Search Count: 228
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.87 Å Release Date: 2026-03-25 Classification: CELL CYCLE Ligands: A1ENX |
 |
Organism: Schizosaccharomyces pombe (strain 972 / atcc 24843)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: POV, 8PE, PO4 |
 |
Organism: Schizosaccharomyces pombe 972h-
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: TRANSPORT PROTEIN Ligands: POV, 8PE, IHP, PO4 |
 |
Crystal Structure Of H5N1 Influenza Polymerase Pb2 Domain In Complex With Compound 3
Organism: Influenza a virus (a/duck/shantou/4610/2003(h5n1))
Method: X-RAY DIFFRACTION Resolution:0.91 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: A1JV9, SO4 |
 |
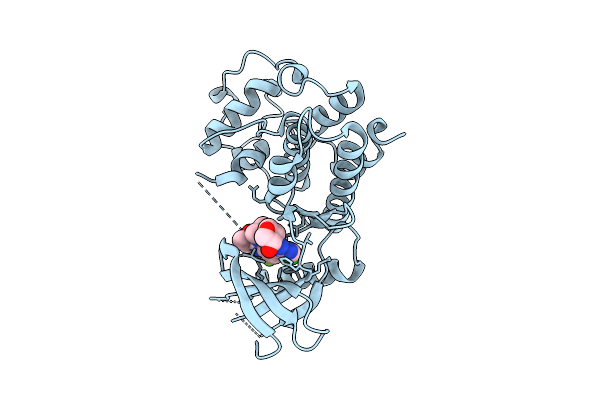
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-12-17 Classification: TRANSFERASE/INHIBITOR Ligands: A1CIF |
 |
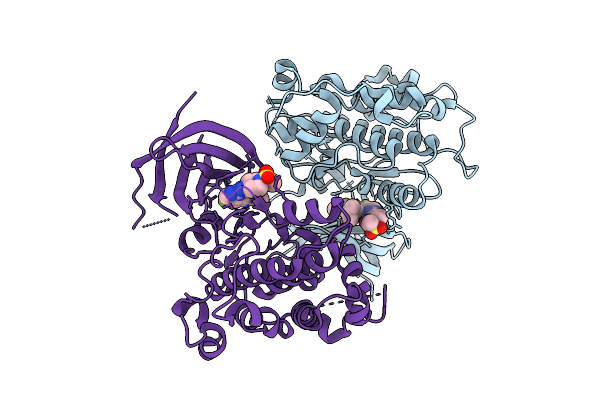
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-17 Classification: TRANSFERASE/INHIBITOR Ligands: A1AZ4 |
 |
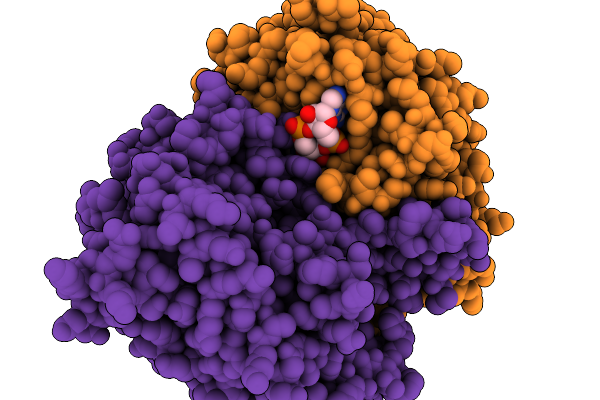
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2025-12-17 Classification: TRANSFERASE/INHIBITOR Ligands: A1CIF |
 |
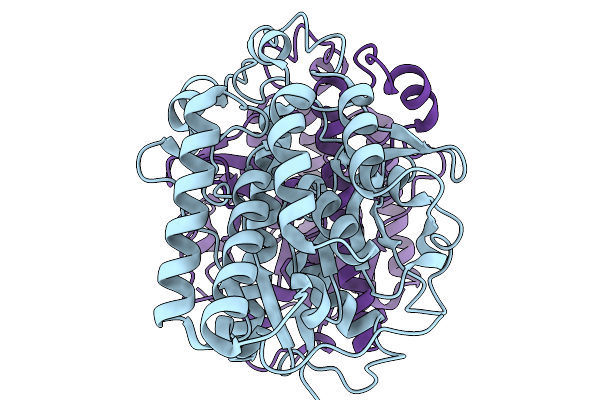
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: HYDROLASE Ligands: 4BW |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2025-11-19 Classification: HYDROLASE |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-11-19 Classification: HYDROLASE |
 |
Crystal Structure Of The Bromodomain Of Human Brd9 In Complex With The Inhibitor Y22076
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-11-19 Classification: PROTEIN BINDING Ligands: A1EKN, DMS |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2025-11-12 Classification: HYDROLASE Ligands: 4BW |
 |
Crystal Structure Of The Bromodomain Of Human Brd9 In Complex With The Inhibitor Y22024
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-11-05 Classification: PROTEIN BINDING Ligands: A1EF0, GOL, MLT |
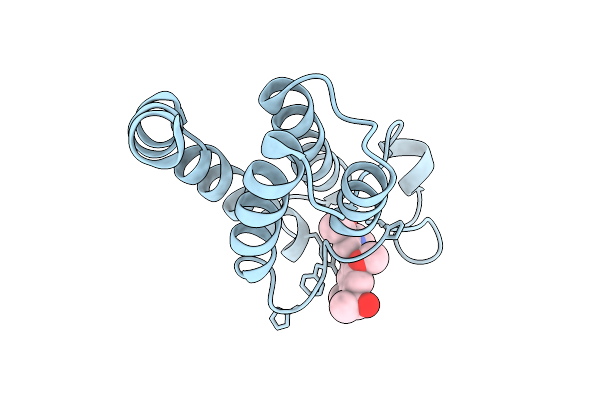 |
Crystal Structure Of The Bromodomain Of Human Brd9 In Complex With The Inhibitor Y22032
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-11-05 Classification: PROTEIN BINDING Ligands: A1EHA |
 |
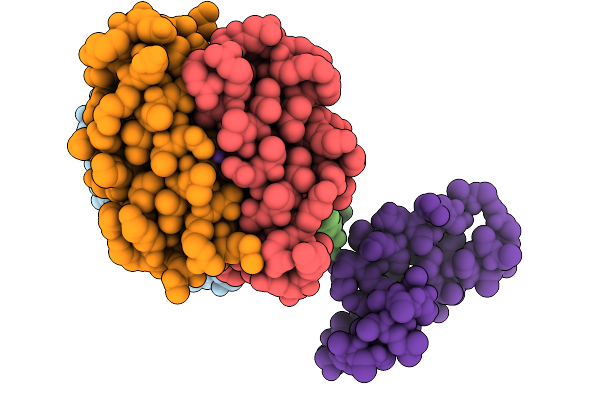
Crystal Structure Of The Bromodomain Of Human Brd9 In Complex With The Inhibitor Y22077
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-11-05 Classification: PROTEIN BINDING Ligands: A1EFZ, EDO |
 |
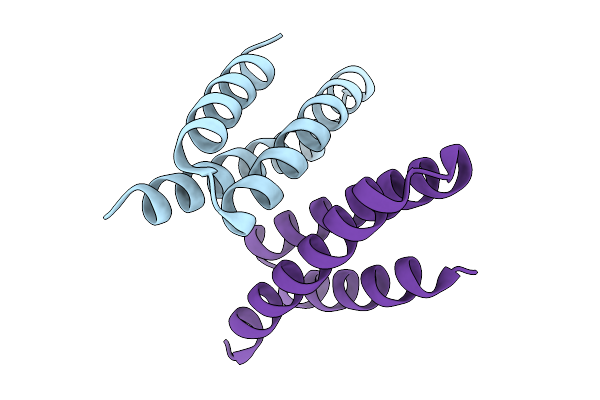
Crystal Structure Of De Novo Designed Light-Responsive Oligomer C2-5 (Lro-C2-5)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-08-27 Classification: DE NOVO PROTEIN |
 |
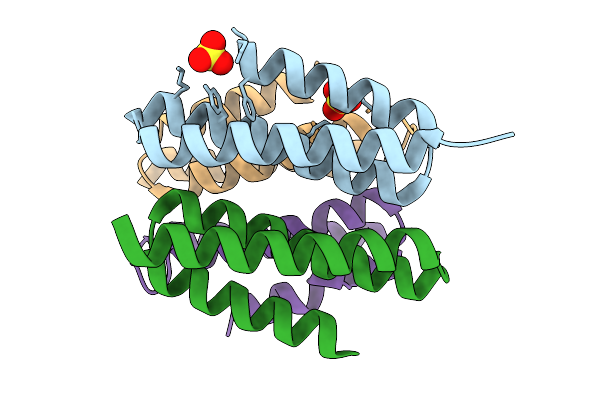
Crystal Structure Of De Novo Designed Light-Responsive Oligomer C2-35 (Lro-C2-35) At Acidic Ph (Ph 4.5)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-08-27 Classification: DE NOVO PROTEIN |
 |
Crystal Structure Of De Novo Designed Light-Responsive Oligomer C2-35 (Lro-C2-35) At Basic Ph (Ph 8.5)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-08-27 Classification: DE NOVO PROTEIN Ligands: SO4 |
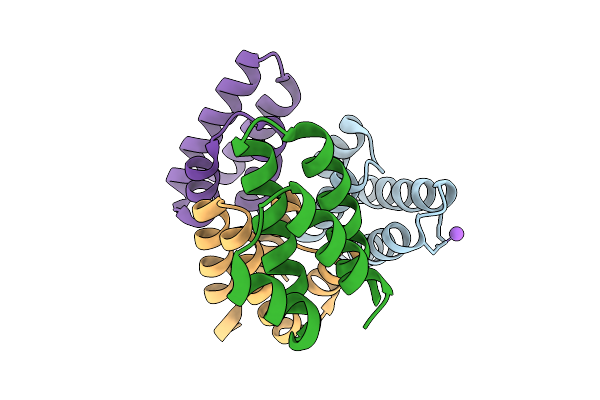 |
Crystal Structure Of De Novo Designed Light-Responsive Oligomer C3-7 (Lro-C3-7)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2025-08-27 Classification: DE NOVO PROTEIN |
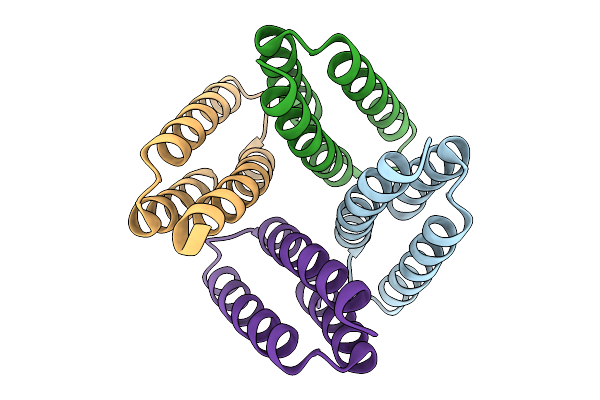 |
Crystal Structure Of De Novo Designed Light-Responsive Oligomer C4-13 (Lro-C4-13)
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-08-27 Classification: DE NOVO PROTEIN |

