Search Count: 8,522
All
Selected
 |
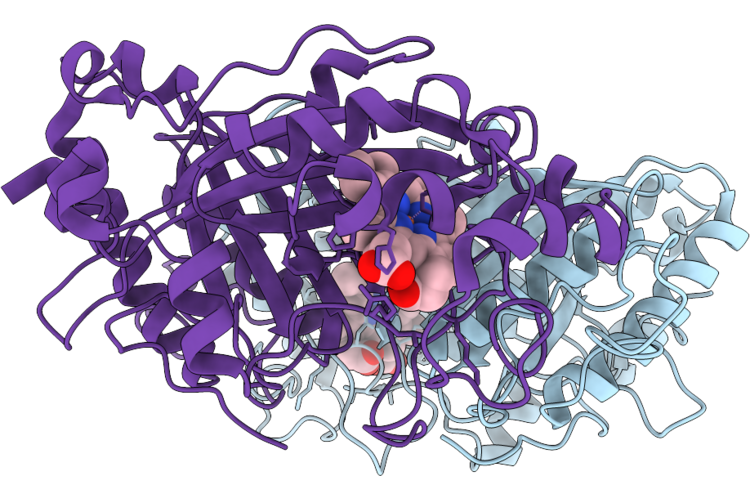
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
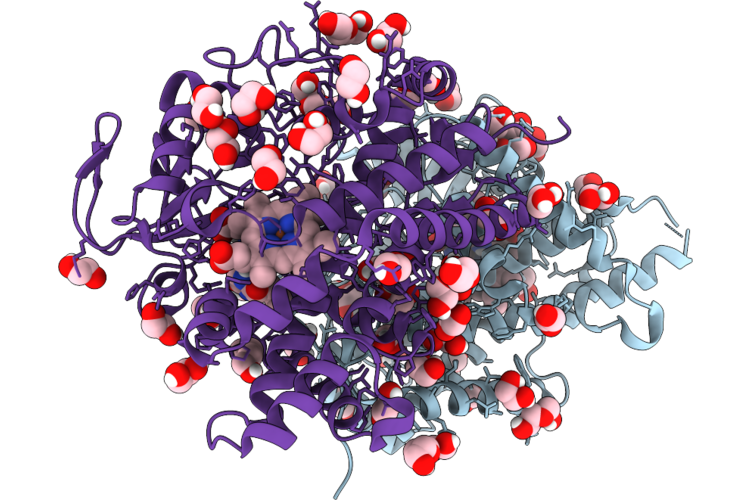
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
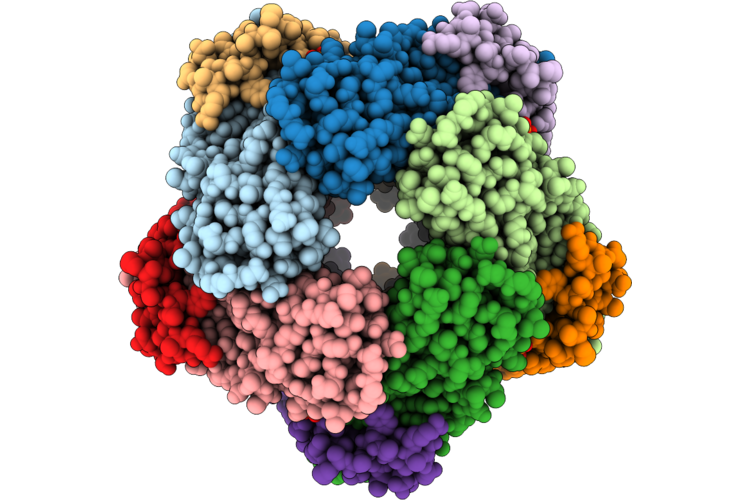
Organism: Streptomyces purpureus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GUN, EDO, PEG, UYM, CL, MG |
 |
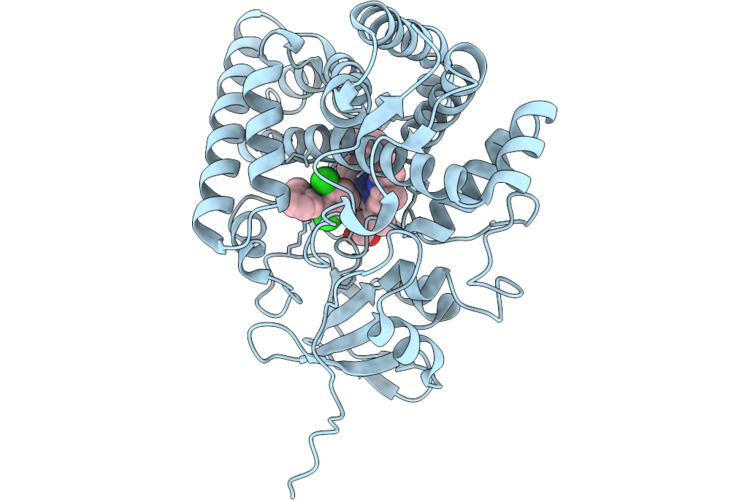
Organism: Streptomyces mobaraensis
Method: ELECTRON MICROSCOPY Resolution:2.01 Å Release Date: 2026-04-22 Classification: BIOSYNTHETIC PROTEIN Ligands: ZN, GTP |
 |
Crystal Structure Of Cyp105A1 R84A Complexed With Diclofenac (Dif) At Room Temperature
Organism: Streptomyces griseolus
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, DIF |
 |
Organism: Streptomyces lividans tk24
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Organism: Streptomyces purpureus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, GOL, EDO, CL, MG, NI |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, MG, O |
 |
Crystal Structure Of Pepstatin A Bound To Rhodotorula Mucilaginosa Aspartic Protease Rmuap1
Organism: Rhodotorula mucilaginosa, Streptomyces argenteolus subsp. toyonakensis
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-15 Classification: HYDROLASE/HYDROLASE INHIBITOR |
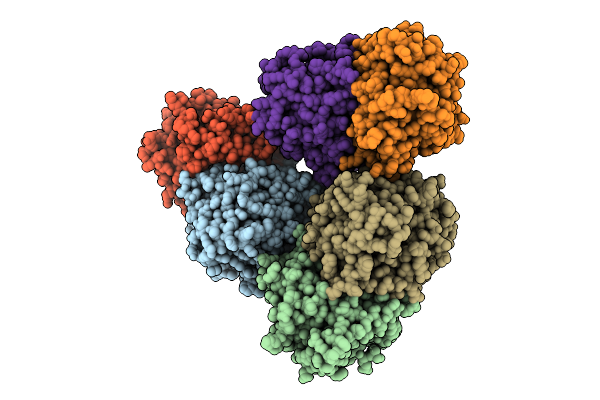 |
Sesterterpene Synthase From Streptomyces Mobaraensis (Sestermobaraene Synthase, Smts1) In Complex With Pyrophosphate (Ppi)
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: LYASE Ligands: MG, POP, PO4 |
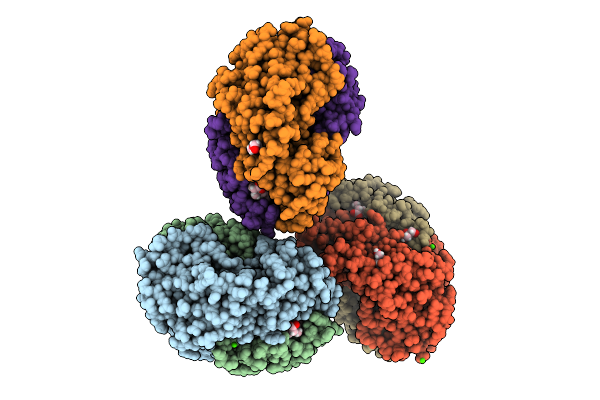 |
Organism: Streptomyces sp. b93
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HEM, GOL, CA, PEG |
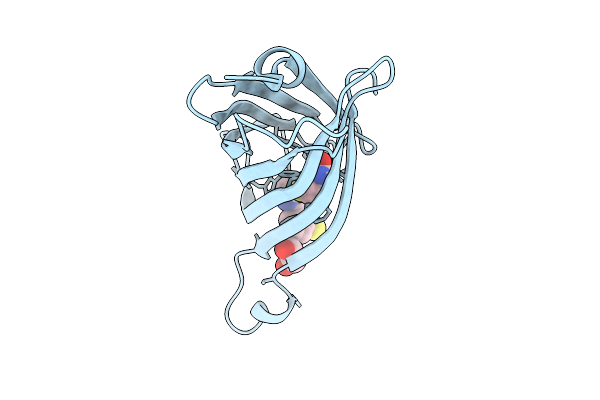 |
Streptavidin K121M With A Thiophenol Cofactor As Artificial Hydrogen Atom Transferase
Organism: Streptomyces avidinii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: A1I8C, EDO |
 |
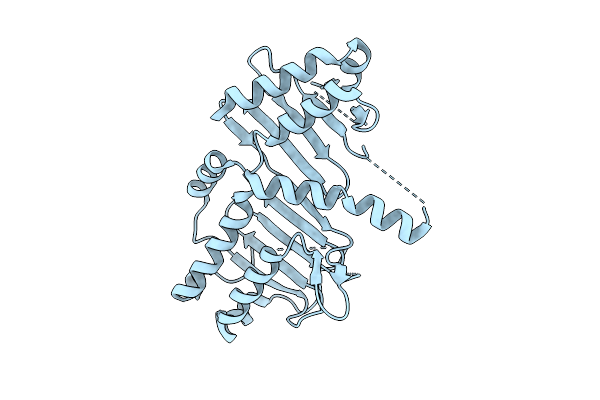
Organism: Streptomyces sp. nl15-2k
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
Organism: Streptomyces novoguineensis
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HOH |
 |
Crystal Structure Of The Cyp105D18 Double Mutant F184A/F191A From Streptomyces Laurentii
Organism: Streptomyces laurentii
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, GOL, DMS, EDO |
 |
Crystal Structure Of The Cyp105D18 Mutant F191A From Streptomyces Laurentii
Organism: Streptomyces laurentii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: HEM, GOL |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And Imidazole
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 2,5,6-Triaminopyrimidin-4(3H)-One.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: A1L8V, IMD, MN |
 |
Crystal Structure Of Cupin-Domain Containing Protein (Azca) Forming Triazine (6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid) Moiety From Streptomyces Mobaraensis In Complex With Manganese And 6-Amino-4-Oxo-4,5-Dihydro-1,3,5-Triazine-2-Carboxylic Acid.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GOL, A1L8W, MN, IMD |
 |
Crystal Structure Of Thiamine Pyrophosphate (Tpp)-Dependent Alpha-Imino Acid Decarboxylase (Azcb And Azcc) From Streptomyces Mobaraensis In Complex With Tpp.
Organism: Streptomyces mobaraensis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-01 Classification: LYASE Ligands: ZN, TPP, MG, GOL |

