Search Count: 39
All
Selected
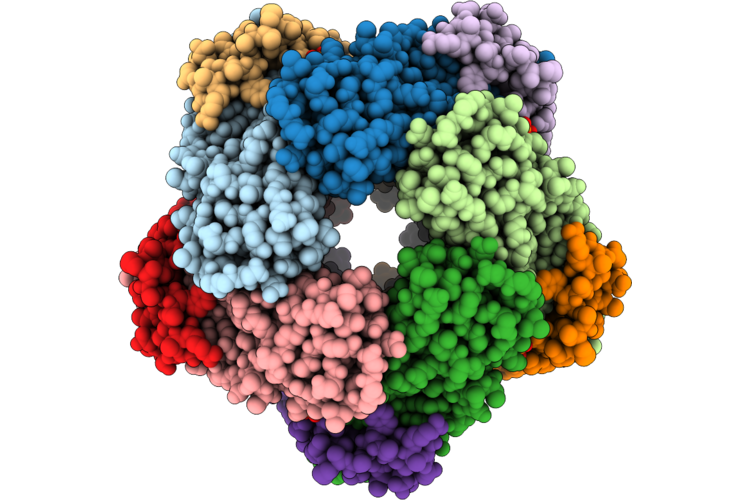 |
Organism: Streptomyces mobaraensis
Method: ELECTRON MICROSCOPY Resolution:2.01 Å Release Date: 2026-04-22 Classification: BIOSYNTHETIC PROTEIN Ligands: ZN, GTP |
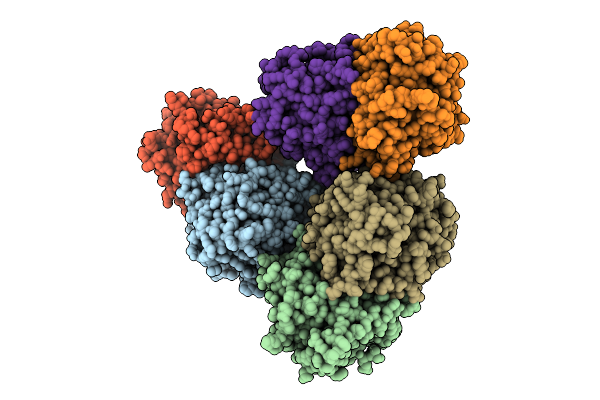 |
Sesterterpene Synthase From Streptomyces Mobaraensis (Sestermobaraene Synthase, Smts1) In Complex With Pyrophosphate (Ppi)
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: LYASE Ligands: MG, POP, PO4 |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And Imidazole
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 2,5,6-Triaminopyrimidin-4(3H)-One.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: A1L8V, IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GOL, A1L8W, MN, IMD |
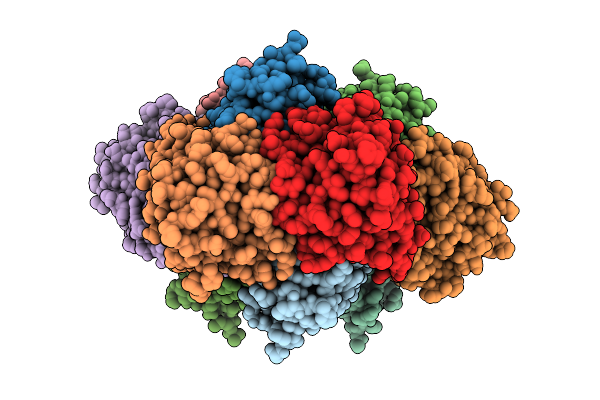 |
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Tpp.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-01 Classification: LYASE Ligands: ZN, TPP, MG, GOL |
 |
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Ttpp And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-01 Classification: LYASE Ligands: A1L8W, ZN, SO4, TZD, MG |
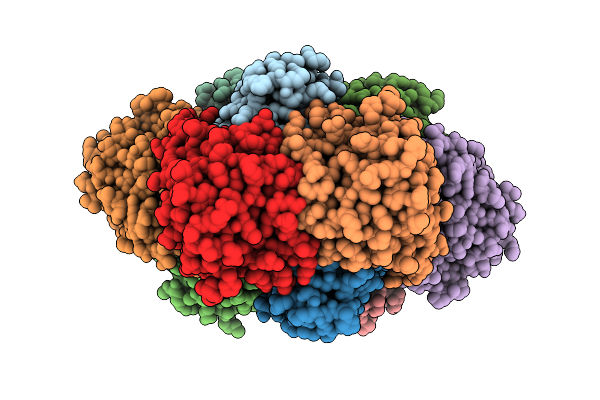 |
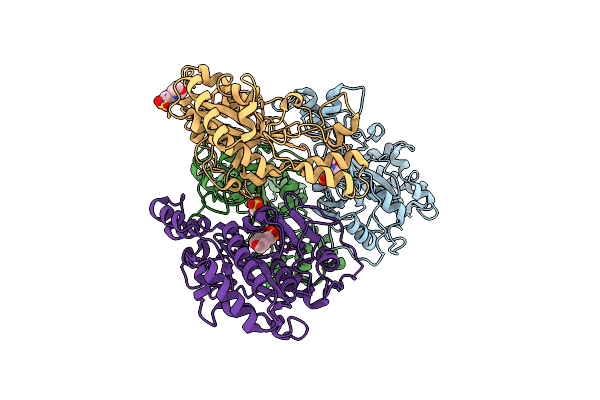
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Tpp And 5-Azacytosine.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2026-04-01 Classification: LYASE Ligands: 5AZ, ZN, TPP, MG, GOL |
 |
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-09-18 Classification: TRANSFERASE Ligands: MES, CL, PEG |
 |
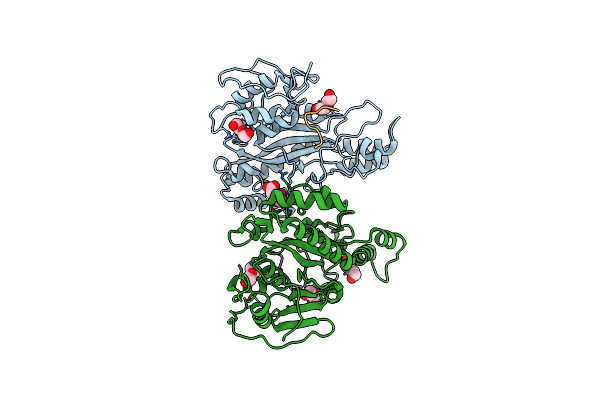
Organism: Streptomyces mobaraensis nbrc 13819 = dsm 40847
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2019-09-04 Classification: ANTIMICROBIAL PROTEIN |
 |
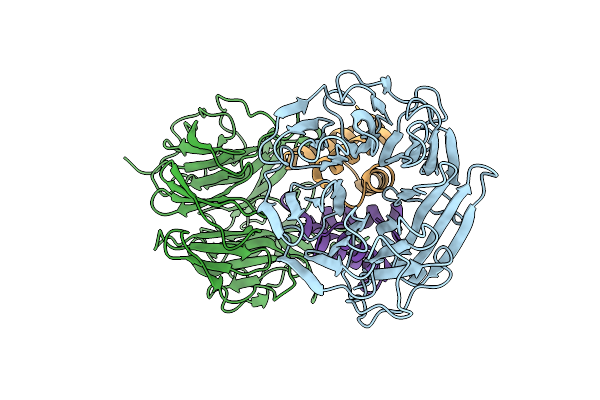
Structure Of A Glutamine Donor Mimicking Inhibitory Peptide Shaped By The Catalytic Cleft Of Microbial Transglutaminase
Organism: Streptomyces mobaraensis, Streptomyces mobaraensis nbrc 13819 = dsm 40847
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2018-10-24 Classification: TRANSFERASE Ligands: FLC, PEG |
 |
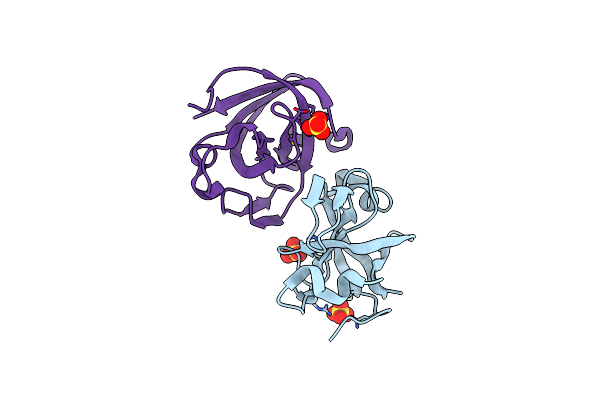
Organism: Streptomyces mobaraensis, Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2018-09-12 Classification: ANTIMICROBIAL PROTEIN |
 |
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2018-03-07 Classification: Protease inhibitor Ligands: SO4 |
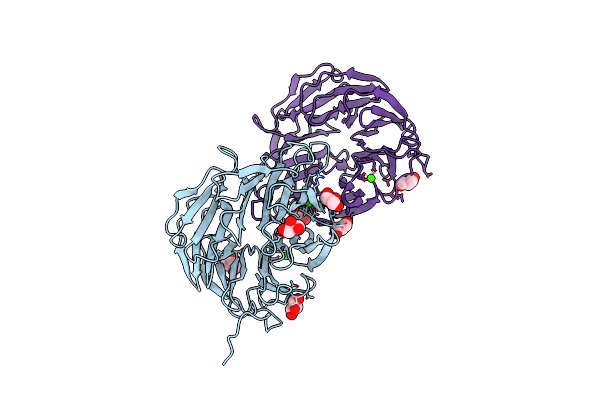 |
Structure Of The Dispase Autolysis Inducing Protein From Streptomyces Mobaraensis
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2016-08-10 Classification: SIGNALING PROTEIN Ligands: GOL, CA |
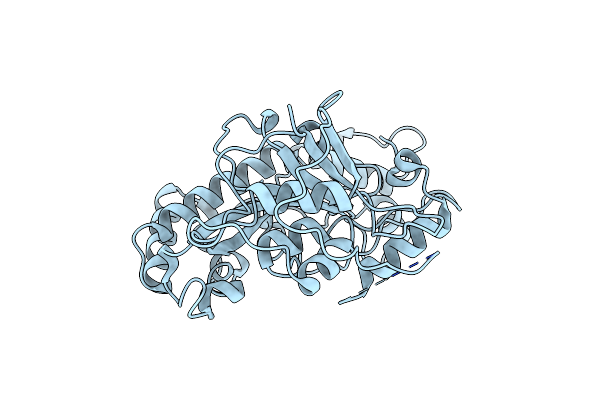 |
Structural Basis For Zymogen Activation And Substrate Binding Of Transglutaminase From Streptomyces Mobaraense
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2010-09-15 Classification: TRANSFERASE |
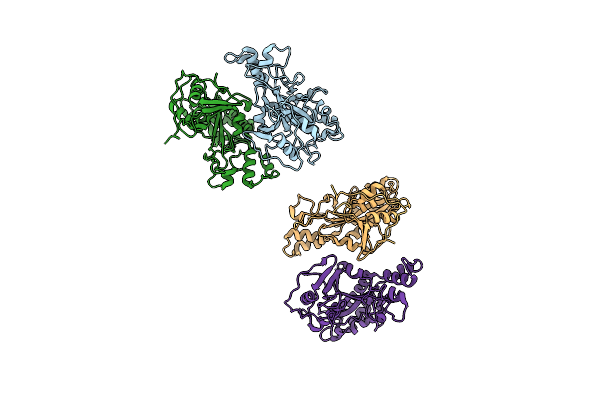 |
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2002-08-27 Classification: TRANSFERASE |
 |
Organism: Streptomyces mobaraensis NBRC 13819 = DSM 40847
Method: Alphafold Release Date: Classification: NA Ligands: NA |
 |
Organism: Streptomyces mobaraensis NBRC 13819 = DSM 40847
Method: Alphafold Release Date: Classification: NA Ligands: NA |
 |
Organism: Streptomyces mobaraensis NBRC 13819 = DSM 40847
Method: Alphafold Release Date: Classification: NA Ligands: NA |
 |
Organism: Streptomyces mobaraensis NBRC 13819 = DSM 40847
Method: Alphafold Release Date: Classification: NA Ligands: NA |

