Search Count: 64
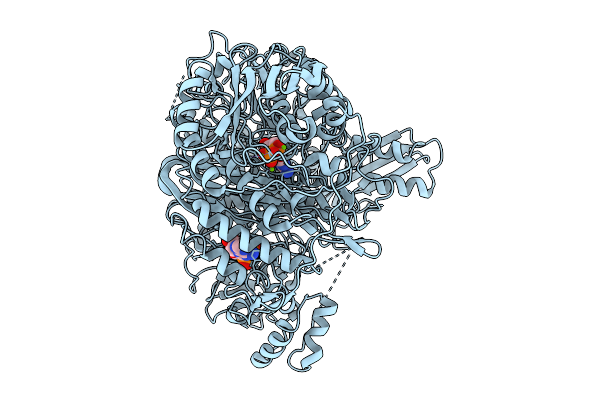 |
Cryo-Em Structure Of Amppnp Bound Human Phosphoribosylformylglycinamidine Synthase
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.31 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: ADP, ANP, MG |
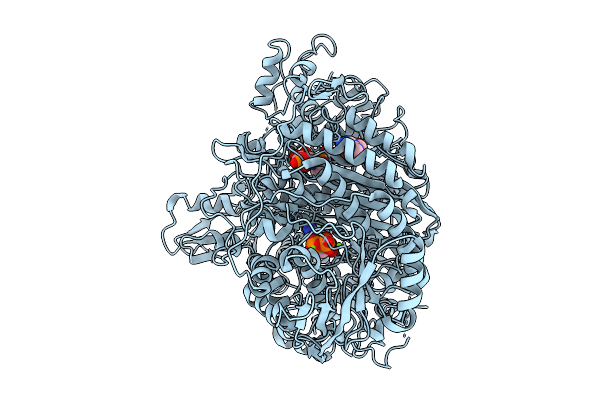 |
Cryo-Em Structure Of Ternary Complex Of Human Phosphoribosylglycinamidine Synthase With The Intermediate (Iminophosphate) And Adp Bound At The Synthase Site.
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: ADP, A1B7I, MG |
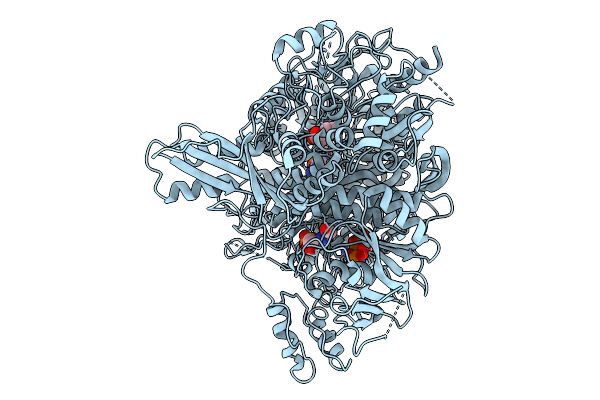 |
Cryo-Em Structure Of Quaternary Complex Of Human Phosphoribosylglycinamidine Synthase With Thioester Intermediate Bound (At Glutaminase Site) And Amppnp And Fgar (At Synthase Site).
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.19 Å Release Date: 2026-02-25 Classification: LIGASE Ligands: FGR, ANP, ADP, GLN, MG |
 |
Organism: Tulane virus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN |
 |
Organism: Tulane virus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-02-25 Classification: VIRAL PROTEIN Ligands: AG7 |
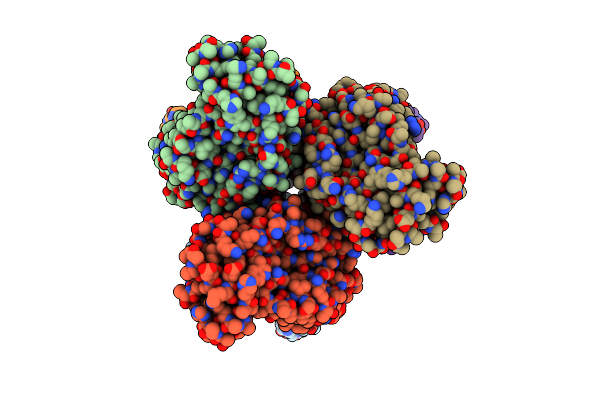 |
Cryo-Em Structure Of Burkholderia Cenocepacia Orotate Phosphoribosyltransferase
Organism: Burkholderia cenocepacia j2315
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: TRANSFERASE Ligands: ACT |
 |
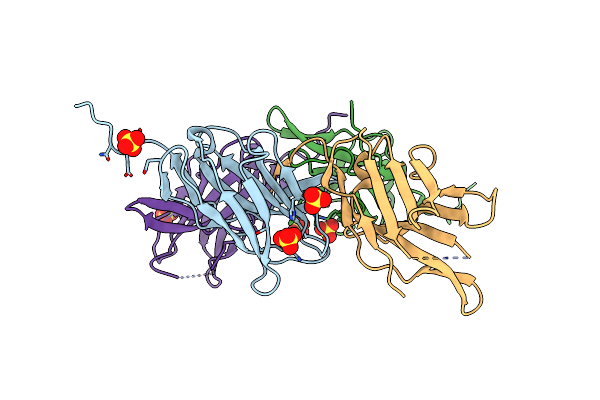
Organism: Rhodococcus sp. (in: high g+c gram-positive bacteria)
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2024-04-17 Classification: HYDROLASE Ligands: TN9, CL, SO4 |
 |
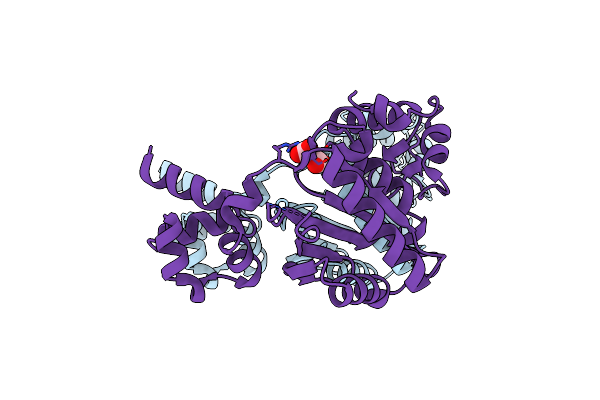
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2023-10-11 Classification: PROTEIN BINDING Ligands: SO4 |
 |
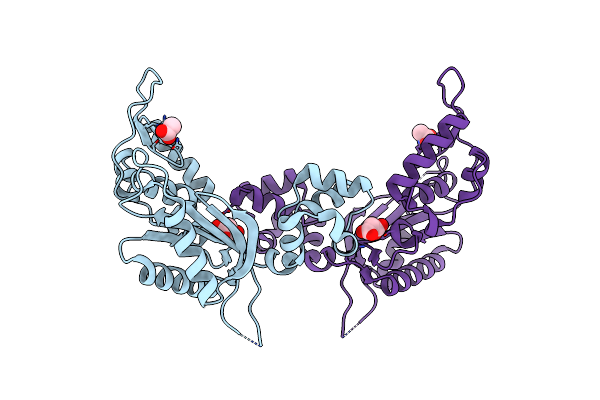
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-02-01 Classification: TRANSFERASE Ligands: MLA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-02-01 Classification: TRANSFERASE Ligands: ACT, MLA |
 |
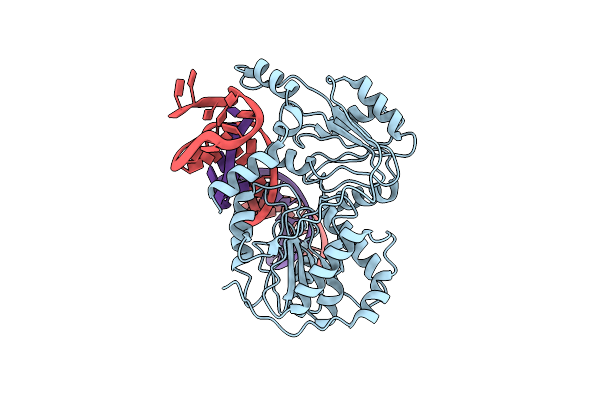
Structure Of The Sars-Cov2 Plpro (C111S) In Complex With A Dimeric Ubv That Inhibits Activity By An Unusual Allosteric Mechanism
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2023-01-25 Classification: VIRAL PROTEIN/SIGNALING PROTEIN Ligands: ZN, BR, NA, CL |
 |
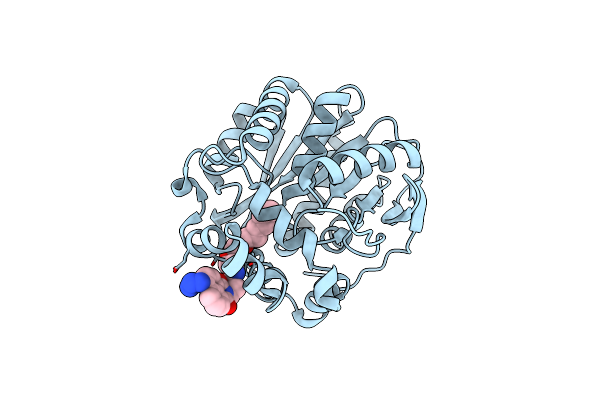
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2022-05-25 Classification: DNA BINDING PROTEIN/DNA |
 |
Organism: Rhodococcus sp.
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: 8MH, CL |
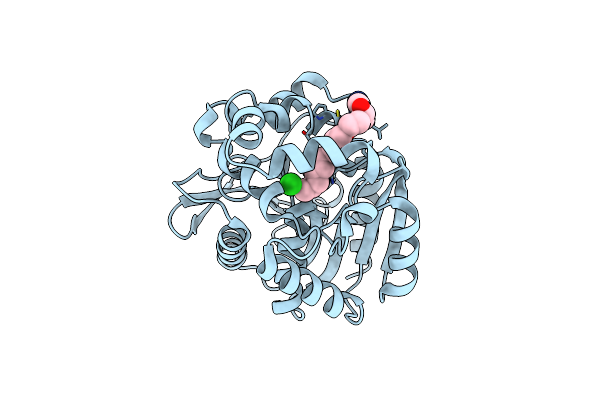 |
Organism: Rhodococcus sp.
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2022-02-02 Classification: HYDROLASE Ligands: 8MS, CL |
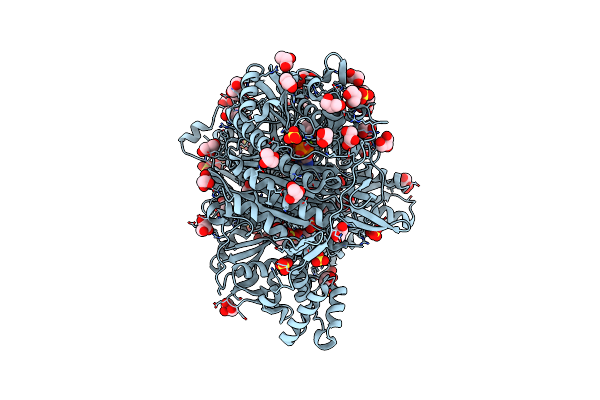 |
Organism: Salmonella typhimurium
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-01-19 Classification: LIGASE Ligands: ADP, GOL, SO4, MG |
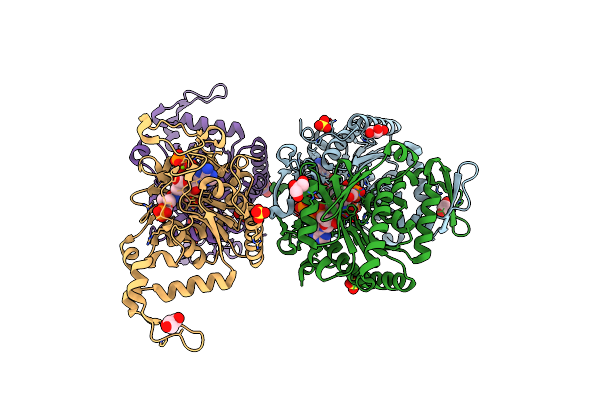 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-11-24 Classification: LIGASE Ligands: ANP, SO4, MG, J1O, GOL |
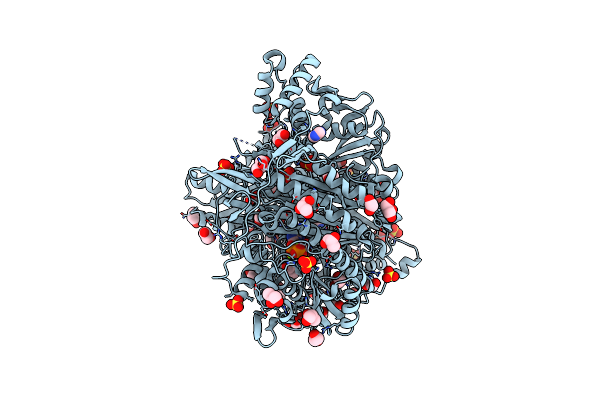 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2021-02-17 Classification: LIGASE Ligands: EDO, IMD, ADP, GLN, MG, SO4, GOL |
 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2021-02-17 Classification: LIGASE Ligands: SO4, MG, ADP, GOL, EDO |
 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2021-02-17 Classification: LIGASE Ligands: ADP, MG, SO4, GOL, EDO |
 |
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2021-02-17 Classification: LIGASE Ligands: ADP, GOL, EDO, SO4, MG |

