Search Count: 655
All
Selected
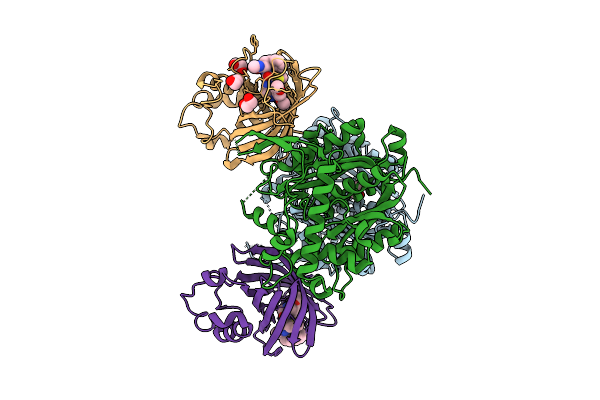 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-15 Classification: PEPTIDE BINDING PROTEIN Ligands: L6U, EDO |
 |
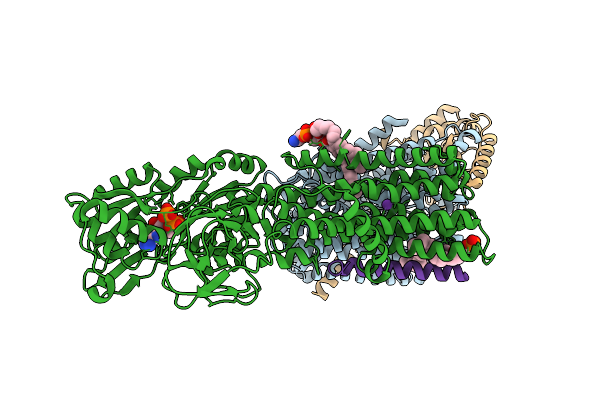
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1 State In Lipid Nanodisc
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Resolution:2.34 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1-Atp State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.28 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, ATP |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E1P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0, MG |
 |
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2P State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.58 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
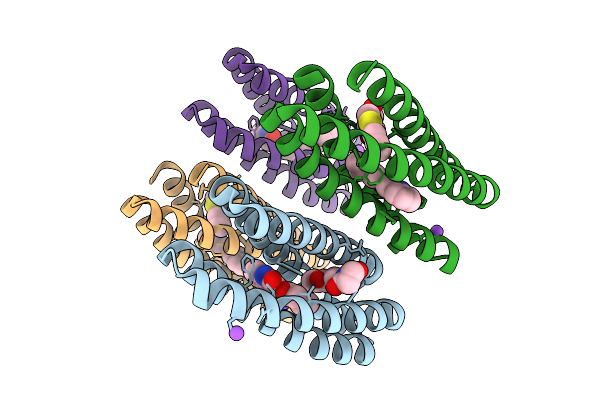
High-Resolution Cryo-Em Structure Of Kdpfabc In The E2 State In Lipid Nanodisc
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.72 Å Release Date: 2026-04-01 Classification: TRANSPORT PROTEIN Ligands: K, 9Y0 |
 |
De Novo Photoenzyme Variant Able-E116C-Txt H49A-L42T (Photoable1 Precursor) In Complex With Substrate
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUV, A1JUW, NA |
 |
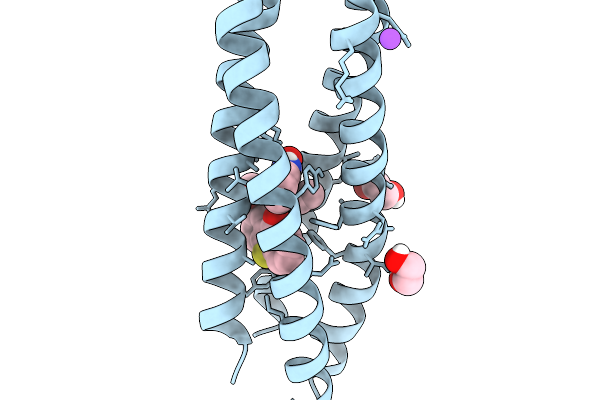
De Novo Photoenzyme Variant Able-E116C-Txt H49A-L42T (Photoable1 Precursor) In Complex With Product Formed In Crystallo After Irradiation
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUX, A1JUW, PGE, EDO, GOL |
 |
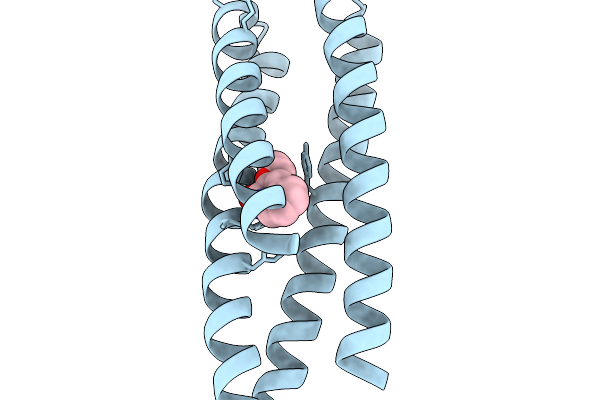
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUV, A1JUW, EDO, NA |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUW, NA, GOL, EDO |
 |
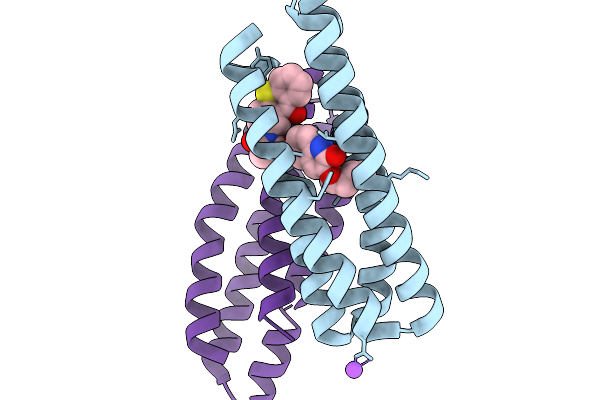
De Novo Photoenzyme Variant Able-E116C-Txt V111H-L53A (Photoable2 Precursor) In Complex With Substrate
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUV, NA |
 |
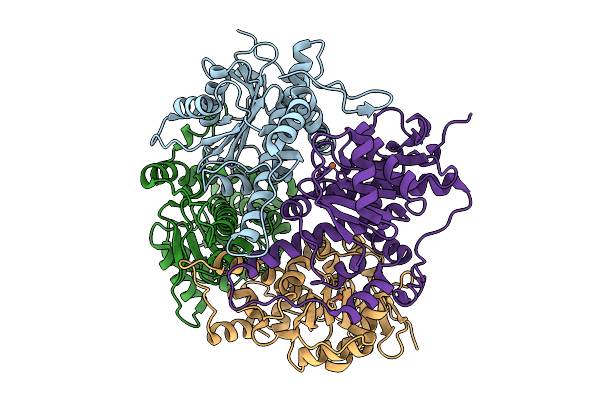
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUV, A1JUW, NA |
 |
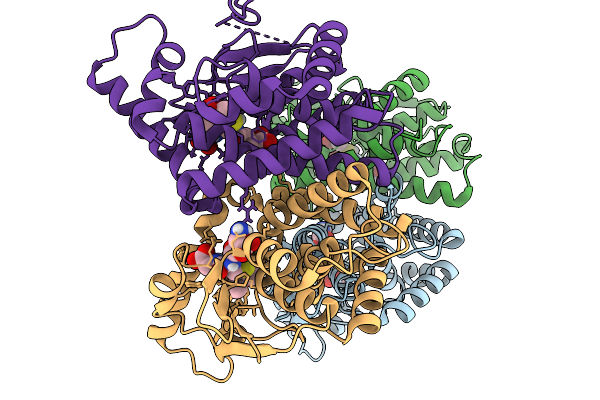
Crystal Structure Of None-Heme Iron Enzyme (Tqam) From Trichoderma Atroviride Bound With Iron
Organism: Trichoderma atroviride
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-28 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
Organism: Gordonia rubripertincta
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-01-14 Classification: TRANSFERASE Ligands: GSH, CAC |
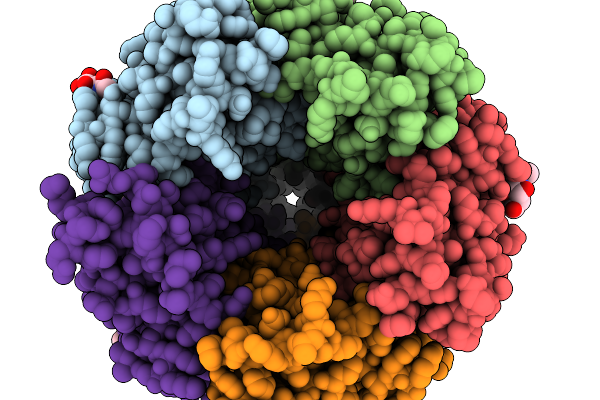 |
Alpha1/Beta Heteromeric Glycine Receptor In The Presence Of 0.200 Mm Strychnine And 500 Nm Ivermectin
Organism: Danio rerio
Method: ELECTRON MICROSCOPY Resolution:2.86 Å Release Date: 2026-01-07 Classification: TRANSPORT PROTEIN Ligands: SY9, NAG |
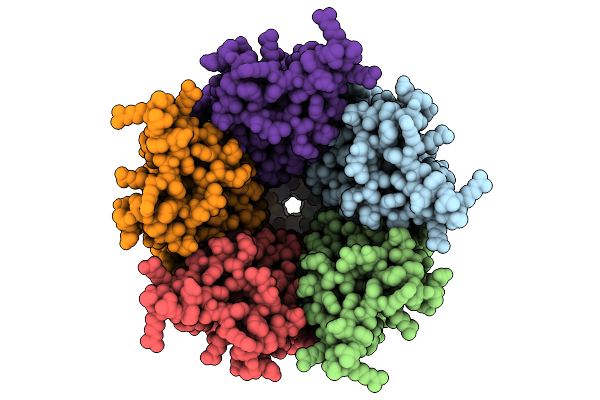 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiiii Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: PEE |
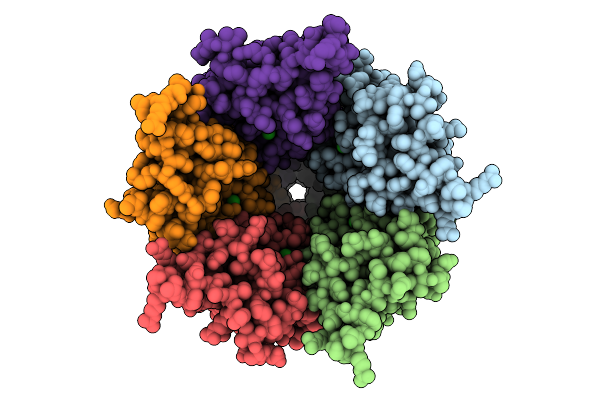 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiiio Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
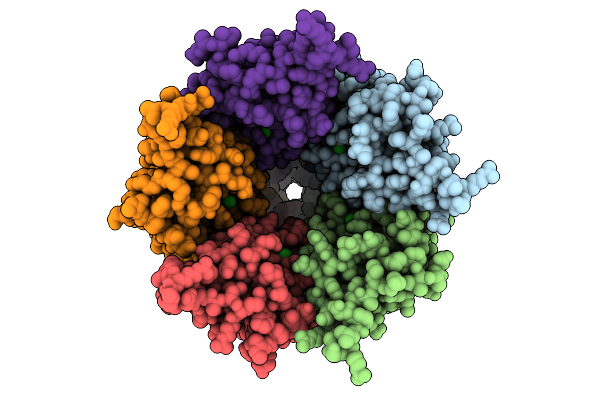 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiioo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
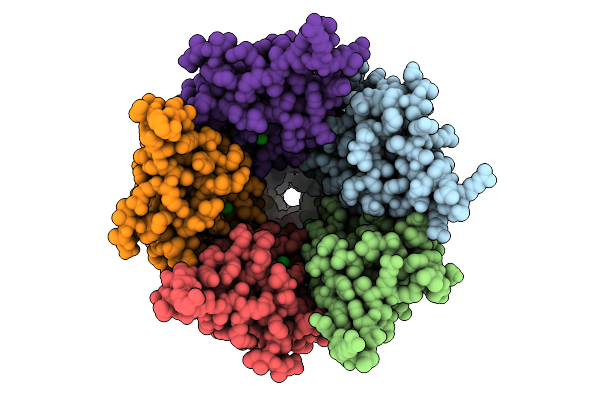 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Iiooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.21 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |
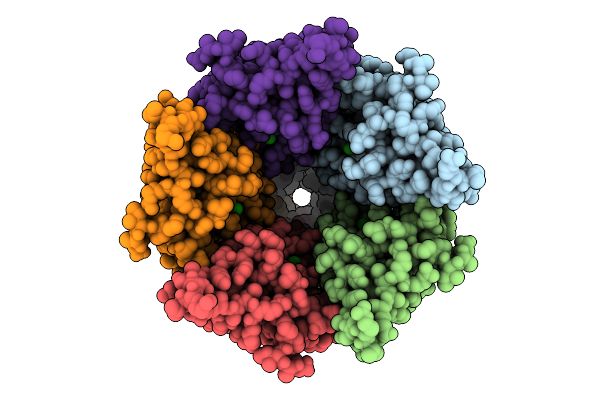 |
Cryo-Em Structure Of Nanodisc (Pe:Ps:Pc) Reconstituted Glic At Ph 4 In Ioooo Conformation
Organism: Gloeobacter violaceus
Method: ELECTRON MICROSCOPY Resolution:3.16 Å Release Date: 2025-10-22 Classification: MEMBRANE PROTEIN Ligands: CL, PEE |

