: Tetrahymena thermophila sb210, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: DNA BINDING PROTEIN/DNA Ligands: SAM |
 |
Cryo-Em Structure Of Unmethylated Dna-Bound Tetrahymena Dna Methyltransferase Complex Mta1C (Mta9-B)
Organism: Tetrahymena thermophila sb210, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-11-05 Classification: DNA BINDING PROTEIN/DNA Ligands: SAM |
 |
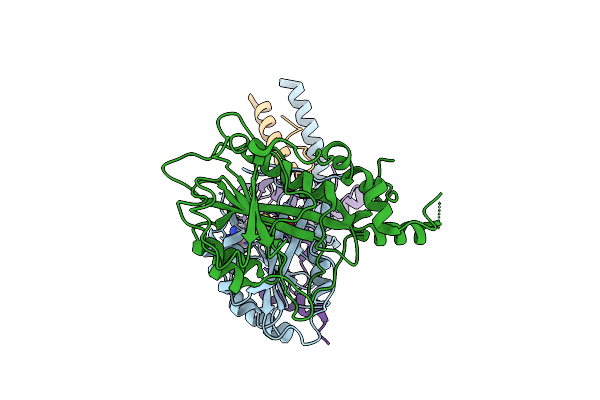
Organism: Tetrahymena thermophila sb210
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: DNA BINDING PROTEIN |
 |
Cryo-Em Structure Of Sah-Bound Tetrahymena Dna Methyltransferase Complex Mta1C
Organism: Tetrahymena thermophila sb210
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: DNA BINDING PROTEIN Ligands: SAH |
 |
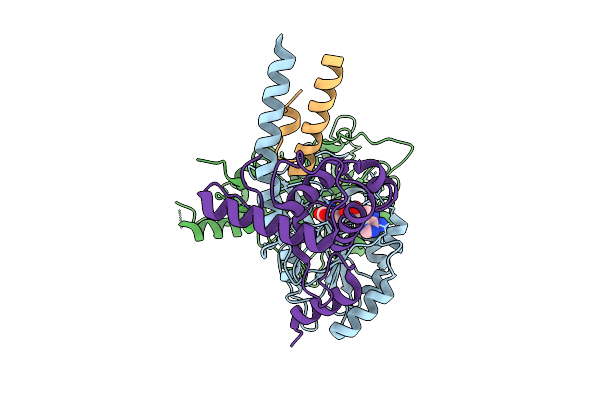
Cryo-Em Structure Of Sam-Bound Tetrahymena Dna Methyltransferase Complex Mta1C
Organism: Tetrahymena thermophila sb210
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: DNA BINDING PROTEIN Ligands: SAM |
 |
Cryo-Em Structure Of Sam-Bound Tetrahymena Dna Methyltransferase Complex Mta1C (D209A)
Organism: Tetrahymena thermophila sb210
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: DNA BINDING PROTEIN Ligands: SAM |
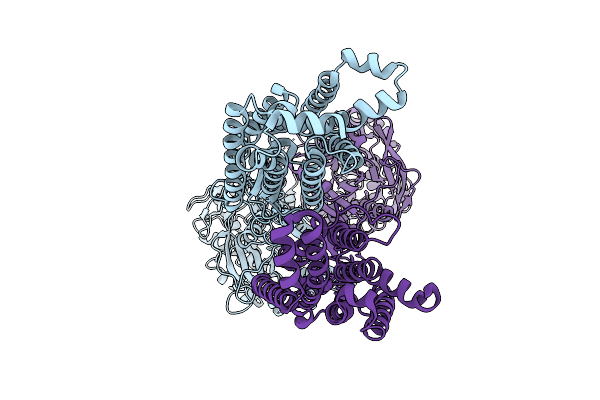 |
The Full-Length Human Sweet Taste Receptor Tas1R2 And Tas1R3 In The Apo State
Organism: Homo sapiens, Discosoma sp. (sea anemone), Branchiostoma lanceolatum
Method: ELECTRON MICROSCOPY Resolution:3.41 Å Release Date: 2025-07-16 Classification: SIGNALING PROTEIN |
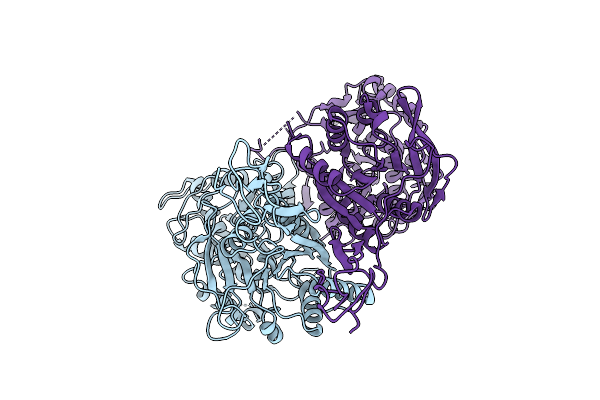 |
The Vft Domains Of Human Sweet Taste Receptor Tas1R2 And Tas1R3 In The Apo State
Organism: Homo sapiens, Discosoma sp. (sea anemone), Branchiostoma lanceolatum
Method: ELECTRON MICROSCOPY Resolution:3.18 Å Release Date: 2025-07-16 Classification: SIGNALING PROTEIN |
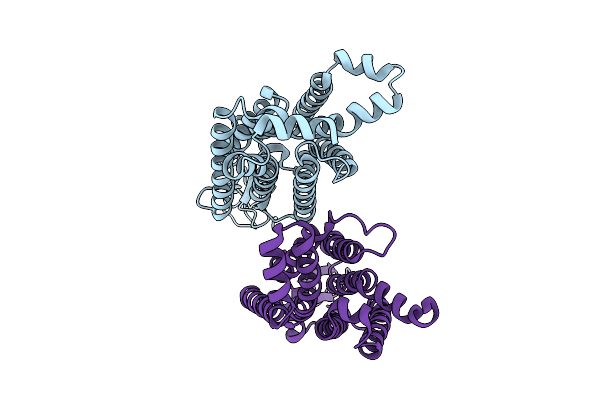 |
The Transmembrane Domains Of Human Sweet Taste Receptor Tas1R2 And Tas1R3 In The Apo State
Organism: Homo sapiens, Discosoma sp. (sea anemone), Branchiostoma lanceolatum
Method: ELECTRON MICROSCOPY Resolution:3.77 Å Release Date: 2025-07-16 Classification: SIGNALING PROTEIN |
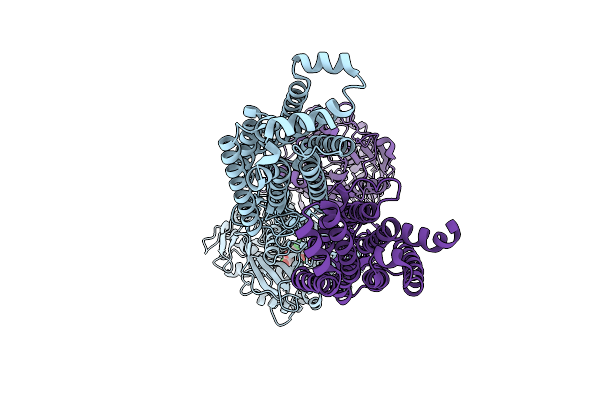 |
The Full-Length Human Sweet Taste Receptor Tas1R2 And Tas1R3 In The Sucralose-Bound State
Organism: Homo sapiens, Discosoma sp., Branchiostoma lanceolatum
Method: ELECTRON MICROSCOPY Resolution:3.59 Å Release Date: 2025-07-16 Classification: SIGNALING PROTEIN |
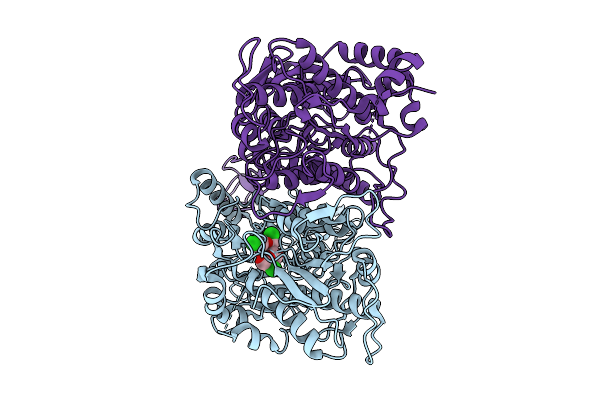 |
The Vft Domains Of Human Sweet Taste Receptor Tas1R2 And Tas1R3 In The Sucralose-Bound State
Organism: Homo sapiens, Discosoma sp., Branchiostoma lanceolatum
Method: ELECTRON MICROSCOPY Resolution:3.33 Å Release Date: 2025-07-16 Classification: SIGNALING PROTEIN |
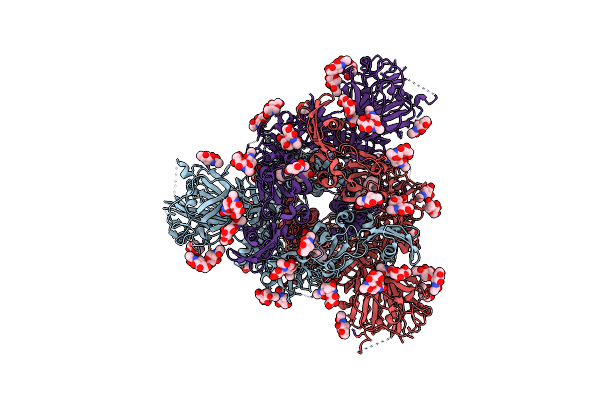 |
Organism: Coronavirus neoromicia/pml-phe1/rsa/2011, Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, EIC |
 |
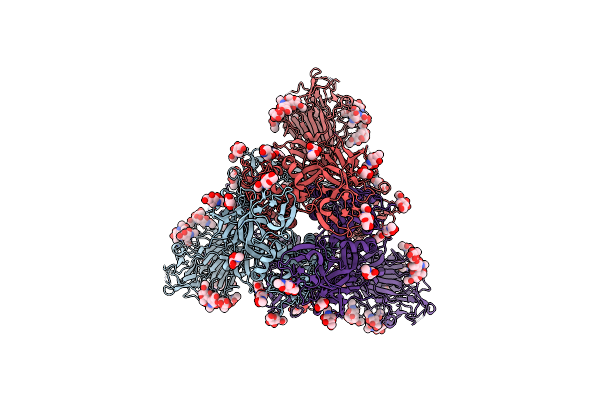
Cryo-Em Structure Of Eu-Hedgehogcov (Erinaceus/Vmc/Deu/2012) S-Trimer In A Locked-2 Conformation
Organism: Betacoronavirus erinaceus/vmc/deu/2012, Enterobacteria phage t4 (bacteriophage t4)
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: NAG, FOL, EIC |
 |
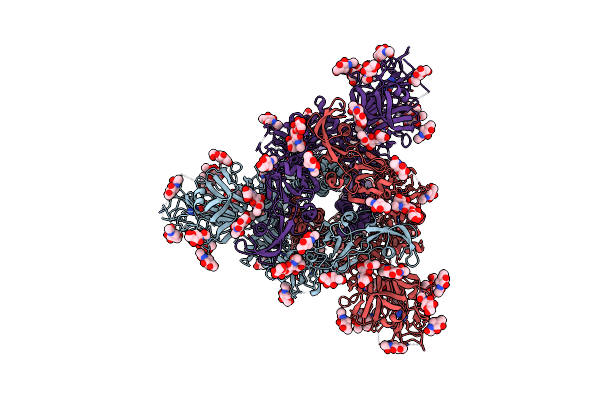
Cryo-Em Structure Of Hku25-Batcov S-Trimer Stabilized With 2P And X1 Disulfide Bond
Organism: Hypsugo bat coronavirus hku25, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: EIC, NAG |
 |
Cryo-Em Structure Of Cn-Hedgehogcov (Hku31/Erinaceus Amurensis/China/2014) S-Trimer In A Locked-2 Conformation
Organism: Erinaceus hedgehog coronavirus hku31, Enterobacteria phage t4
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: VIRAL PROTEIN Ligands: FOL, NAG |

